Search Count: 48
All
Selected
 |
Gii.23/24/25 Noroviruses Recognize Glycans Via A Conventional Glycan-Binding Site
Organism: Norovirus
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-03-04 Classification: VIRAL PROTEIN |
 |
Gii.23/24/25 Noroviruses Recognize Glycans Via A Conventional Glycan-Binding Site
Organism: Norovirus gii
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-02-25 Classification: VIRAL PROTEIN |
 |
Organism: Norovirus gii
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-02-25 Classification: VIRAL PROTEIN |
 |
Organism: Norovirus gii
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2026-02-25 Classification: VIRAL PROTEIN Ligands: P |
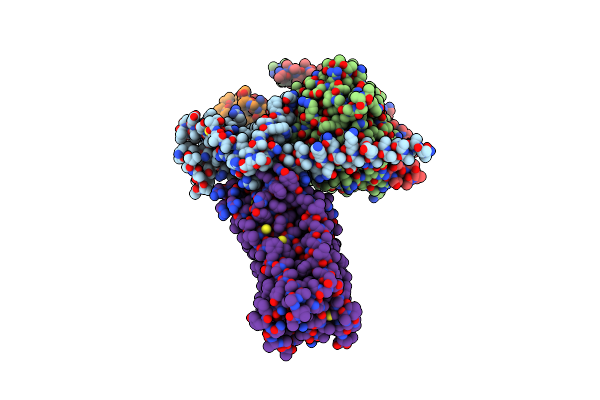 |
Organism: Homo sapiens, Lama glama
Method: ELECTRON MICROSCOPY Release Date: 2024-01-10 Classification: SIGNALING PROTEIN Ligands: Z2C |
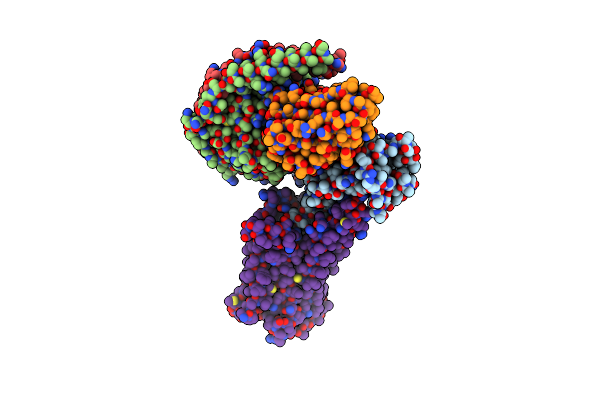 |
Organism: Homo sapiens, Lama glama
Method: ELECTRON MICROSCOPY Release Date: 2024-01-10 Classification: SIGNALING PROTEIN Ligands: YZV |
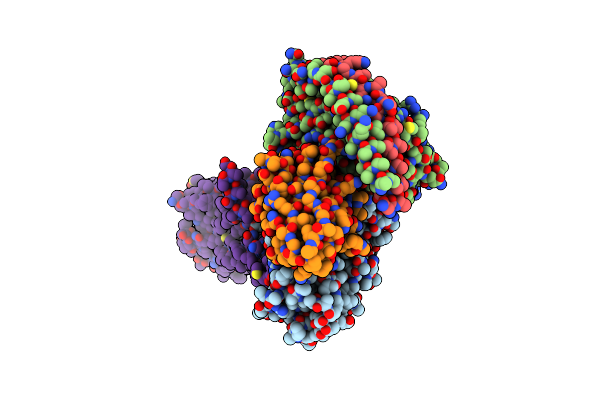 |
Organism: Homo sapiens, Lama glama
Method: ELECTRON MICROSCOPY Release Date: 2024-01-10 Classification: SIGNALING PROTEIN Ligands: P2E |
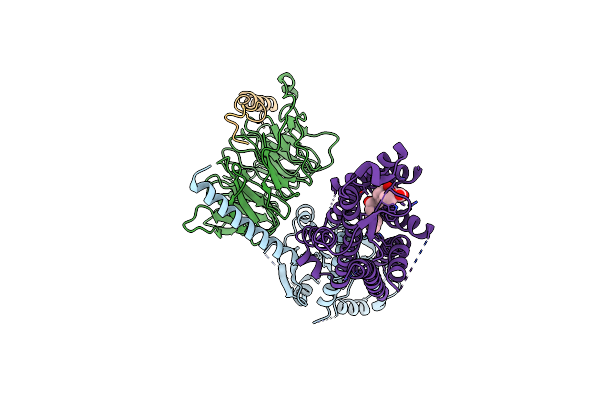 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-01-10 Classification: SIGNALING PROTEIN Ligands: P2E |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-01-03 Classification: MEMBRANE PROTEIN Ligands: YX9 |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-01-03 Classification: MEMBRANE PROTEIN Ligands: P2E |
 |
Functional And Structural Characterization Of Norovirus Gii.6 In Recognizing Histo-Blood Group Antigens
Organism: Norovirus gii
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2023-06-07 Classification: VIRAL PROTEIN Ligands: GOL |
 |
