Search Count: 417
All
Selected
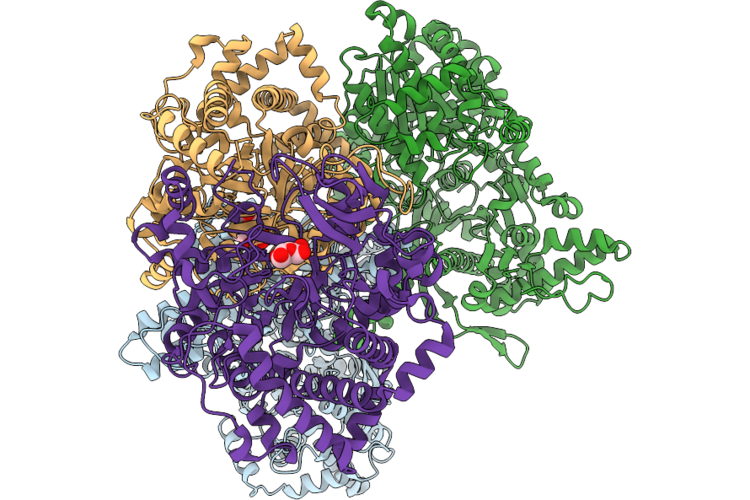 |
Cryo-Em Structure Of Maltose And Glucose Bound Rice Phs1(R709A)-Dpe1(D391A) Complex
Organism: Oryza sativa japonica group
Method: ELECTRON MICROSCOPY Resolution:3.07 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: GLC |
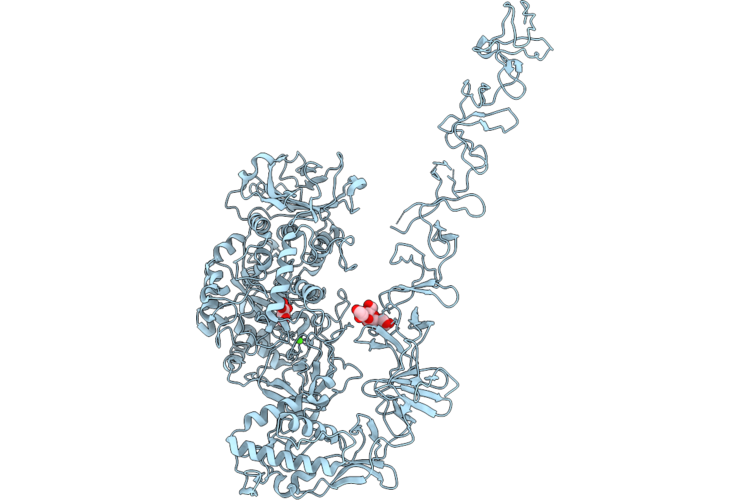 |
Structure Of Full-Length Streptococcus Mutans Gtfd Active Site Mutant (D465A, D584A) With Active Site Glucose
Organism: Streptococcus mutans
Method: X-RAY DIFFRACTION Resolution:3.15 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: CA, GLC, EDO |
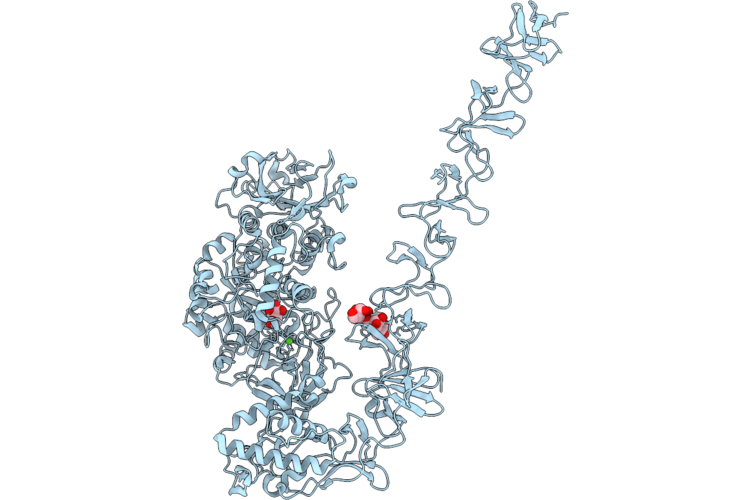 |
Structure Of Full-Length Streptococcus Mutans Gtfd In Complex With Dextran 5000 In Domain V
Organism: Streptococcus mutans
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: CA, GLC |
 |
Crystal Structure Of Gh57 Family Amylopullulanase From Aquifex Aeolicus Mutant D352N In Complex With Maltoheptaose
Organism: Aquifex aeolicus vf5
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: GLC, GOL |
 |
Sucrose Hydrolase Suxb From Xanthomonas Oryzae Pv. Oryzae In Complex With Glucose
Organism: Xanthomonas oryzae pv. oryzae
Method: X-RAY DIFFRACTION Resolution:2.95 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: K, GLC |
