Search Count: 304
All
Selected
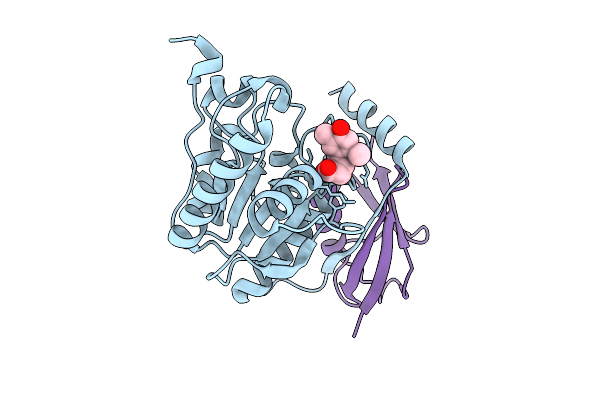 |
Crystal Structure Of Human Thymine Dna Glycosylase Tdg In Complex With A Covalent Inhibitor (1S, 5R)-C-2711
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.14 Å Release Date: 2026-03-11 Classification: DNA BINDING PROTEIN Ligands: A1LYG |
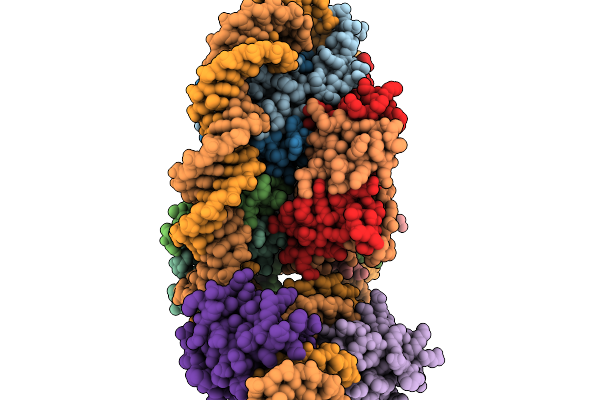 |
Structure Of Two Human Elf2 Transcription Factors In Complex With A Nucleosome
Organism: Xenopus laevis, Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN |
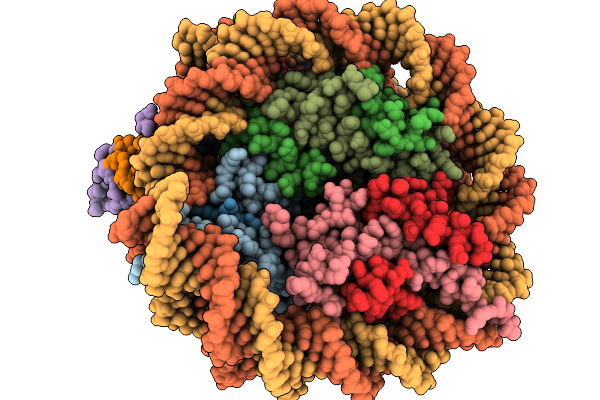 |
The 1:1 Cryo-Em Structure Of Bap1/Asxl1-K351Ub In Complex With H2Ak119Ub Nucleosome
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: NUCLEAR PROTEIN/DNA |
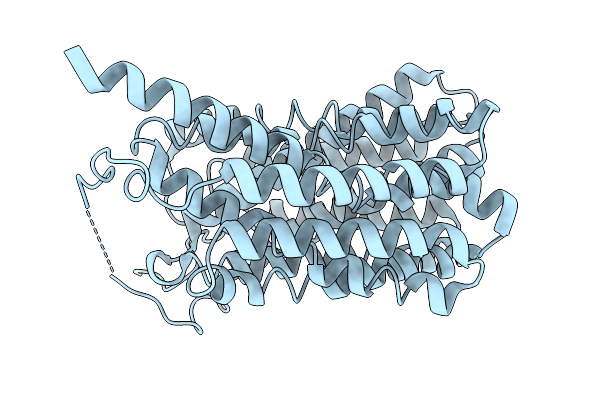 |
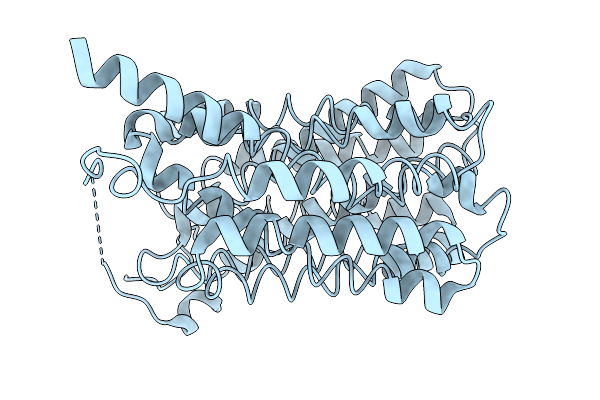
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.07 Å Release Date: 2026-01-07 Classification: TRANSPORT PROTEIN |
 |
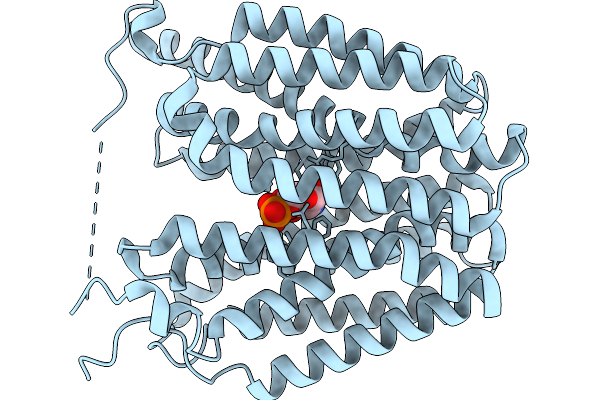
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.16 Å Release Date: 2026-01-07 Classification: TRANSPORT PROTEIN |
 |
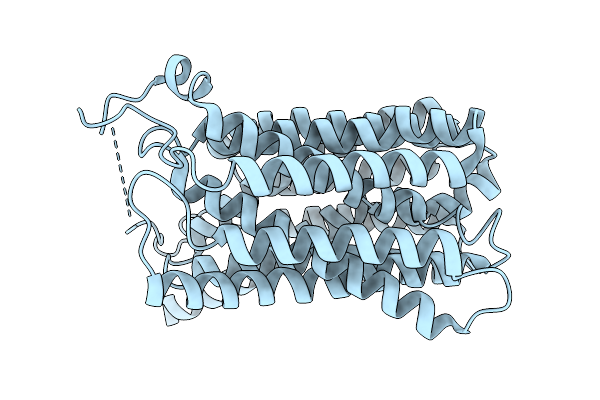
