Search Count: 18
All
Selected
 |
Joint X-Ray/Neutron Structure Of Wild-Type Bacillus Halodurans Rnase H1 In The Apo-Form
Organism: Halalkalibacterium halodurans
Method: X-RAY DIFFRACTION, NEUTRON DIFFRACTION Resolution:2.48 Å, 1.90 Å Release Date: 2026-03-18 Classification: HYDROLASE Ligands: SO4, DOD |
 |
Joint X-Ray/Neutron Structure Of D132N Bacillus Halodurans Rnase H1 In The Apo-Form
Organism: Halalkalibacterium halodurans
Method: X-RAY DIFFRACTION, NEUTRON DIFFRACTION Resolution:2.4 Å, 2.20 Å Release Date: 2026-03-18 Classification: HYDROLASE Ligands: SO4, DOD |
 |
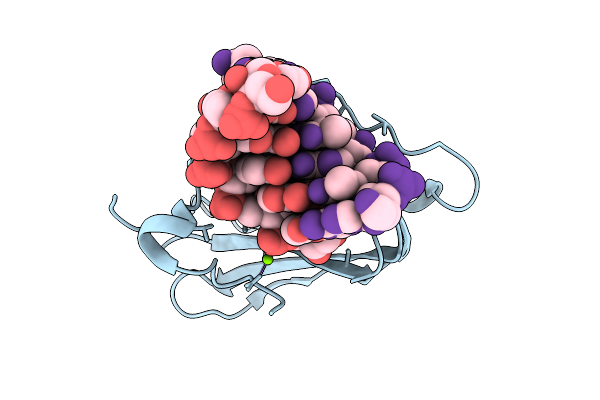
Room-Temperature X-Ray Structure Of D132N Bacillus Halodurans Rnase H1 In Complex With Rna/Dna Duplex
Organism: Halalkalibacterium halodurans
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-03-18 Classification: HYDROLASE/RNA/DNA Ligands: MG, PO3 |
 |
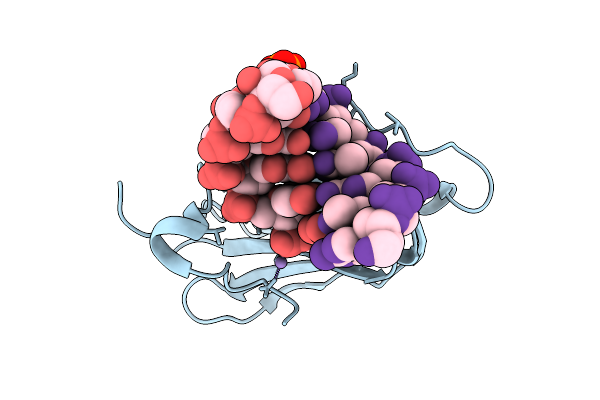
Room-Temperature X-Ray Structure Of D132N Bacillus Halodurans Rnase H1 In Complex With Complementary Rna/Dna Duplex
Organism: Halalkalibacterium halodurans
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-03-18 Classification: HYDROLASE/RNA/DNA Ligands: MG |
 |
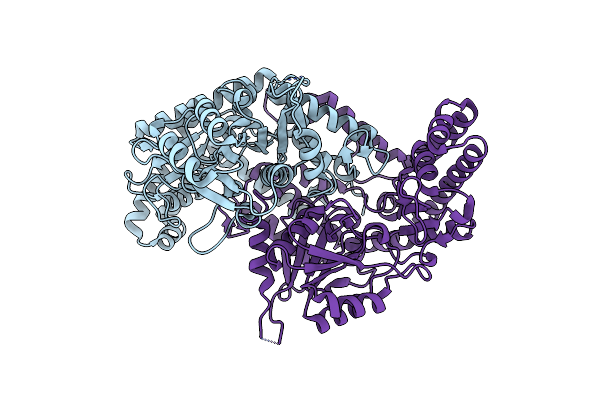
100K X-Ray Structure Of Mixed Metal D132N Bacillus Halodurans Rnase H1 Complex With Rna/Dna Duplex
Organism: Halalkalibacterium halodurans
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-18 Classification: HYDROLASE/RNA/DNA Ligands: MN, MG, PO3 |
 |
