Search Count: 103
All
Selected
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.46 Å Release Date: 2026-02-18 Classification: DE NOVO PROTEIN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-02-18 Classification: DE NOVO PROTEIN |
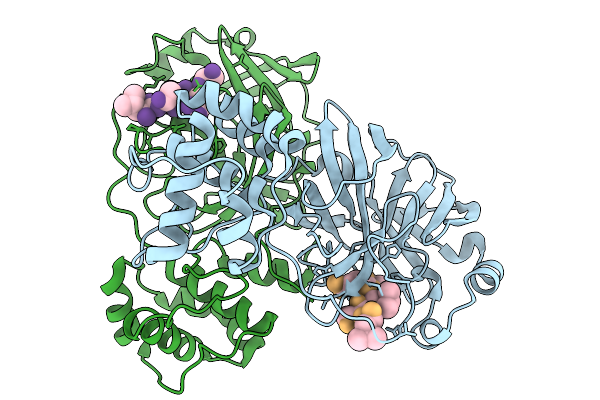 |
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.43 Å Release Date: 2026-02-04 Classification: VIRAL PROTEIN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.47 Å Release Date: 2026-01-07 Classification: TRANSFERASE/TRANSFERASE INHIBITOR Ligands: A1CMW, MTA |
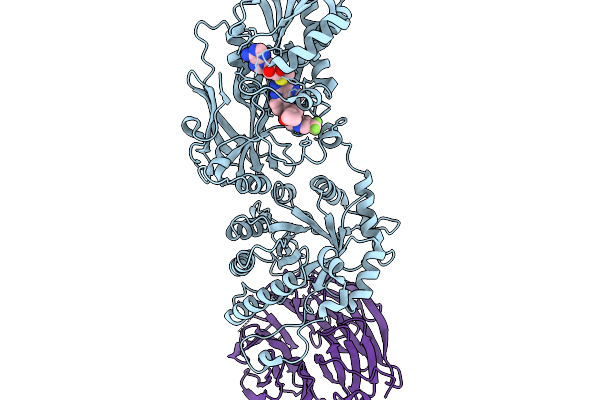 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.36 Å Release Date: 2026-01-07 Classification: TRANSFERASE/TRANSFERASE INHIBITOR Ligands: A1CMZ, MTA |
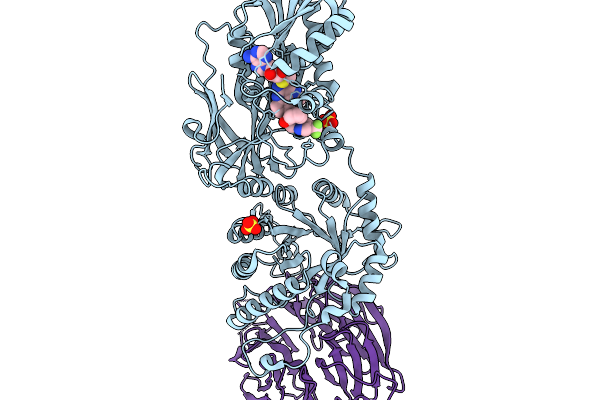 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.04 Å Release Date: 2026-01-07 Classification: TRANSFERASE/TRANSFERASE INHIBITOR Ligands: MTA, A1CM0, SO4 |
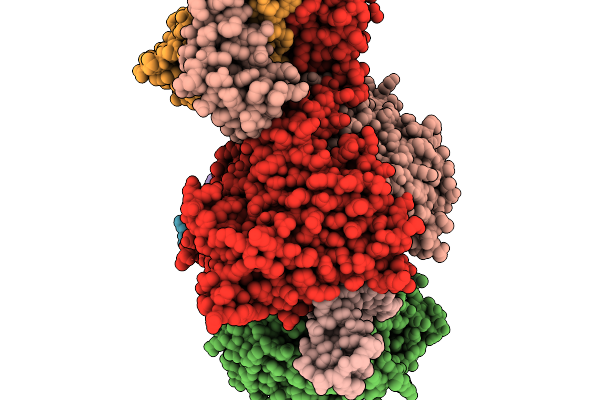 |
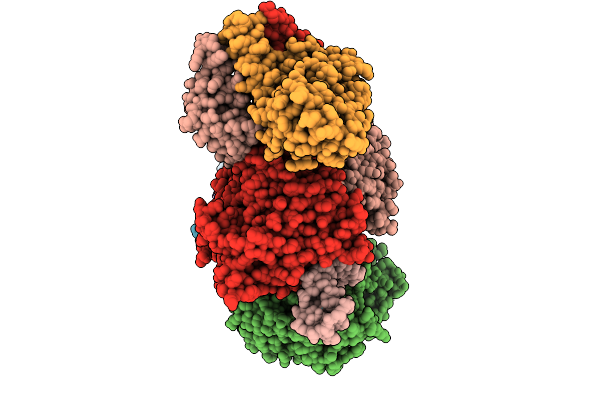
