Search Count: 183
All
Selected
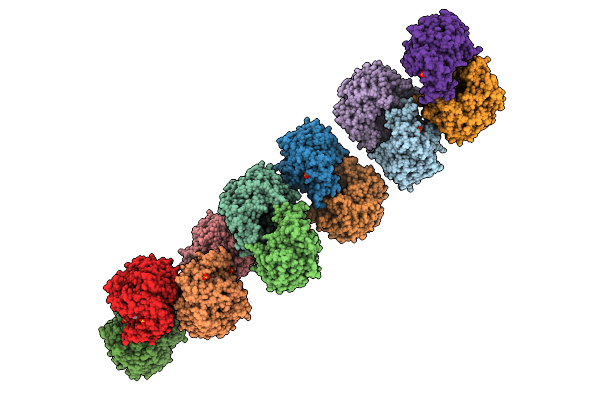 |
Organism: Caldicellulosiruptor sp.
Method: X-RAY DIFFRACTION Resolution:2.34 Å Release Date: 2025-10-15 Classification: HYDROLASE Ligands: GOL, SO4 |
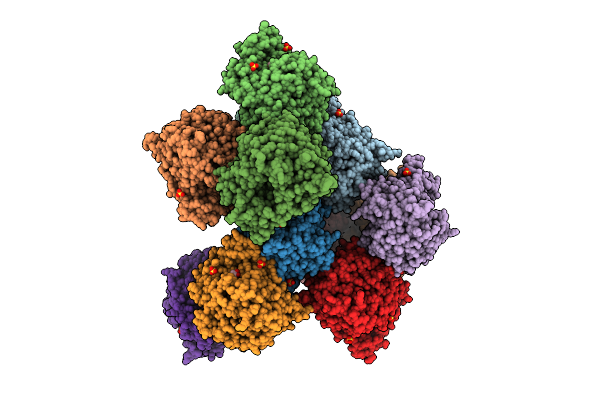 |
Crystal Structure Of Beta-Glucosidase Cabgl Mutant E163Q In Complex With Glucose
Organism: Caldicellulosiruptor sp.
Method: X-RAY DIFFRACTION Resolution:2.26 Å Release Date: 2025-10-15 Classification: HYDROLASE Ligands: GOL, BGC, SO4 |
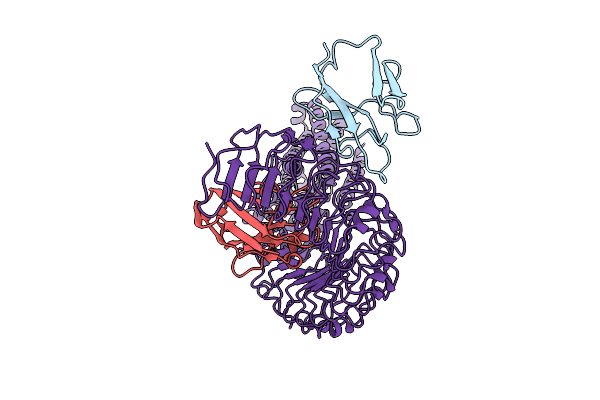 |
Organism: Camelus bactrianus, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-30 Classification: MEMBRANE PROTEIN |
 |
Organism: Camelus ferus, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.05 Å Release Date: 2025-07-16 Classification: MEMBRANE PROTEIN |
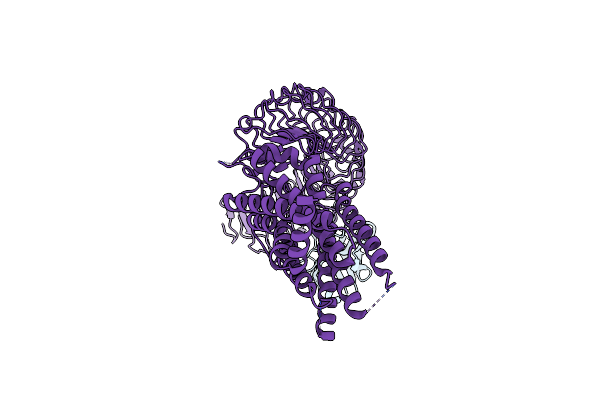 |
Organism: Homo sapiens, Camelus bactrianus
Method: ELECTRON MICROSCOPY Release Date: 2025-01-15 Classification: MEMBRANE PROTEIN |
 |
