Search Count: 58
All
Selected
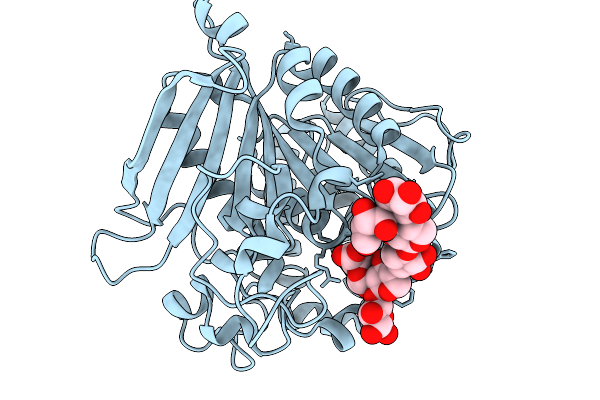 |
The Glucuronyl Esterase Otce15A From Opitutus Terrae In Complex With A Heptasaccharide
Organism: Opitutus terrae pb90-1
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-01-14 Classification: HYDROLASE |
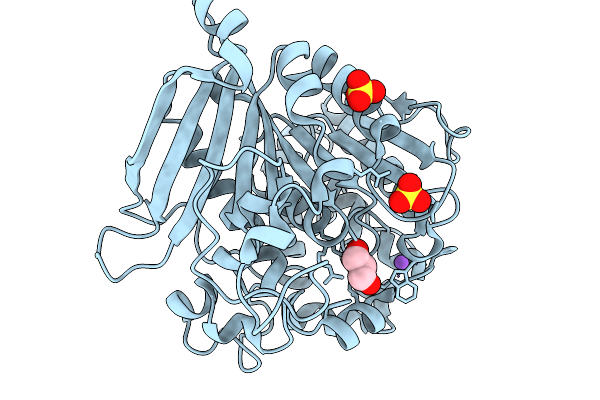 |
Organism: Opitutus terrae pb90-1
Method: X-RAY DIFFRACTION Resolution:1.39 Å Release Date: 2026-01-14 Classification: HYDROLASE Ligands: GOL, SO4, NA |
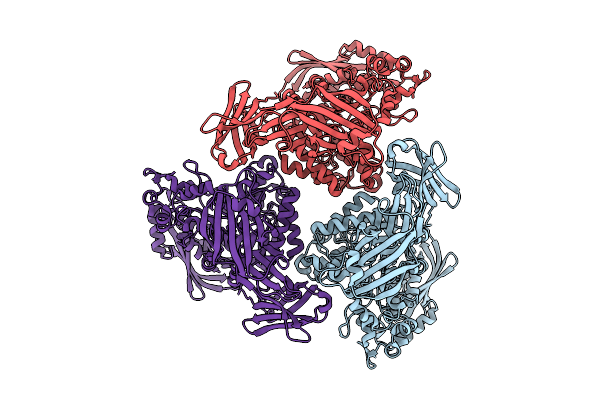 |
Active Conformation Of A Redox-Regulated Glycoside Hydrolase (Capgh2B) From The Gh2 Family
Organism: Metagenome
Method: ELECTRON MICROSCOPY Resolution:2.62 Å Release Date: 2025-11-12 Classification: HYDROLASE |
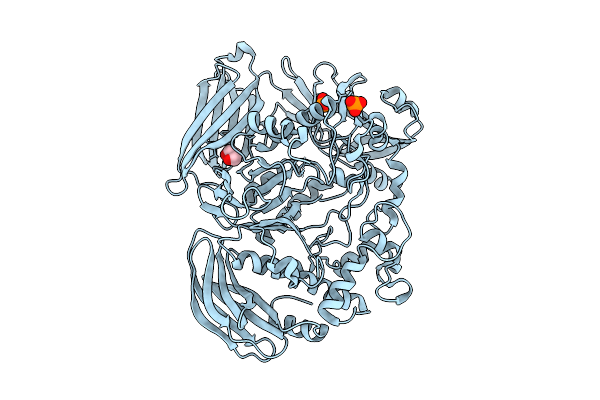 |
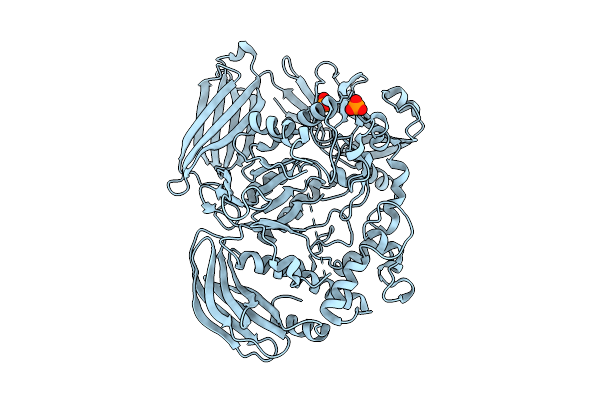
Crystal Structure Of The Inactive Conformation Of A Glycoside Hydrolase (Capgh2B) From The Gh2 Family In The Space Group I213 At 2.05 A
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: PO4, GOL |
 |
Crystal Structure Of The Inactive Conformation Of A Glycoside Hydrolase (Capgh2B - E553Q Mutant) From The Gh2 Family In The Space Group I213 At 2.6 A
