Search Count: 158
All
Selected
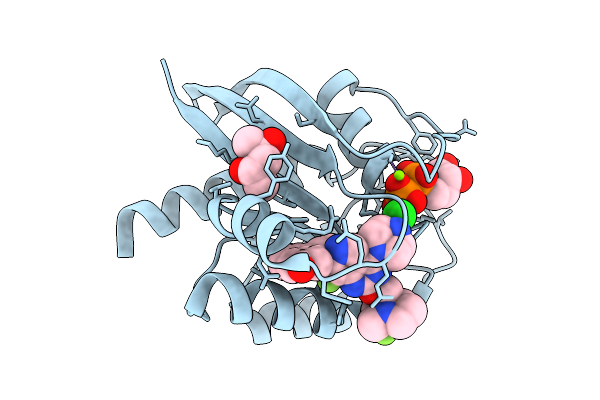 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-01-21 Classification: ONCOPROTEIN Ligands: GDP, MG, 6IC, MPD, CL |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-01-21 Classification: ONCOPROTEIN Ligands: GNP, MG, 6IC, GOL, SO4, CL, NA |
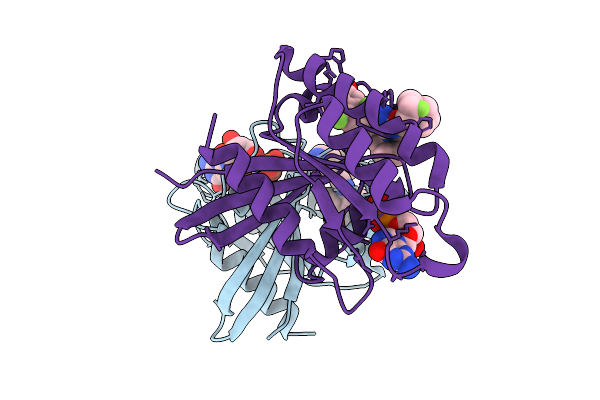 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-01-21 Classification: ONCOPROTEIN Ligands: GDP, MG, 6IC, CL |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-01-21 Classification: ONCOPROTEIN Ligands: GNP, MG, 6IC, SO4, GOL, CL |
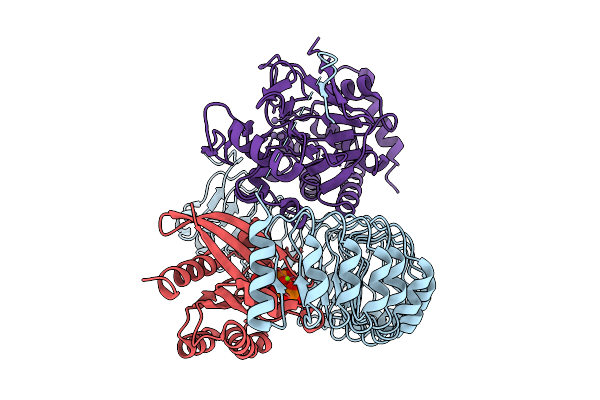 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2026-01-21 Classification: SIGNALING PROTEIN Ligands: GNP, MG, MN |
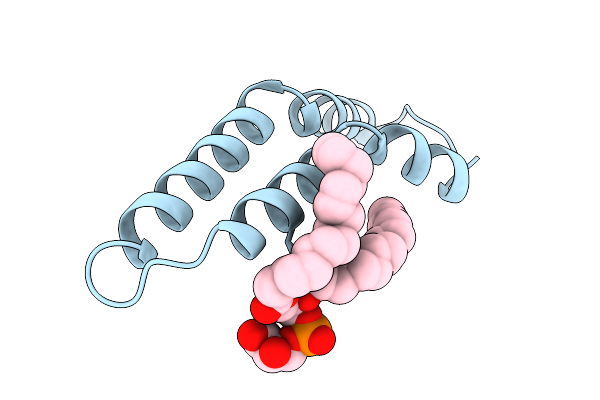 |
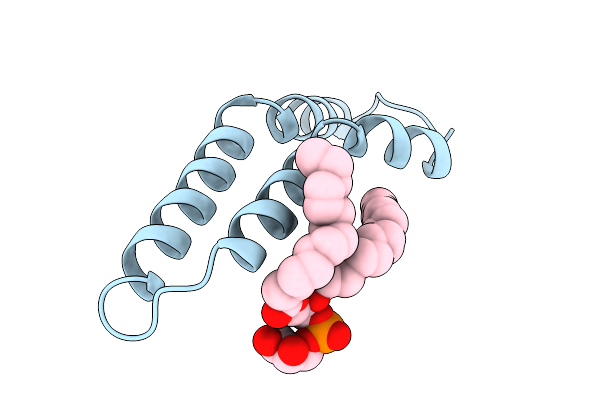
Organism: salmonella enterica subsp. enterica serovar typhi
Method: ELECTRON MICROSCOPY Resolution:3.15 Å Release Date: 2025-12-24 Classification: PROTEIN FIBRIL Ligands: LHG |
 |
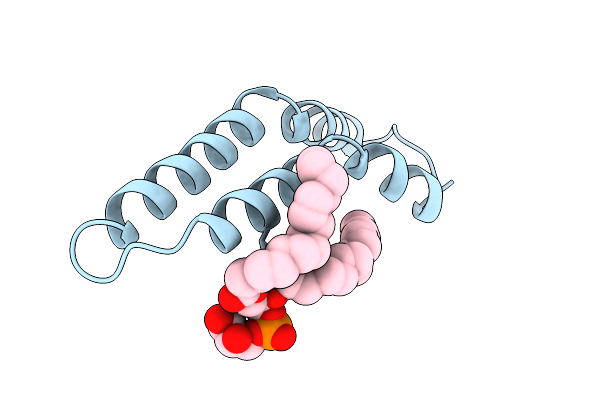
Organism: Salmonella enterica subsp. enterica serovar typhi
Method: ELECTRON MICROSCOPY Resolution:2.88 Å Release Date: 2025-12-24 Classification: PROTEIN FIBRIL Ligands: LHG |
 |
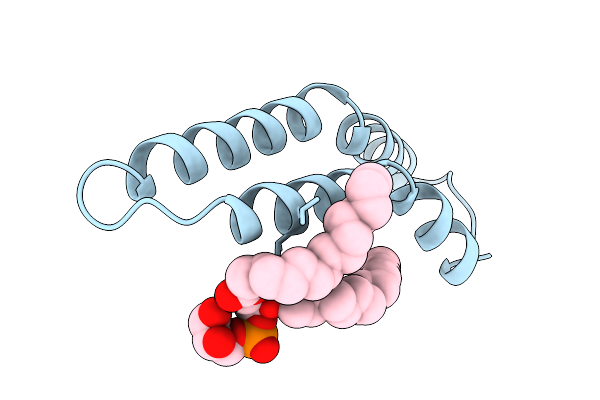
Organism: Salmonella enterica subsp. enterica serovar typhi
Method: ELECTRON MICROSCOPY Resolution:2.89 Å Release Date: 2025-12-24 Classification: PROTEIN FIBRIL Ligands: LHG |
 |
Organism: Salmonella enterica subsp. enterica serovar typhi
Method: ELECTRON MICROSCOPY Resolution:3.11 Å Release Date: 2025-12-24 Classification: PROTEIN FIBRIL Ligands: LHG |
 |
Crystal Structure Of Wild-Type Egfr In Complex With The Reversible Inhibitor Sevabertinib (Bay 2927088)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.14 Å Release Date: 2025-10-15 Classification: TRANSFERASE Ligands: SIN, CL, DMS, EDO, A1JBI |
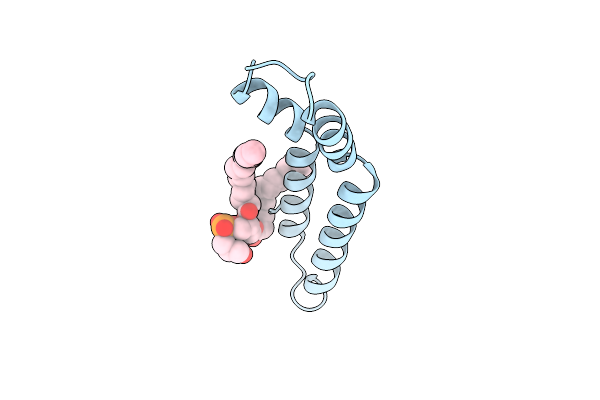 |
Organism: Salmonella