Organism: Camelus bactrianus, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-01-15 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens, Camelus bactrianus
Method: ELECTRON MICROSCOPY Release Date: 2024-12-04 Classification: MEMBRANE PROTEIN |
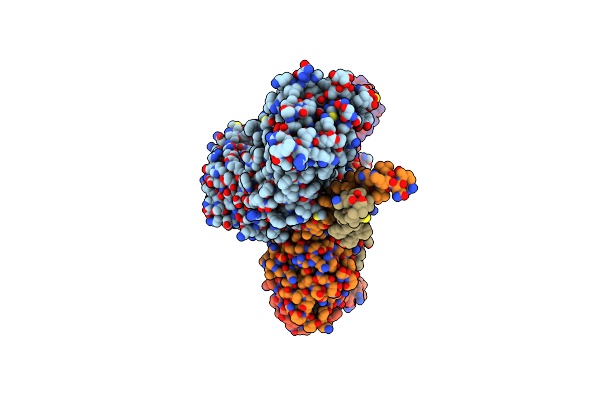 |
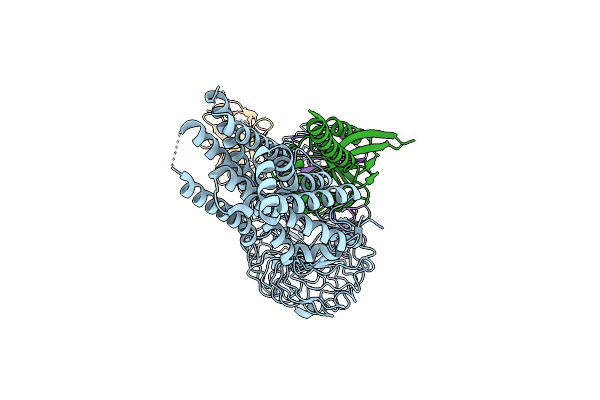
Organism: Homo sapiens, Camelus bactrianus
Method: ELECTRON MICROSCOPY Release Date: 2024-12-04 Classification: MEMBRANE PROTEIN |
 |
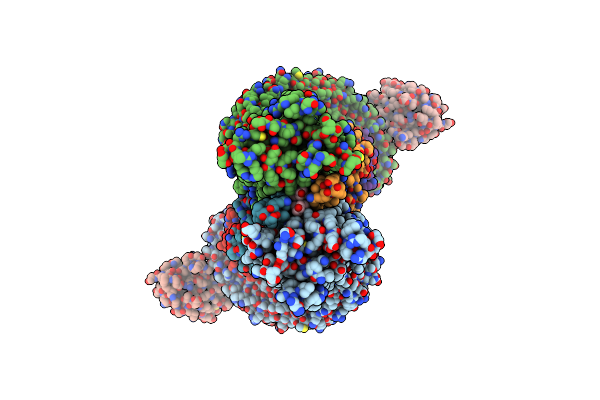
Organism: Homo sapiens, Camelus bactrianus
Method: ELECTRON MICROSCOPY Release Date: 2024-12-04 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens, Camelus bactrianus
Method: ELECTRON MICROSCOPY Release Date: 2024-12-04 Classification: MEMBRANE PROTEIN Ligands: CLR |
 |
Crystal Structure Of Cellulosomal Double-Dockerin Module Of Clo1313_0689 From Clostridium Thermocellum
Organism: Acetivibrio thermocellus dsm 1313
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2024-04-03 Classification: PROTEIN BINDING Ligands: CA |