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: PO4 |
 |
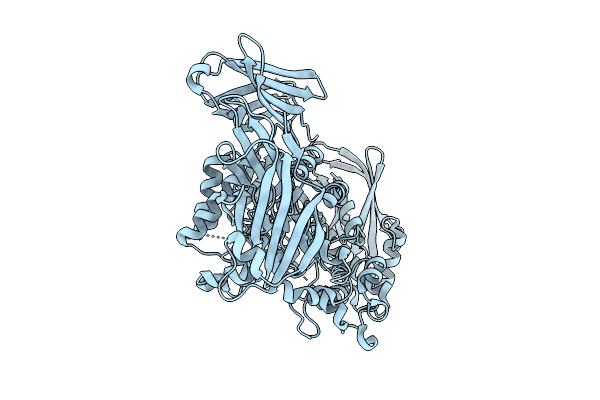
Crystal Structure Of The Inactive Conformation Of A Glycoside Hydrolase (Capgh2B) From The Gh2 Family In The Space Group I213 At 2.75 A
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: PO4, TAU |
 |
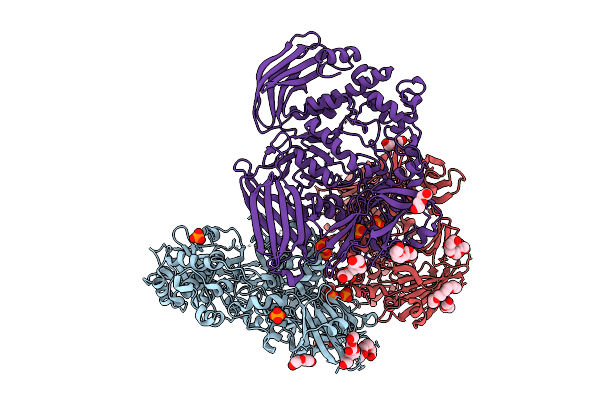
Crystal Structure Of The Inactive Conformation Of A Glycoside Hydrolase (Capgh2B) From The Gh2 Family In The Space Group R3 At 2.45 A
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2025-11-12 Classification: HYDROLASE |
 |
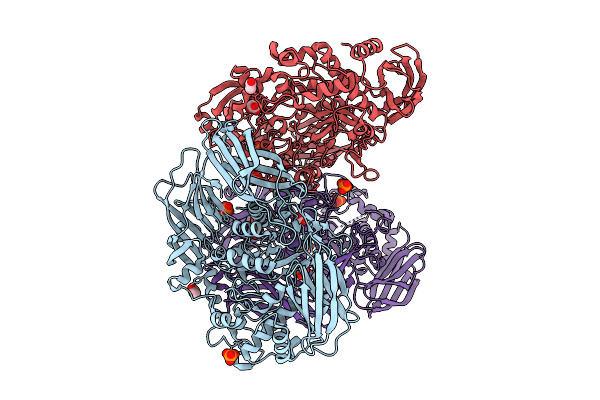
Crystal Structure Of The Inactive Conformation Of A Glycoside Hydrolase (Capgh2B) From The Gh2 Family In The Space Group I212121 At 2.65 A
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: PO4, PEG, PG4 |
 |
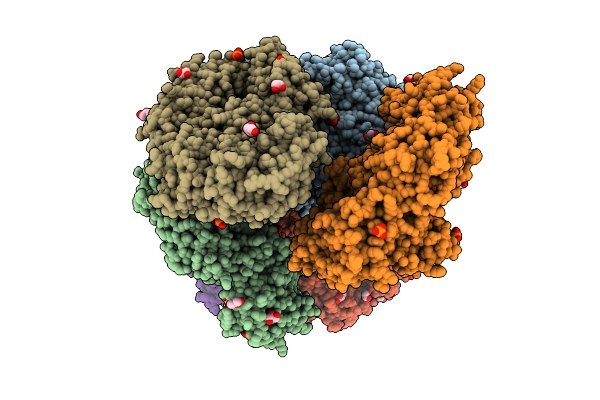
Crystal Structure Of The Inactive Conformation Of A Glycoside Hydrolase (Capgh2B - E553Q Mutant) From The Gh2 Family In The Space Group P3121 At 3.05 A
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:3.05 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: PO4, ACT, MLI |
 |
