Search Count: 17
All
Selected
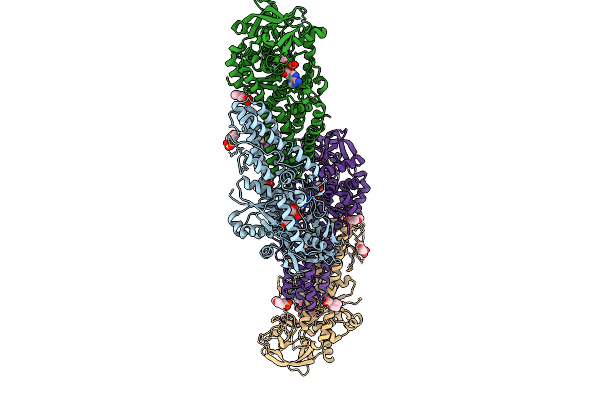 |
Organism: Streptomyces candidus
Method: X-RAY DIFFRACTION Resolution:2.23 Å Release Date: 2025-12-31 Classification: LIGASE Ligands: MES, ZN, GSU |
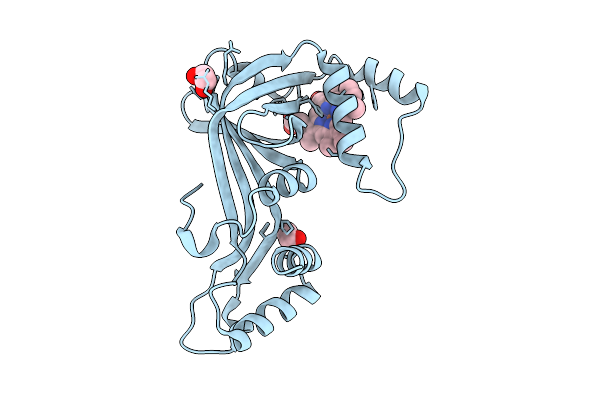 |
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, EDO |
 |
N-Hydroxylamine Dehydratase (Nohd) T98A/K167A Mutant Crystal Structure With Heme And N-Hydroxylated Ornithine (2.5H Soak)
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: EDO, HEM, ONH |
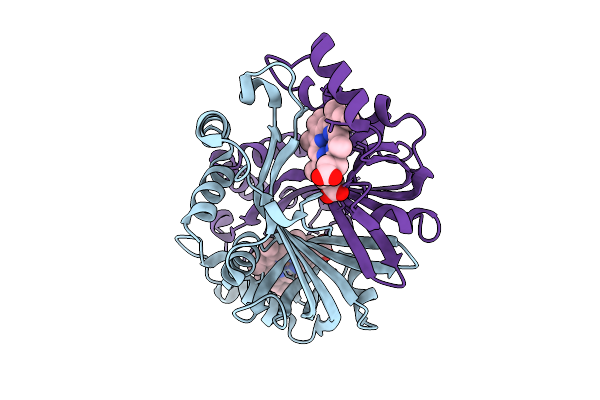 |
N-Hydroxylamine Dehydratase (Nohd) T98A/K167A Mutant Crystal Structure With Heme And N-Hydroxylated Ornithine (5H Soak)
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, ONH |
 |
N-Hydroxylamine Dehydratase (Nohd) R144Y/V98A (A1/A2) Mutant Crystal Structure With Heme
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, GOL, SO4 |
 |
N-Hydroxylamine Dehydratase (Nohd) H2F/F4P/P5S/R6Y/A59N/R144A (P1/P2/A1) Mutant Crystal Structure With Heme
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, EDO |
 |
N-Hydroxylamine Dehydratase (Nohd) H2F/F4P/P5S/R6Y/R144Y/V96A (P1/A1/A2) Mutant Crystal Structure With Heme
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, GOL, SO4 |
 |
N-Hydroxylamine Dehydratase (Nohd) A59N/R144Y/V96A (P2/A1/A2) Mutant Crystal Structure With Heme
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:2.13 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM |
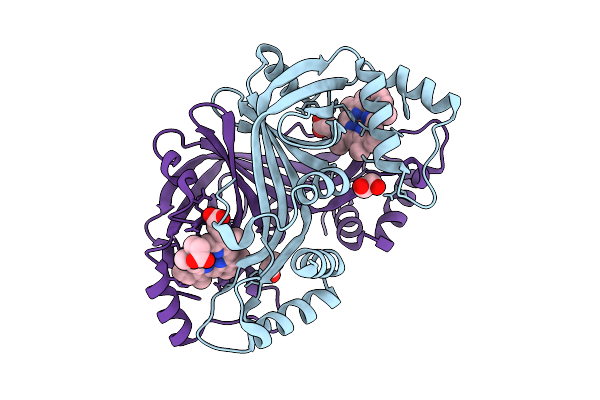 |
N-Hydroxylamine Dehydratase (Nohd) H2F/F4P/P5S/R6Y/A59N/R144Y/V96A (P1/P2/A1/A2) Mutant Crystal Structure With Heme
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, EDO |
 |
N-Hydroxylamine Dehydratase (Nohd) H2F/F4P/P5S/R6Y/A59N/R144Y/V96A/D172E/L172Q/D173P/R176V (P1/P2/A1/A2/A3) Mutant Crystal Structure With Heme And N-Hydroxylated Ornithine
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, BTB, ONH |
 |
Organism: Streptomyces griseus subsp. griseus
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2025-01-22 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, GOL |
 |
Organism: Streptomyces griseus subsp. griseus
Method: X-RAY DIFFRACTION Resolution:1.92 Å Release Date: 2025-01-22 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, ONH, GOL |
 |
Organism: Pseudoalteromonas luteoviolacea strain hi1
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2019-08-07 Classification: BIOSYNTHETIC PROTEIN Ligands: NQ7, PGE |
 |
O2-, Plp-Dependent L-Arginine Hydroxylase Rohp 4-Hydroxy-2-Ketoarginine Complex
Organism: Streptomyces cattleya (strain atcc 35852 / dsm 46488 / jcm 4925 / nbrc 14057 / nrrl 8057)
Method: X-RAY DIFFRACTION Resolution:1.53 Å Release Date: 2018-03-07 Classification: BIOSYNTHETIC PROTEIN Ligands: EDO, PEG, EGV |
 |
Organism: Streptomyces cattleya (strain atcc 35852 / dsm 46488 / jcm 4925 / nbrc 14057 / nrrl 8057)
Method: X-RAY DIFFRACTION Resolution:1.51 Å Release Date: 2018-03-07 Classification: BIOSYNTHETIC PROTEIN Ligands: EDO, PGE |
 |
Organism: Streptomyces cattleya
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2018-03-07 Classification: BIOSYNTHETIC PROTEIN Ligands: EDO, PEG, EJ1 |
 |
Organism: Streptomyces cattleya (strain atcc 35852 / dsm 46488 / jcm 4925 / nbrc 14057 / nrrl 8057)
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2018-03-07 Classification: BIOSYNTHETIC PROTEIN Ligands: EDO, EMS, PEG |

