Search Count: 100
All
Selected
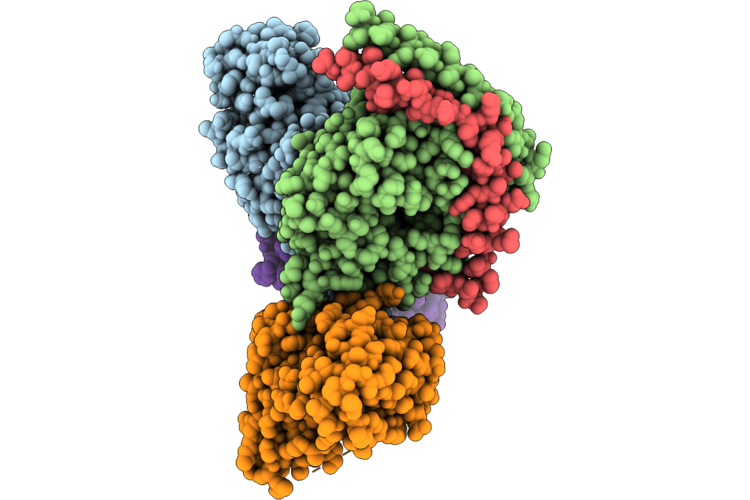 |
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-04-22 Classification: SIGNALING PROTEIN Ligands: GOQ |
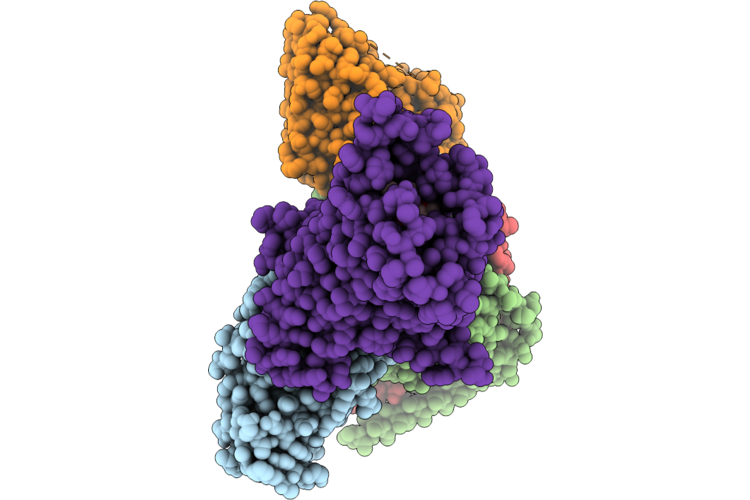 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2026-04-22 Classification: SIGNALING PROTEIN Ligands: GOQ |
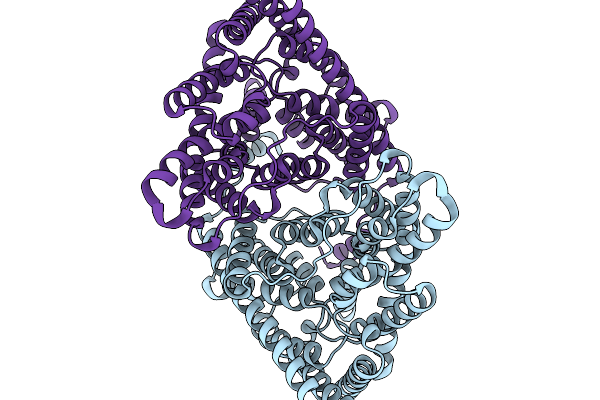 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-01-07 Classification: TRANSPORT PROTEIN Ligands: CL |
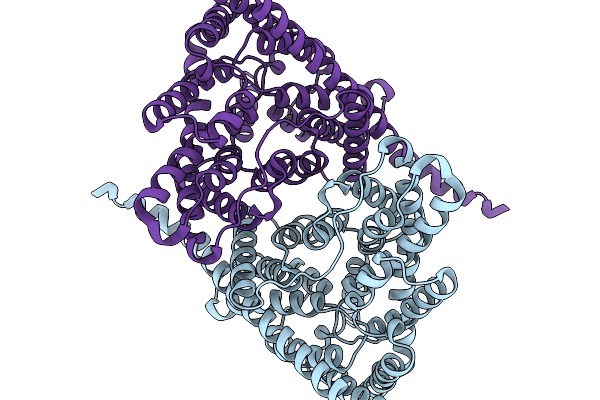 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-01-07 Classification: TRANSPORT PROTEIN Ligands: CL |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-01-07 Classification: TRANSPORT PROTEIN Ligands: CL |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-01-07 Classification: TRANSPORT PROTEIN Ligands: CL |
 |
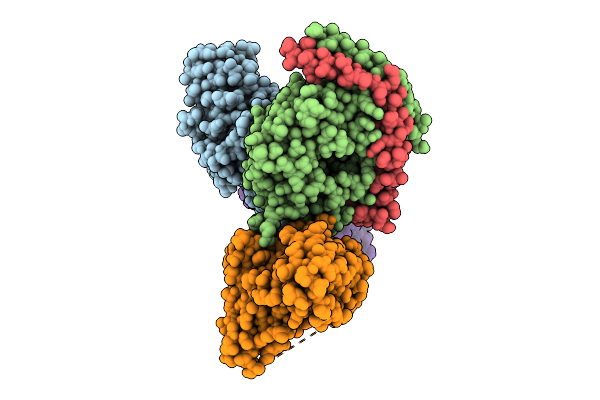
Local Refinement Of Drd2 Bound To Lsd In Complex With A Mini-Goa And Scfv16 Obtained By Cryo-Electron Microscopy (Cryoem)
Organism: Escherichia coli, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.28 Å Release Date: 2025-09-17 Classification: MEMBRANE PROTEIN Ligands: 7LD |
 |
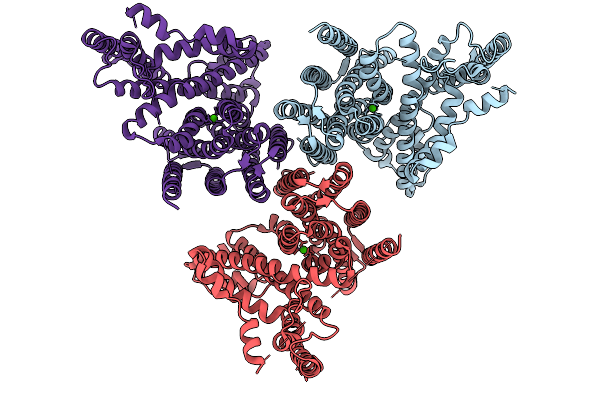
Global Reconstruction Of Drd2 Bound To Lsd In Complex With A Mini-Goa And Scfv16 Obtained By Cryo-Electron Microscopy (Cryoem)
Organism: Homo sapiens, Escherichia coli, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:2.32 Å Release Date: 2025-09-17 Classification: MEMBRANE PROTEIN Ligands: 7LD |
 |