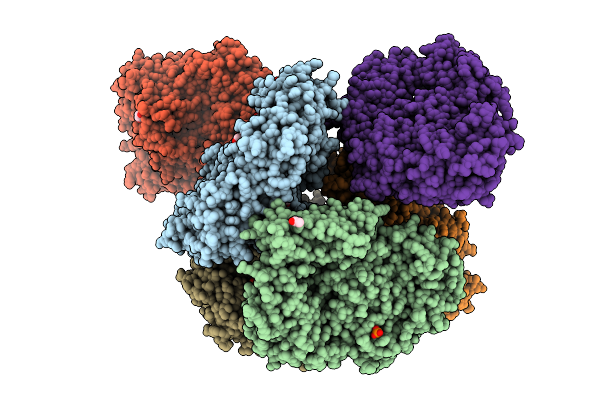
Crystal Structure Of The Inactive Conformation Of A Glycoside Hydrolase (Capgh2B) From The Gh2 Family In The Space Group P1 At 2.40 A
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: PO4, GOL |
 |
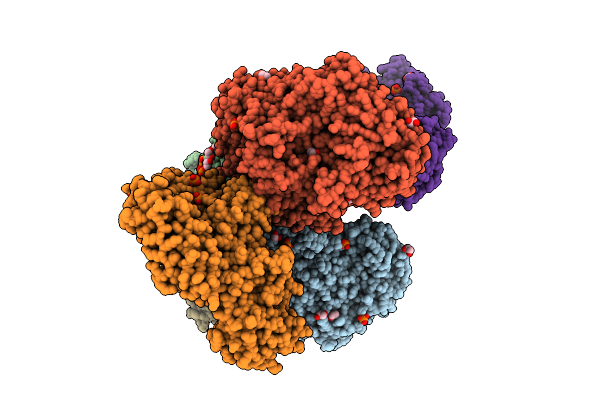
Crystal Structure Of The Inactive Conformation Of A Glycoside Hydrolase (Capgh2B) From The Gh2 Family In The Space Group P1 At 2.15 A
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: PO4, GOL, ACT |
 |
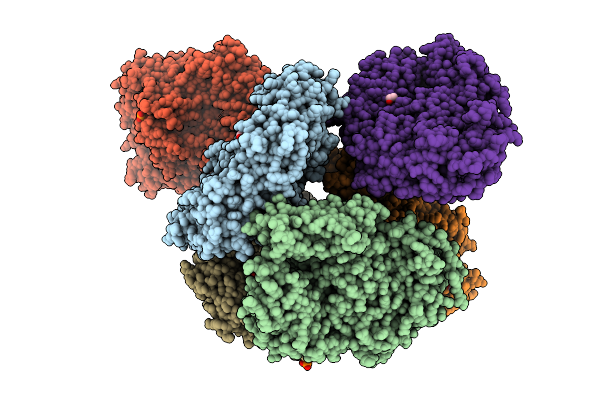
Crystal Structure Of The Inactive Conformation Of A Glycoside Hydrolase (Capgh2B - E553Q Mutant) From The Gh2 Family In The Space Group P1 At 2.25 A
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: PO4, EDO |
 |
Crystal Structure Of The Inactive Conformation Of A Glycoside Hydrolase (Capgh2B - E465A Mutant) From The Gh2 Family In The Space Group P1 At 3.1 A
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:3.10 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: PO4, EDO |
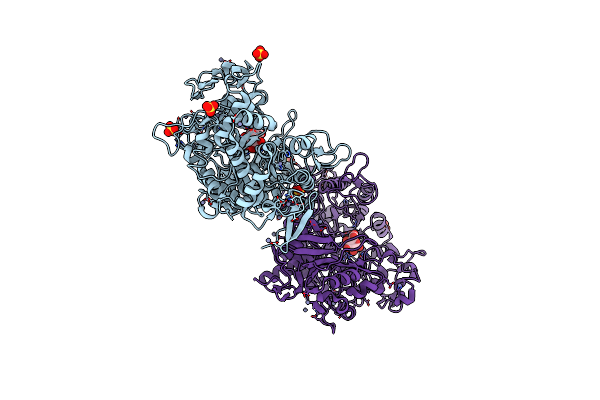 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.37 Å Release Date: 2022-11-16 Classification: HYDROLASE Ligands: ZN, SO4, C3G, BDP |