Room-Temperature X-Ray Structure Of Human Mitochondrial Serine Hydroxymethyltransferase (Hshmt2)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2023-08-16 Classification: TRANSFERASE Ligands: CL |
 |
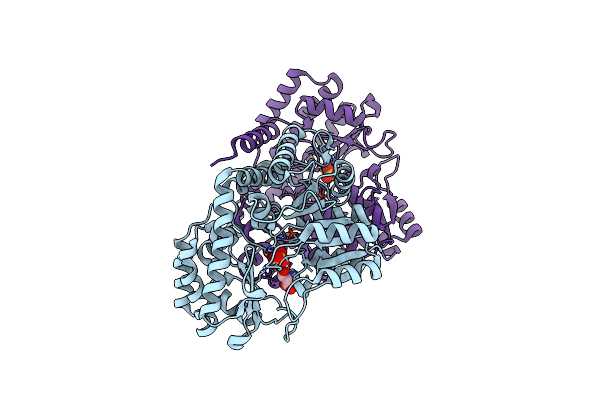
Room-Temperature X-Ray Structure Of Thermus Thermophilus Serine Hydroxymethyltransferase (Shmt) Bound With D-Ser In A Pseudo-Michaelis Complex
Organism: Thermus thermophilus hb8
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2023-08-16 Classification: TRANSFERASE Ligands: DSN, SO4 |
 |
Joint X-Ray/Neutron Structure Of Thermus Thermophilus Serine Hydroxymethyltransferase (Tthshmt) In Internal Aldimine State With L-Ser Bound In A Pre-Michalis Complex
Organism: Thermus thermophilus hb8
Method: X-RAY DIFFRACTION, NEUTRON DIFFRACTION Resolution:2.30 Å, 2.00 Å Release Date: 2023-08-16 Classification: TRANSFERASE Ligands: SO4, SER, DOD |
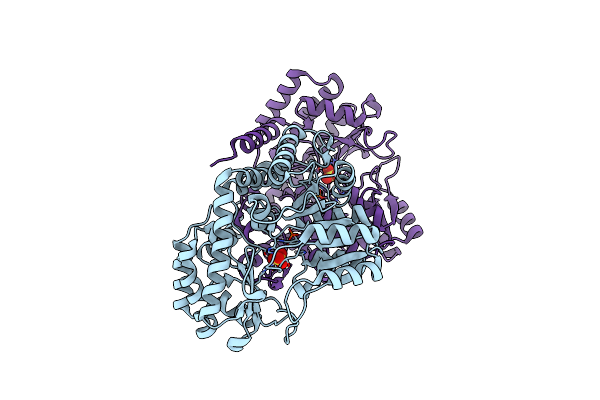 |
Joint X-Ray/Neutron Structure Of Thermus Thermophilus Serine Hydroxymethyltransferase (Tthshmt) In Internal Aldimine State
Organism: Thermus thermophilus hb8
Method: X-RAY DIFFRACTION, NEUTRON DIFFRACTION Resolution:2.30 Å, 2.00 Å Release Date: 2023-08-16 Classification: TRANSFERASE Ligands: SO4, DOD |
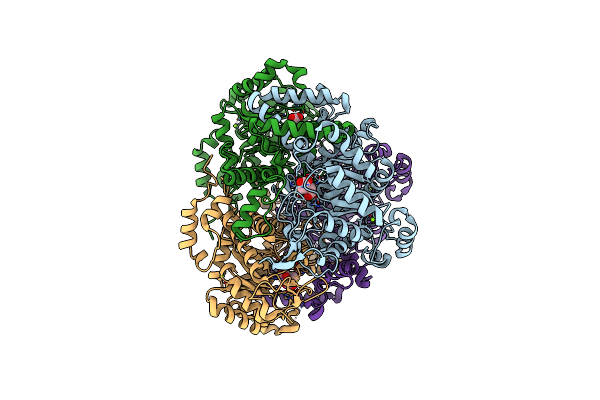 |
Crystal Structure Of Itaconate Modified Mycobaterium Tuberculosis Isocitrate Lyase
Organism: Mycobacterium tuberculosis (strain atcc 25618 / h37rv)
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2020-10-21 Classification: LYASE Ligands: ITN, MG |
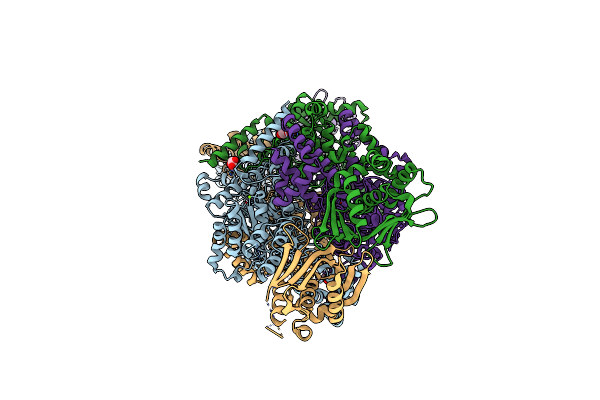 |
Organism: Mycobacterium tuberculosis (strain cdc 1551 / oshkosh)