 |
Atomic Resolution (1.02 A) Xfel Structure Of Nitrite-Bound Copper Nitrite Reductase From Bradyrhizobium Sp. Determined By Serial Femtosecond Rotation Crystallography (Sf-Rox) At 100 K
Organism: Bradyrhizobium diazoefficiens usda 110
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: CU, NO2, GLC, FRU, SO4 |
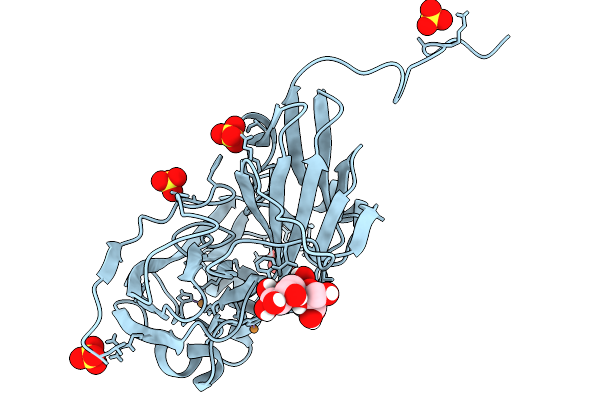 |
Atomic Resolution (1.00 A) Xfel Structure Of As-Isolated Copper Nitrite Reductase From Bradyrhizobium Sp. At High Ph (7.3) Determined By Serial Femtosecond Rotation Crystallography (Sf-Rox) At 100 K
Organism: Bradyrhizobium diazoefficiens usda 110
Method: X-RAY DIFFRACTION Resolution:1.00 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: CU, GLC, FRU, SO4 |
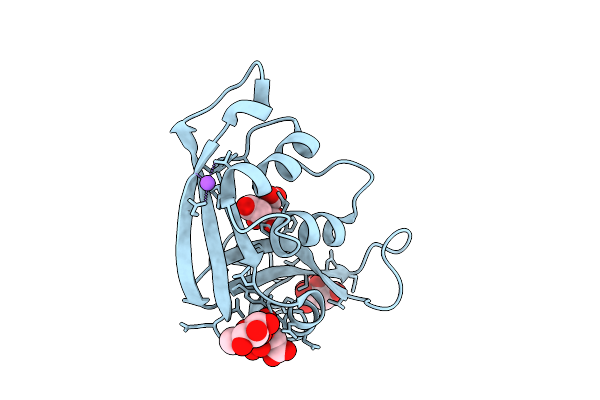 |
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.31 Å Release Date: 2026-03-18 Classification: PROTEIN BINDING Ligands: CIT, GLC, NA, CL |
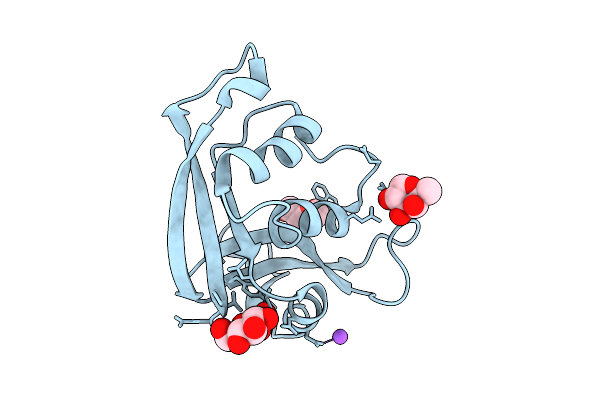 |
Structural Characterization Of Glucose- And Fucose-Binding Sites In Human Rnase2
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.01 Å Release Date: 2026-03-18 Classification: PROTEIN BINDING Ligands: PEG, FUC, ACT, GLC, NA |
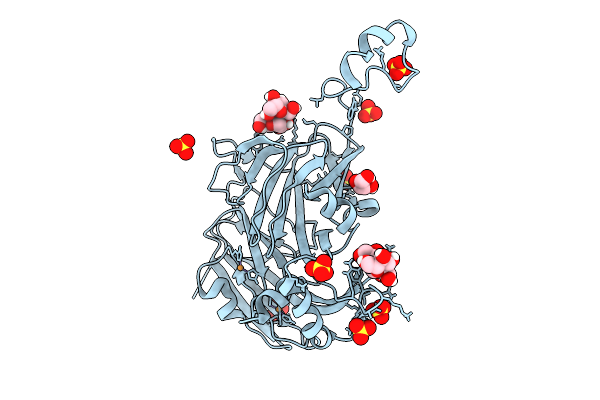 |