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2026-01-07 Classification: TRANSPORT PROTEIN Ligands: GLP |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.47 Å Release Date: 2026-01-07 Classification: TRANSPORT PROTEIN |
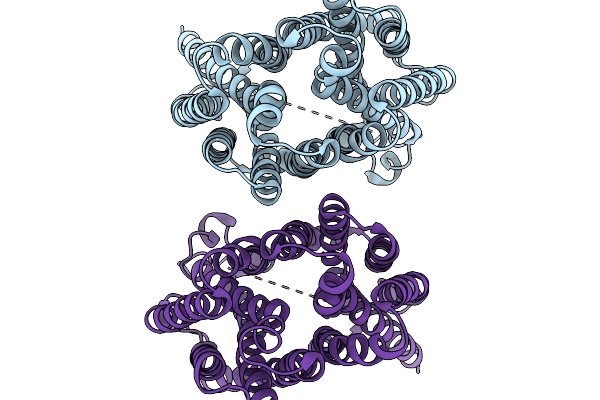 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.12 Å Release Date: 2026-01-07 Classification: TRANSPORT PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.89 Å Release Date: 2026-01-07 Classification: TRANSPORT PROTEIN Ligands: PO4 |
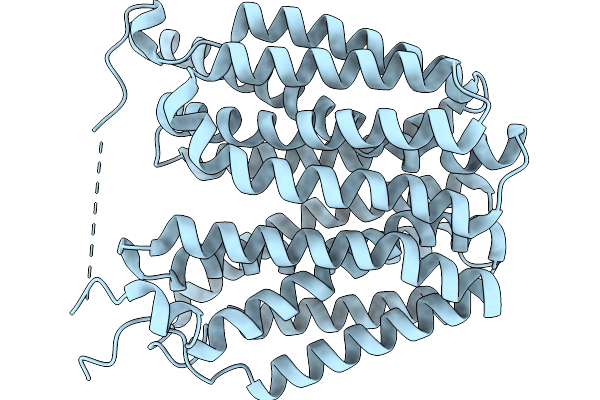 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-07 Classification: TRANSPORT PROTEIN |
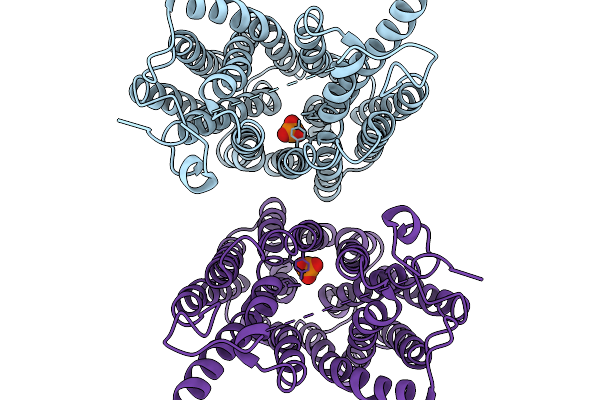 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.89 Å Release Date: 2026-01-07 Classification: TRANSPORT PROTEIN Ligands: PO4 |
 |
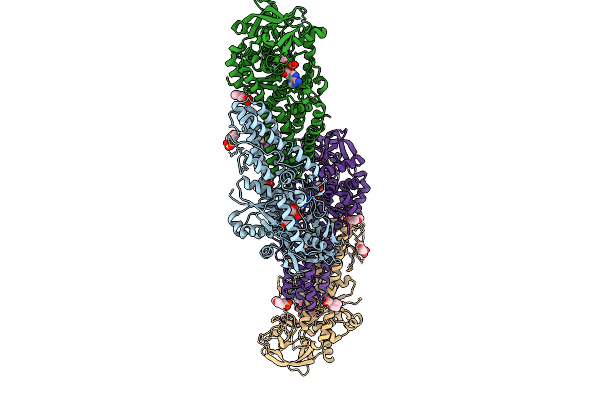
Cryo-Em Structure Of Rnf20/Rnf40-Rad6A-Ub In Complex With H2Bs112Glcnac Nucleosome
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.48 Å Release Date: 2025-12-31 Classification: NUCLEAR PROTEIN Ligands: ZN, NAG |
 |
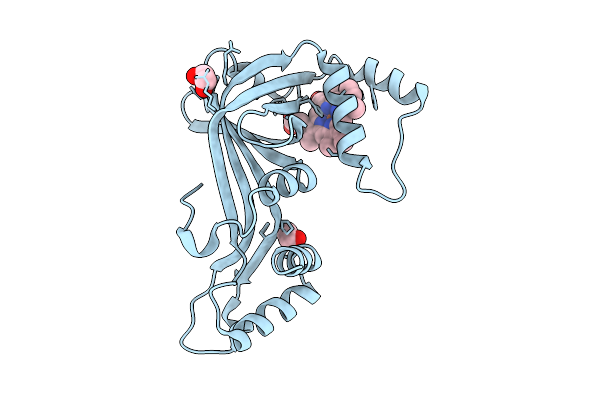
Organism: Streptomyces candidus
Method: X-RAY DIFFRACTION Resolution:2.23 Å Release Date: 2025-12-31 Classification: LIGASE Ligands: MES, ZN, GSU |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.59 Å Release Date: 2025-12-24 Classification: LIGASE |
 |
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, EDO |
 |
N-Hydroxylamine Dehydratase (Nohd) T98A/K167A Mutant Crystal Structure With Heme And N-Hydroxylated Ornithine (2.5H Soak)
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: EDO, HEM, ONH |
 |
N-Hydroxylamine Dehydratase (Nohd) T98A/K167A Mutant Crystal Structure With Heme And N-Hydroxylated Ornithine (5H Soak)
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, ONH |
 |
N-Hydroxylamine Dehydratase (Nohd) R144Y/V98A (A1/A2) Mutant Crystal Structure With Heme
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, GOL, SO4 |
 |
N-Hydroxylamine Dehydratase (Nohd) H2F/F4P/P5S/R6Y/A59N/R144A (P1/P2/A1) Mutant Crystal Structure With Heme
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, EDO |
 |
N-Hydroxylamine Dehydratase (Nohd) H2F/F4P/P5S/R6Y/R144Y/V96A (P1/A1/A2) Mutant Crystal Structure With Heme
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, GOL, SO4 |