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2025-09-10 Classification: MEMBRANE PROTEIN Ligands: CA |
 |
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: MEMBRANE PROTEIN Ligands: CA |
 |
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: MEMBRANE PROTEIN |
 |
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: MEMBRANE PROTEIN |
 |
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Resolution:4.29 Å Release Date: 2025-09-10 Classification: MEMBRANE PROTEIN |
 |
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Resolution:2.97 Å Release Date: 2025-09-10 Classification: MEMBRANE PROTEIN Ligands: CA |
 |
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:3.19 Å Release Date: 2025-04-02 Classification: MEMBRANE PROTEIN Ligands: 9UJ, A1LYC |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.36 Å Release Date: 2025-04-02 Classification: MEMBRANE PROTEIN Ligands: A1LYD |
 |
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-03-05 Classification: SIGNALING PROTEIN Ligands: A1AIW |
 |
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-03-05 Classification: SIGNALING PROTEIN Ligands: KCA |
 |
Organism: Homo sapiens, Mus musculus, Synthetic construct, Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-03-05 Classification: TRANSPORT PROTEIN/IMMUNE SYSTEM/INHIBITOR Ligands: A1BM8 |
 |
Organism: Homo sapiens, Mus musculus, Synthetic construct, Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:3.11 Å Release Date: 2025-03-05 Classification: TRANSPORT PROTEIN/IMMUNE SYSTEM Ligands: I2R |