Atomic Resolution (1.15 A) Xfel Structure Of As-Isolated Copper Nitrite Reductase From Bradyrhizobium Sp. At Low Ph (5.5) Determined By Serial Femtosecond Rotation Crystallography (Sf-Rox) At 100 K
Organism: Bradyrhizobium sp. ors 375
Method: X-RAY DIFFRACTION Resolution:1.15 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: CU, GLC, FRU, SO4, GOL |
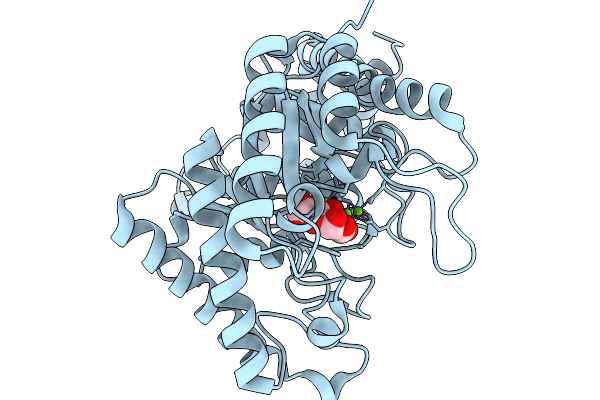 |
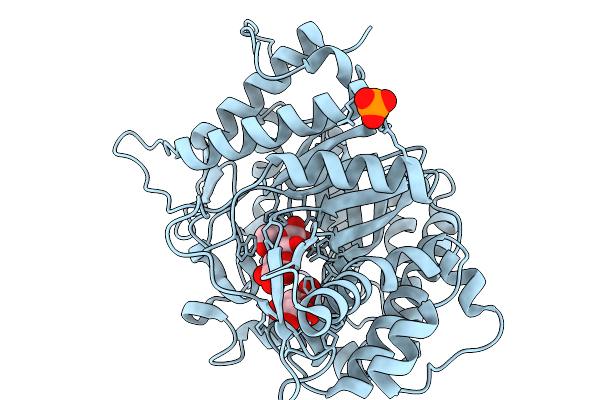
Xylose Isomerase Collected At 20C Using Time-Resolved Serial Synchrotron Crystallography With Glucose At 180 Seconds
Organism: Streptomyces rubiginosus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-02-18 Classification: ISOMERASE Ligands: GLC, GLO, MG |
 |
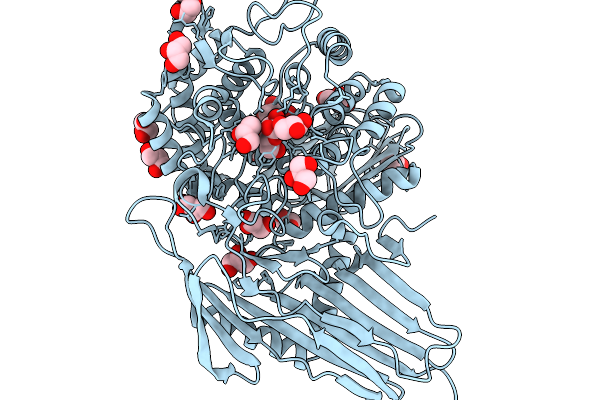
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: GLC, BGC, PO4 |
 |
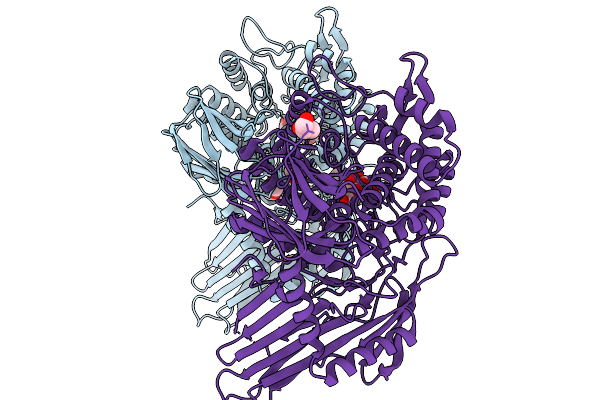
Crystal Structure Of The C-Terminally Truncated Human E574Q-Pgghg Mutant In Complex With Glucose
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.36 Å Release Date: 2026-02-11 Classification: SUGAR BINDING PROTEIN Ligands: CL, GOL, BGC, GLC, PEG |
 |
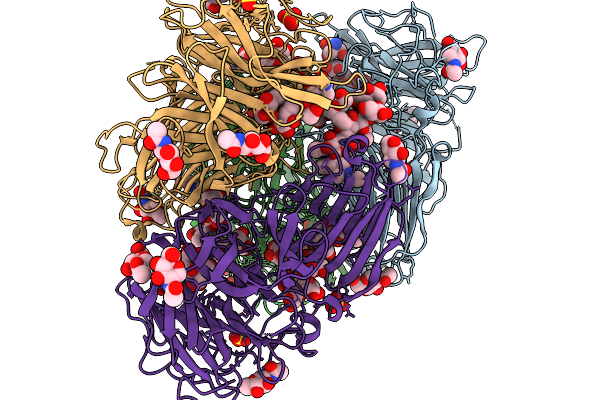
Crystal Structure Of C-Terminally Truncated Human Pgghg In Complex With Glucose.