Method: ELECTRON MICROSCOPY Resolution:2.24 Å Release Date: 2025-03-26 Classification: PROTEIN FIBRIL Ligands: LHG |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-01-15 Classification: PROTEIN FIBRIL Ligands: X41 |
 |
Cryo-Em Structure Of The Decameric Trat Surface Exclusion Lipoprotein From Escherichia Coli (F Plasmid)
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2024-12-11 Classification: MEMBRANE PROTEIN |
 |
Cryo-Em Structure Of The Decameric Trat Surface Exclusion Lipoprotein From Klebsiella Pneumoniae (Pkpqil Plasmid)
Organism: Klebsiella pneumoniae
Method: ELECTRON MICROSCOPY Release Date: 2024-12-11 Classification: MEMBRANE PROTEIN Ligands: DGA |
 |
Crystal Structure Of Chlorite Dismutase At 3000 Ev Based On Analytical Absorption Corrections
Organism: Cyanothece sp. pcc 7425
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2024-07-03 Classification: OXIDOREDUCTASE Ligands: HEM, SO4, GOL, CL |
 |
Crystal Structure Of Ompk36 Gd At 3500 Ev Based On Spherical Harmonics Absorption Corrections
Organism: Klebsiella pneumoniae
Method: X-RAY DIFFRACTION Resolution:2.34 Å Release Date: 2024-06-19 Classification: MEMBRANE PROTEIN Ligands: SO4 |
 |
Organism: Klebsiella pneumoniae
Method: X-RAY DIFFRACTION Resolution:2.34 Å Release Date: 2024-06-19 Classification: MEMBRANE PROTEIN Ligands: SO4 |
 |
Crystal Structure Of Chlorite Dismutase At 3000 Ev Based On Spherical Harmonics Absorption Corrections
Organism: Cyanothece sp. pcc 7425
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2024-06-19 Classification: OXIDOREDUCTASE Ligands: HEM, SO4, GOL, CL |
 |
Crystal Structure Of Chlorite Dismutase At 3000 Ev With No Absorption Corrections
Organism: Cyanothece sp. pcc 7425
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2024-06-19 Classification: OXIDOREDUCTASE Ligands: HEM, SO4, GOL, CL |
 |
Crystal Structure Of Chlorite Dismutase At 3000 Ev Based On A Combination Of Spherical Harmonics And Analytical Absorption Corrections
Organism: Cyanothece sp. pcc 7425
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2024-06-19 Classification: OXIDOREDUCTASE Ligands: HEM, SO4, GOL, CL |