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-01-07 Classification: DNA BINDING PROTEIN/RNA/DNA Ligands: ATP, MG |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.50 Å Release Date: 2025-12-31 Classification: DNA BINDING PROTEIN/RNA/DNA Ligands: ATP, MG |
 |
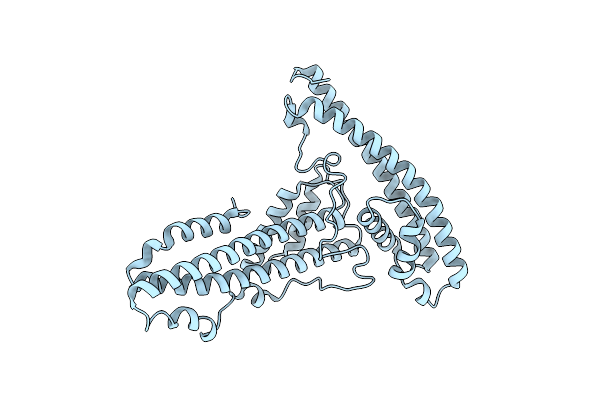
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:3.40 Å Release Date: 2025-12-03 Classification: DNA BINDING PROTEIN Ligands: ZN |
 |
Organism: Plasmodium falciparum 3d7
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-05-14 Classification: CELL INVASION |
 |
Organism: Kutzneria sp. 744
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2025-01-29 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
Organism: Kutzneria sp. 744
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-01-29 Classification: OXIDOREDUCTASE Ligands: HEM, ONH |
 |
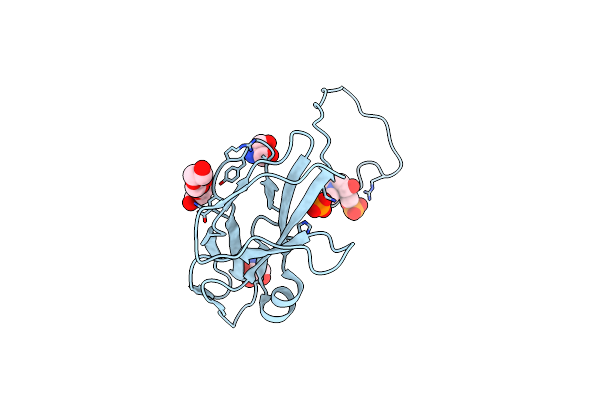
Organism: Kutzneria sp. 744
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2025-01-29 Classification: OXIDOREDUCTASE Ligands: HEM, A1ECN |
 |
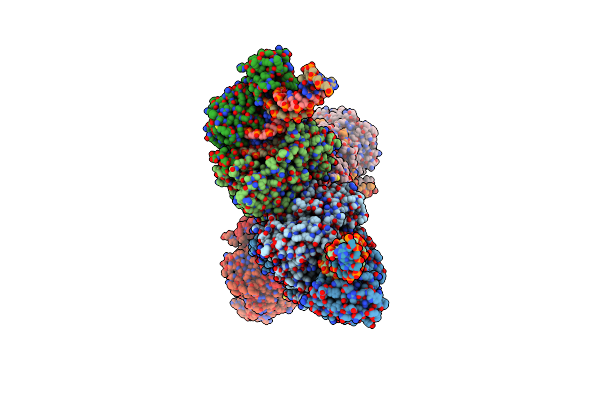
Organism: Nocardia seriolae