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2019-08-14 Classification: LYASE Ligands: GOL, MG |
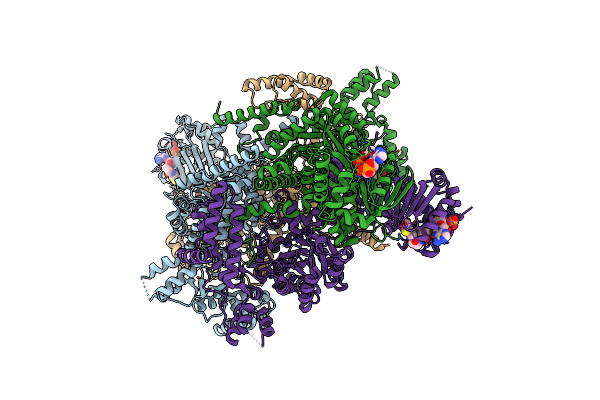 |
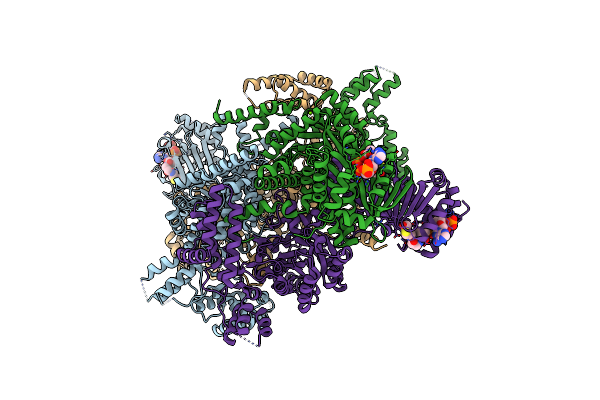
Crystal Structure Of Mycobacterium Tuberculosis Icl2 In Complex With Acetyl-Coa, Form I
Organism: Mycobacterium tuberculosis (strain cdc 1551 / oshkosh)
Method: X-RAY DIFFRACTION Resolution:2.67 Å Release Date: 2019-08-14 Classification: LYASE Ligands: ACO |
 |
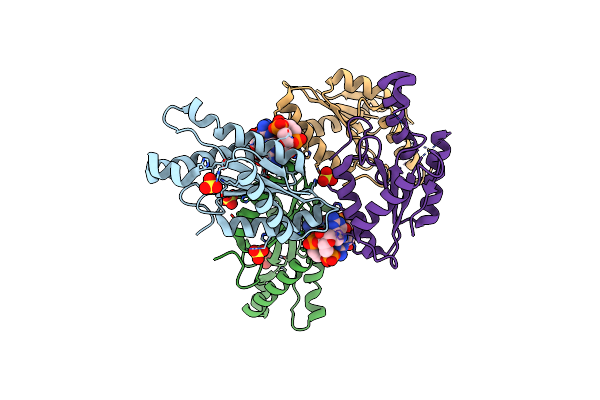
Crystal Structure Of Mycobacterium Tuberculosis Icl2 In Complex With Acetyl-Coa
Organism: Mycobacterium tuberculosis (strain cdc 1551 / oshkosh)
Method: X-RAY DIFFRACTION Resolution:2.36 Å Release Date: 2019-08-14 Classification: LYASE Ligands: ACO, MG |
 |
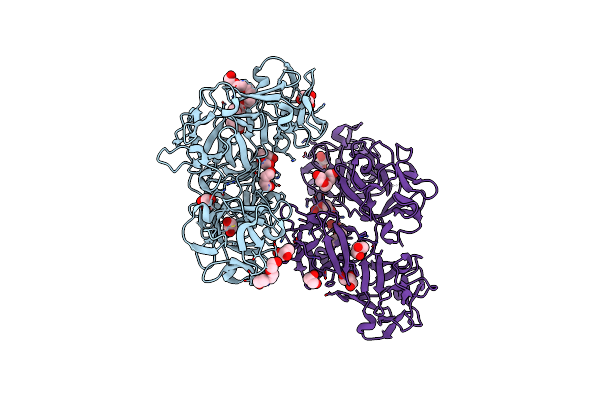
Crystal Structure Of The C-Di-Gmp-Bound Ggdef Domain Of P. Fluorescens Gcbc
Organism: Pseudomonas fluorescens (strain pf0-1)
Method: X-RAY DIFFRACTION Resolution:2.99 Å Release Date: 2015-12-30 Classification: MEMBRANE PROTEIN Ligands: C2E, SO4 |
 |
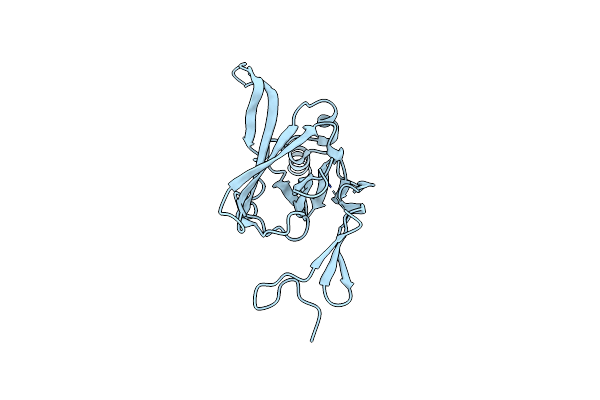
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2011-06-29 Classification: STRUCTURAL PROTEIN Ligands: GOL, 12P, 1PE, PG0 |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2005-03-08 Classification: METAL TRANSPORT |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.41 Å Release Date: 2005-03-08 Classification: ALLERGEN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2000-01-10 Classification: IMMUNE SYSTEM Ligands: NAG |