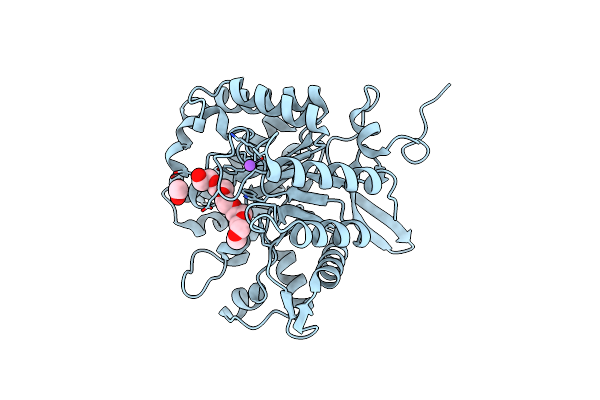 |
Crystal Structure Of A Novel Gh5 Enzyme Retrieved From Capybara Gut Metagenome
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2022-11-02 Classification: HYDROLASE Ligands: 1PG, ACT, NA, CL |
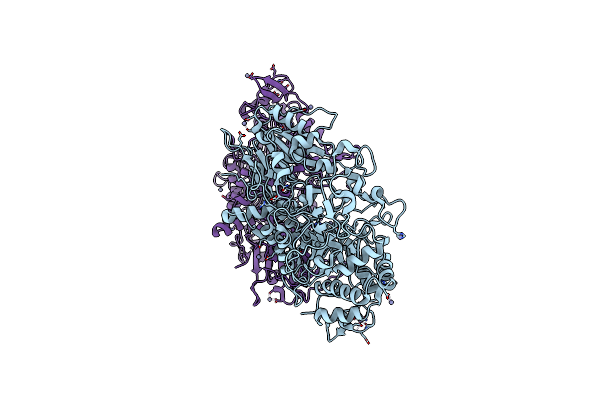 |
Organism: Unidentified
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2022-10-12 Classification: HYDROLASE Ligands: ZN |
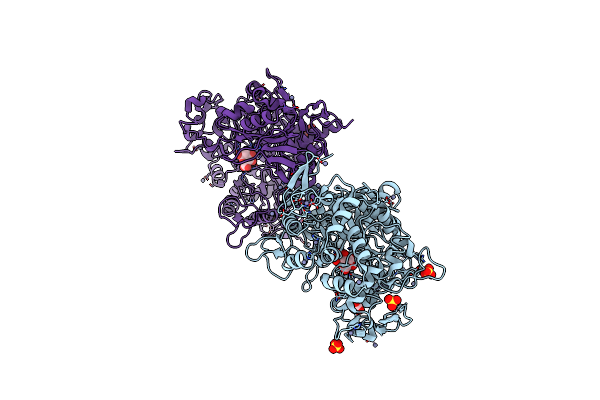 |
Organism: Unidentified
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2022-10-12 Classification: HYDROLASE Ligands: BDP, ACT, ZN, SO4 |
 |
Crystal Structure Of Fluorescein-Di-Beta-D-Glucuronide Bound To A Mutant Of Sn243 (D415A)
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2022-10-12 Classification: HYDROLASE Ligands: B9I, ZN, ACT, SO4 |
 |
Crystal Structure Of Para-Nitrophenyl-Beta-D-Glucuronide Bound To A Mutant Of Sn243 (D415A)
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.41 Å Release Date: 2022-10-12 Classification: HYDROLASE Ligands: ACT, C3G, ZN, SO4 |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2022-10-12 Classification: HYDROLASE Ligands: ZN, SO4 |