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2024-11-13 Classification: HYDROLASE Ligands: UMP, PO4, TRS, PEG |
 |
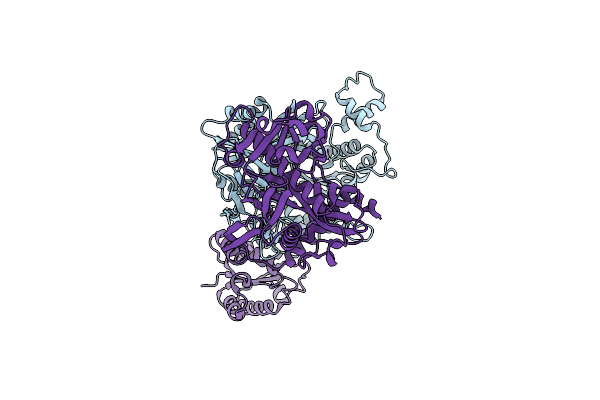
Organism: Thermoflavifilum thermophilum, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2023-11-08 Classification: DNA BINDING PROTEIN/DNA/RNA |
 |
Organism: Thermoflavifilum thermophilum
Method: ELECTRON MICROSCOPY Release Date: 2023-10-18 Classification: DNA BINDING PROTEIN |
 |
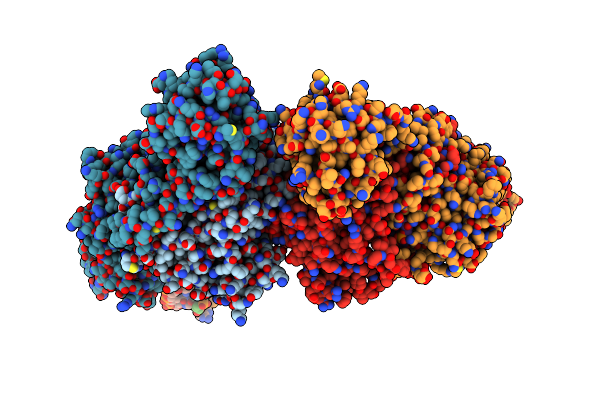
Organism: Thermoflavifilum thermophilum, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2023-10-18 Classification: DNA BINDING PROTEIN/DNA/RNA |
 |
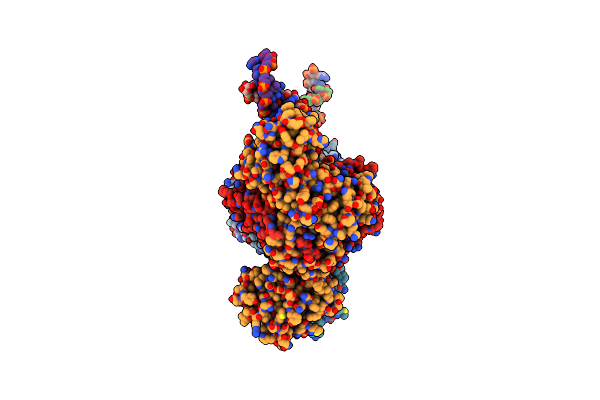
Organism: Thermoflavifilum thermophilum, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2023-10-18 Classification: DNA BINDING PROTEIN/DNA/RNA |
 |
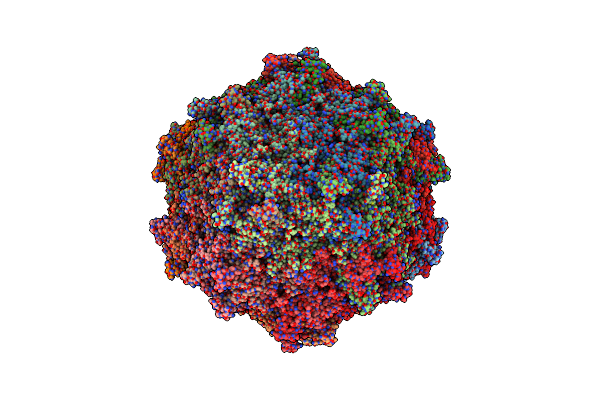
Organism: Thermoflavifilum thermophilum, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2023-10-18 Classification: DNA/RNA/CELL INVASION |
 |
Organism: Canine minute virus
Method: ELECTRON MICROSCOPY Release Date: 2023-08-30 Classification: VIRUS |