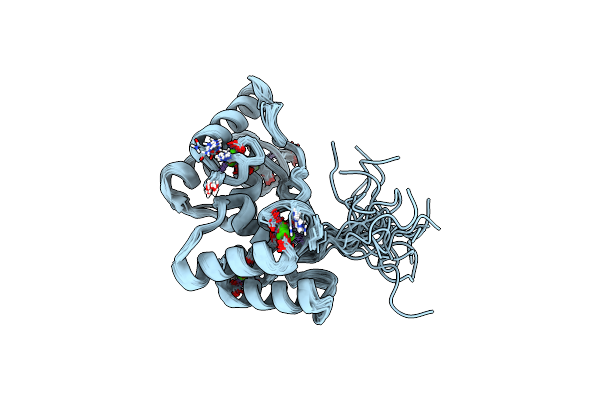 |
Solution Nmr Structure Of Cellulosomal Double-Dockerin Module Of Clo1313_0689 From Clostridium Thermocellum
Organism: Acetivibrio thermocellus dsm 1313
Method: SOLUTION NMR Release Date: 2024-04-03 Classification: PROTEIN BINDING Ligands: CA |
 |
Organism: Rhodococcus opacus
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2024-01-31 Classification: HYDROLASE Ligands: SO4 |
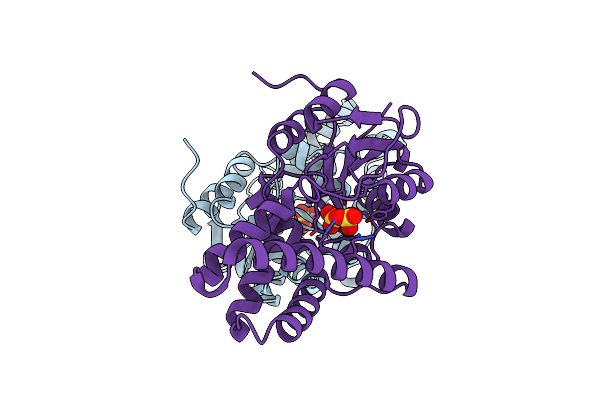 |
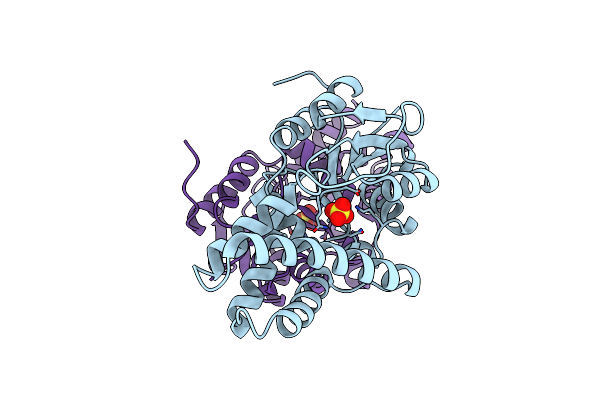
Crystal Structure Of Cis-Epoxysuccinate Hydrolases Rhcesh[L] Complexed With Sulfate Ions
Organism: Rhodococcus opacus
Method: X-RAY DIFFRACTION Resolution:1.94 Å Release Date: 2024-01-31 Classification: HYDROLASE Ligands: SO4 |
 |
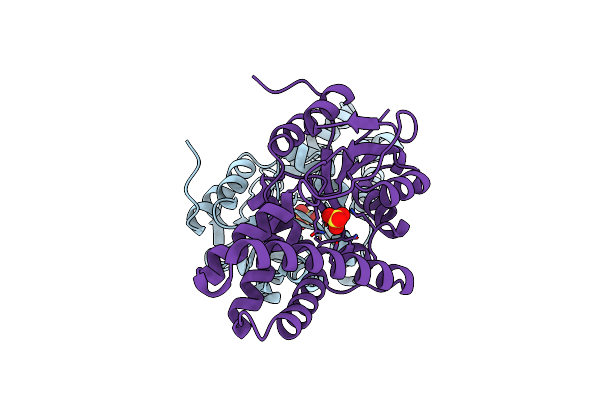
Crystal Structure Of Cis-Epoxysuccinate Hydrolases Rhcesh[L] Mutant D193A Complexed With Sulfate Ions
Organism: Rhodococcus opacus
Method: X-RAY DIFFRACTION Resolution:2.06 Å Release Date: 2024-01-31 Classification: HYDROLASE Ligands: SO4 |
 |