Functional And Structural Characterization Of Norovirus Gii.6 In Recognizing Histo-Blood Group Antigens
Organism: Norovirus gii
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2023-05-03 Classification: VIRAL PROTEIN |
 |
Cryo-Em Structure Of The Relaxin Receptor Rxfp1 In Complex With Heterotrimeric Gs
Organism: Homo sapiens, Lama glama
Method: ELECTRON MICROSCOPY Release Date: 2023-02-15 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2022-09-14 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2022-09-14 Classification: MEMBRANE PROTEIN |
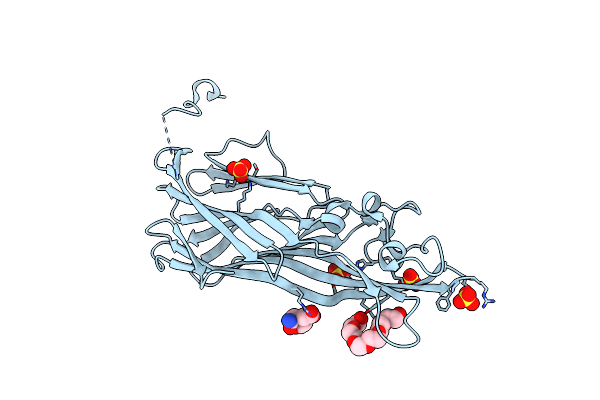 |
Crystal Structure Of Leukotoxin Luke From Staphylococcus Aureus At 1.9 Angstrom Resolution
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2022-04-06 Classification: TOXIN Ligands: P6G, MHA, SO4 |
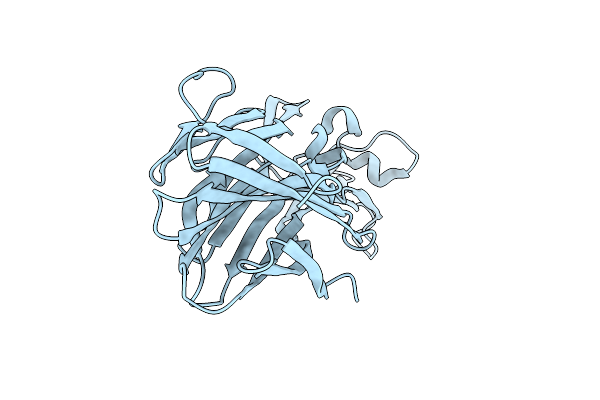 |
Crystal Structure Of Leukotoxin Luke From Staphylococcus Aureus At 1.5 Angstrom Resolution
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:1.46 Å Release Date: 2022-04-06 Classification: TOXIN Ligands: CL |
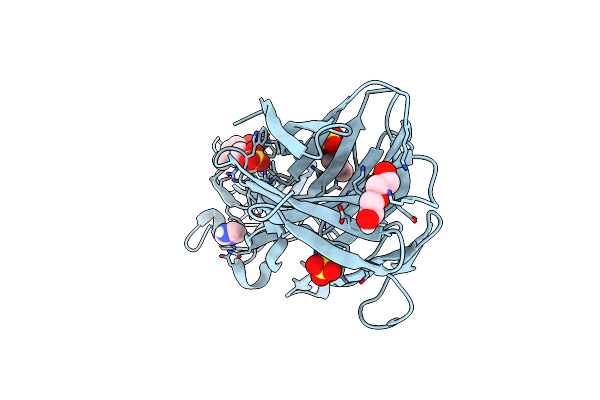 |
Crystal Structure Of Leukotoxin Luke From Staphylococcus Aureus In Complex With P-Cresyl Sulfate
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2022-04-06 Classification: TOXIN Ligands: PEG, IMD, SO4, 6EI |
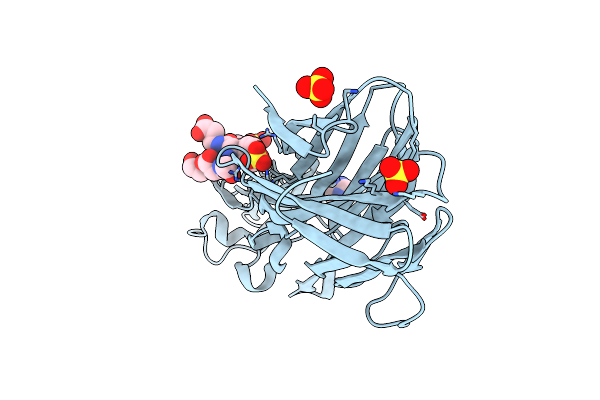 |
Crystal Structure Of Leukotoxin Luke From Staphylococcus Aureus In Complex With A Doubly Sulfated Ccr2 N-Terminal Peptide
Organism: Staphylococcus aureus, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2022-04-06 Classification: TOXIN Ligands: IMD, SO4 |
 |
Crystal Structure Of Leukotoxin Luke From Staphylococcus Aureus In Complex With A Sulfated Ackr1 N-Terminal Peptide
Organism: Staphylococcus aureus, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2022-04-06 Classification: TOXIN |

