Search Count: 23
All
Selected
 |
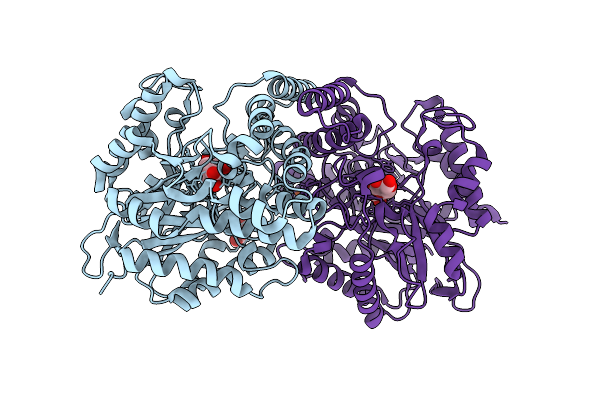
Cryo-Em Structure Of Collagenase H (E416Q Mutant) From Hathewaya Histolytica Bound To C-Terminal Region Of Collagen Model Peptide (Pro-Hyp-Gly)10
Organism: Hathewaya histolytica, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: METAL BINDING PROTEIN Ligands: CA, ZN |
 |
Organism: Hathewaya histolytica
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: METAL BINDING PROTEIN Ligands: ZN, CA |
 |
Cryo-Em Structure Of Collagenase H (E416Q Mutant) From Hathewaya Histolytica In Complex With Collagen Model Peptide (Pro-Hyp-Gly)10
Organism: Hathewaya histolytica, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: METAL BINDING PROTEIN Ligands: CA, ZN |
 |
Cryo-Em Structure Of Collagenase H (E416Q Mutant) From Hathewaya Histolytica In Complex With Collagen Model Peptide (Pro-Hyp-Gly)12
Organism: Hathewaya histolytica, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: METAL BINDING PROTEIN Ligands: CA, ZN |
 |
Cryo-Em Structure Of Collagenase H (E416Q Mutant) From Hathewaya Histolytica Bound To C-Terminal Region Of The Collagen-Binding Protein Colh (Pro-Pro-Gly)10
Organism: Hathewaya histolytica, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: METAL BINDING PROTEIN Ligands: CA, ZN |
 |
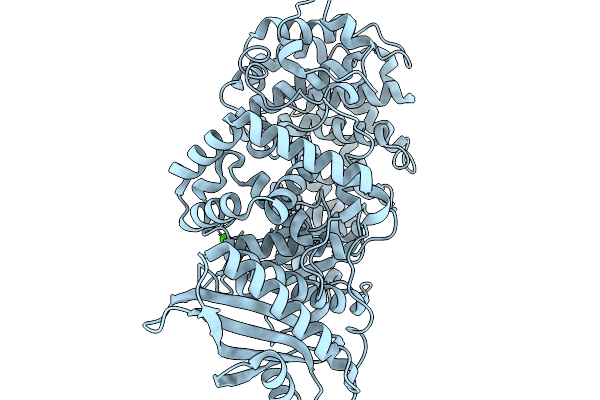
Crystal Structure Of Substrate Bound Gh5_22 Exo-Beta-Xylosidase From The Seaweed-Derived Thermophile Geobacillus Thermodenitrificans Os27
Organism: Geobacillus thermodenitrificans
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: GOL, FE |
 |
Crystal Structure Of Gh5_22 Exo-Beta-Xylosidase From The Seaweed-Derived Thermophile Geobacillus Thermodenitrificans Os27
Organism: Geobacillus thermodenitrificans
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: FE |
 |
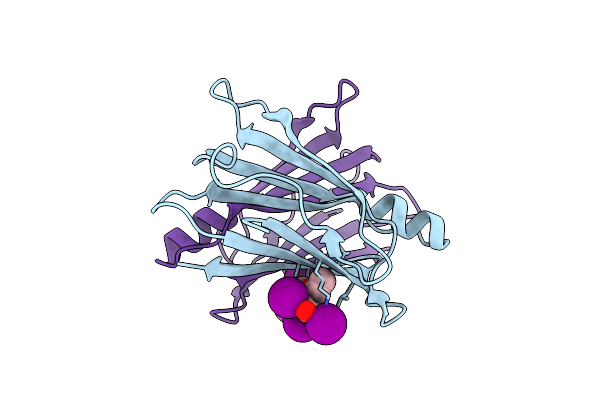
Room Temperature Crystal Structure Of Collagenase Colh From Hathewaya Histolytica At 2.7 Angstrom Resolution
Organism: Hathewaya histolytica
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-01-14 Classification: HYDROLASE Ligands: ZN, CA |
 |
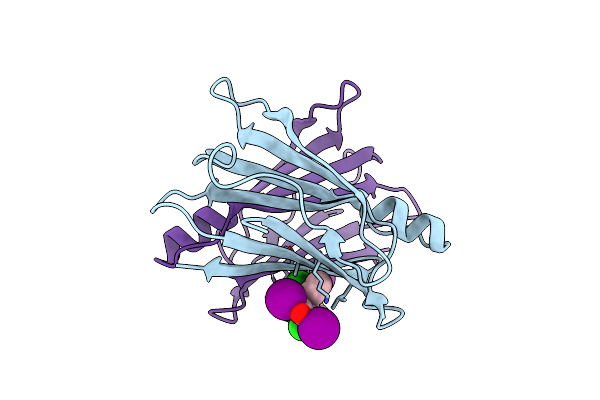
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2024-05-15 Classification: TRANSPORT PROTEIN Ligands: WGH |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2024-05-15 Classification: TRANSPORT PROTEIN Ligands: WGJ |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.51 Å Release Date: 2024-05-15 Classification: TRANSPORT PROTEIN Ligands: WH0 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2023-06-28 Classification: TRANSPORT PROTEIN Ligands: PJO |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2023-06-28 Classification: TRANSPORT PROTEIN Ligands: R75, CA |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2023-06-28 Classification: TRANSPORT PROTEIN Ligands: PKK |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2023-06-28 Classification: TRANSPORT PROTEIN Ligands: PKQ |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2019-03-27 Classification: TRANSCRIPTION Ligands: B0X |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2019-03-27 Classification: TRANSCRIPTION Ligands: B1O |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2018-10-17 Classification: TRANSCRIPTION Ligands: RTE, CL |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2018-10-17 Classification: TRANSCRIPTION Ligands: RTF, CL |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2010-08-04 Classification: TRANSCRIPTION Ligands: ACY, MPD |