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.61 Å Release Date: 2026-02-11 Classification: SUGAR BINDING PROTEIN Ligands: CL, GLC, BGC, GOL, PEG |
 |
Structure Of D81A-Fructofuranosidase From Purpureocilum Lilacinum In Complex With Raffinose
Organism: Komagataella phaffii cbs 7435
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: FRU, TRS, NAG, GLC, GLA, SO4 |
 |
Structure Of D81A-Fructofuranosidase From Purpureocilum Lilacinum In Complex With Sucrose
Organism: Komagataella phaffii cbs 7435
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: GLC, FRU, SO4, TRS, NAG, GOL |
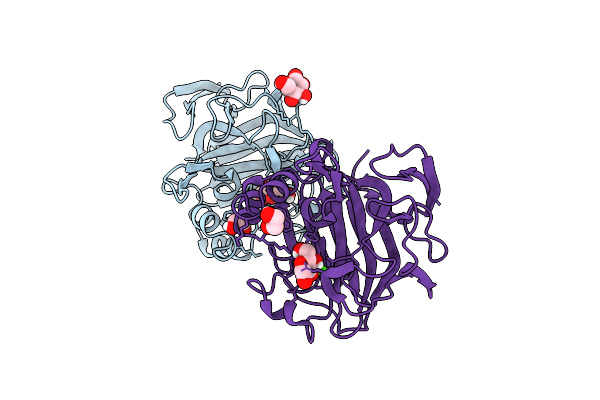 |
3-Keto-Glycoside Eliminase/Hydratase In Komplex With 2-Hydroxy-3-Keto-Glucal
Organism: Bacteroides thetaiotaomicron
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2025-10-29 Classification: LYASE Ligands: A1IBU, EDO, CA, GLC |
 |
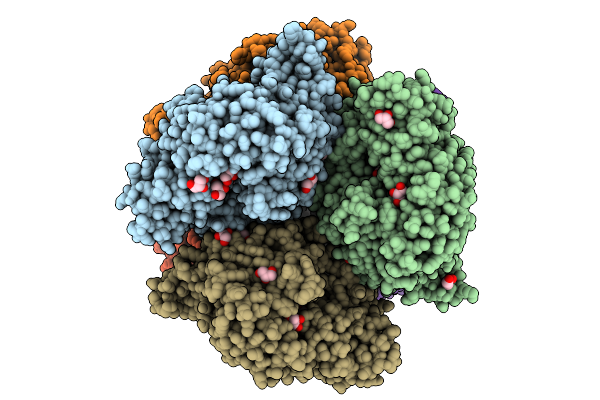
Structural Basis Of The Bifunctionality Of M. Salinexigens Zyf650T Glucosylglycerol Phosphorylase In Glucosylglycerol Catabolism
Organism: Marinobacter salinexigens
Method: X-RAY DIFFRACTION Resolution:2.72 Å Release Date: 2025-09-10 Classification: STRUCTURAL PROTEIN Ligands: GLC, GOL, NA, G1P |
 |
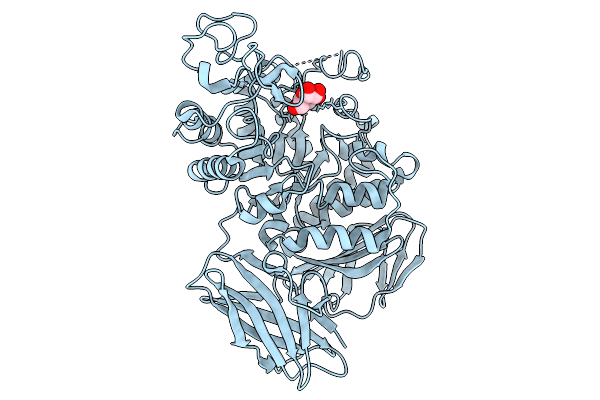
Structural Basis Of The Bifunctionality Of M. Salinexigens Zyf650T Glucosylglycerol Phosphorylase In Glucosylglycerol Catabolism
Organism: Marinobacter salinexigens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-09-10 Classification: STRUCTURAL PROTEIN Ligands: GOL, GLC, PEG |
 |
Structure Of A Truncated Loopa Mutant From The Human Gut Flora K. Grimontii Apg In Complex With Glucose
Organism: Klebsiella grimontii
Method: X-RAY DIFFRACTION Resolution:2.39 Å Release Date: 2025-09-03 Classification: HYDROLASE Ligands: GLC |