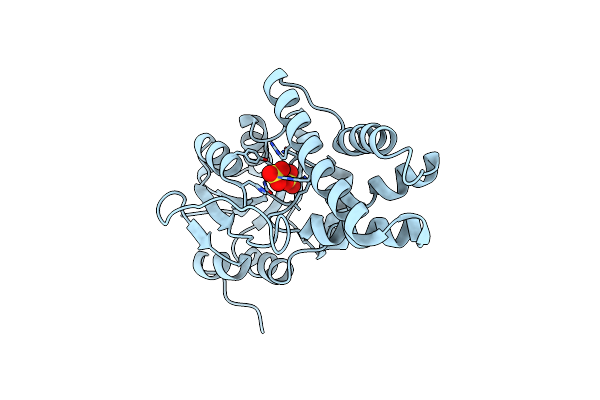
Organism: Rhodococcus opacus
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2024-01-31 Classification: HYDROLASE Ligands: SO4 |
 |
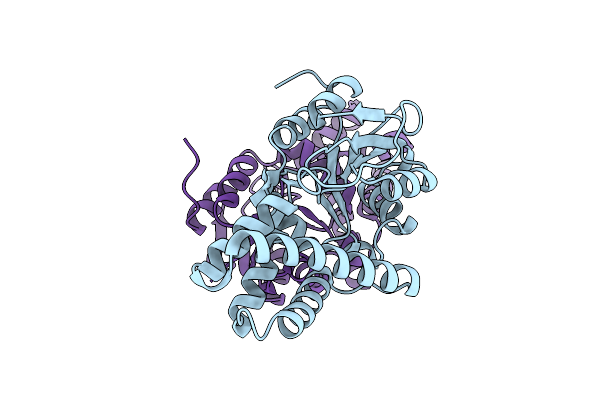
Crystal Structure Of Cis-Epoxysuccinate Hydrolases Rhcesh[L] Mutant D18N Complexed With Sulfate Ions
Organism: Rhodococcus opacus
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2024-01-31 Classification: HYDROLASE Ligands: SO4 |
 |
Organism: Rhodococcus opacus
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2024-01-31 Classification: HYDROLASE |
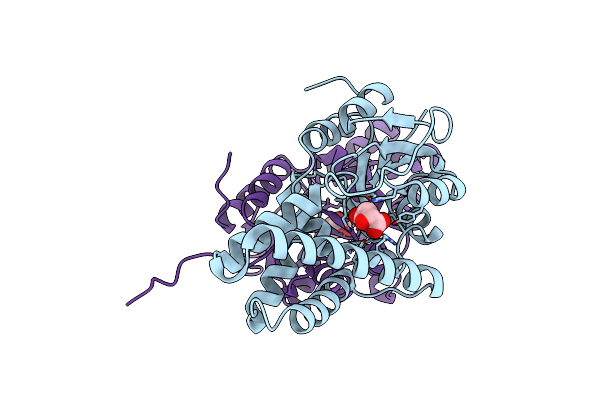 |
Crystal Structure Of Cis-Epoxysuccinate Hydrolases Rhcesh[L] Mutant E212Q Complexed With L-Ta.
Organism: Rhodococcus opacus
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2024-01-31 Classification: HYDROLASE Ligands: TLA |
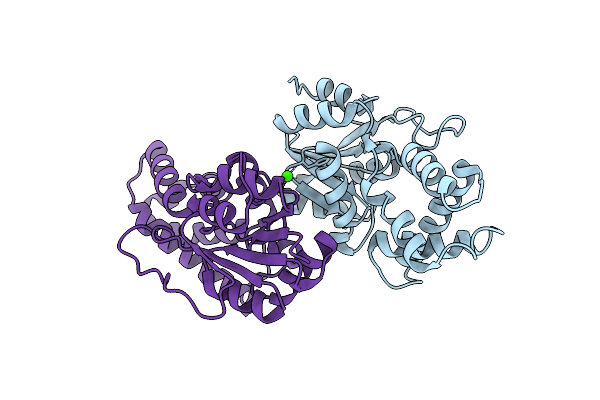 |
Organism: Klebsiella sp. bk-58
Method: X-RAY DIFFRACTION Resolution:2.02 Å Release Date: 2024-01-31 Classification: HYDROLASE Ligands: CA |

