Search Count: 48
All
Selected
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-01-07 Classification: TRANSFERASE/TRANSFERASE INHIBITOR Ligands: A1C0G, SO4 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.97 Å Release Date: 2026-01-07 Classification: TRANSFERASE/TRANSFERASE INHIBITOR Ligands: A1C0I, FRU, SO4, GOL |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-01-07 Classification: TRANSFERASE/TRANSFERASE INHIBITOR Ligands: A1C0K, FRU, SO4 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.11 Å Release Date: 2026-01-07 Classification: TRANSFERASE/TRANSFERASE INHIBITOR Ligands: A1C0J, SO4 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.02 Å Release Date: 2026-01-07 Classification: TRANSFERASE/TRANSFERASE INHIBITOR Ligands: A1C0H, FRU |
 |
Structure Of The C-Terminal Beta Helix Domain Of The Bdellovibrio Bacteriovorus Bd3182 Fibre
Organism: Bdellovibrio bacteriovorus hd100
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2023-10-25 Classification: CELL ADHESION |
 |
Structure Of The C-Terminal Domain Of The Bdellovibrio Bacteriovorus Bd2133 Fibre
Organism: Bdellovibrio bacteriovorus hd100
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2023-10-25 Classification: CELL ADHESION Ligands: SO4, NA, CL |
 |
Organism: Bdellovibrio bacteriovorus hd100
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2023-10-25 Classification: CELL ADHESION |
 |
Organism: Bdellovibrio bacteriovorus hd100
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2023-10-25 Classification: CELL ADHESION |
 |
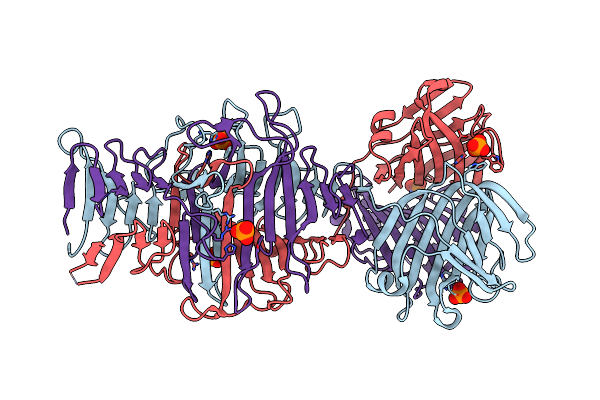
Structure Of The C-Terminal Domains Of The Bdellovibrio Bacteriovorus Bd2439 Fibre In Complex With Glcnac
Organism: Bdellovibrio bacteriovorus hd100
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2023-10-25 Classification: CELL ADHESION Ligands: IOD, PG4, PEG, NAG, SO4, B3P |
 |
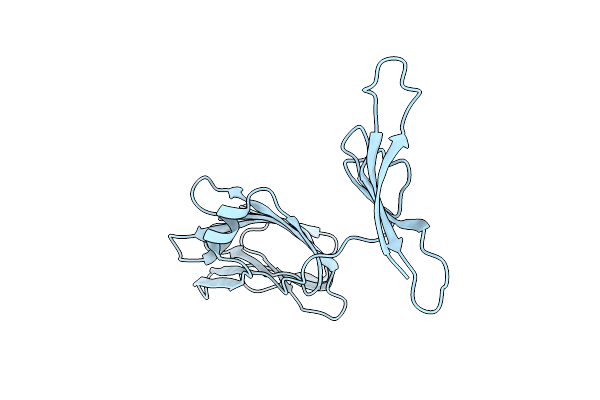
Organism: Bdellovibrio bacteriovorus hd100
Method: X-RAY DIFFRACTION Resolution:2.56 Å Release Date: 2023-10-25 Classification: CELL ADHESION Ligands: PO4 |
 |
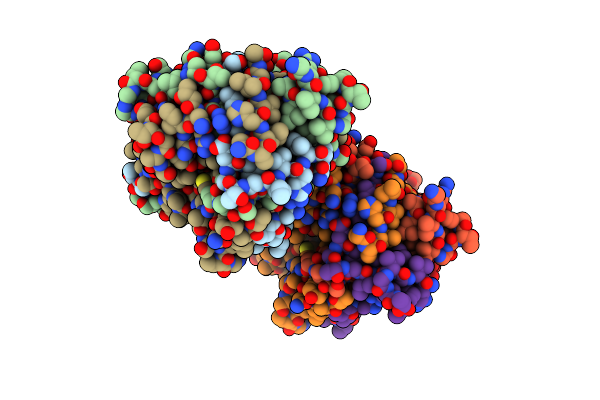
Structure Of The C-Terminal Domains Of The Bdellovibrio Bacteriovorus Bd1334 Fibre
Organism: Bdellovibrio bacteriovorus hd100
Method: X-RAY DIFFRACTION Resolution:1.41 Å Release Date: 2023-10-25 Classification: CELL ADHESION |
 |
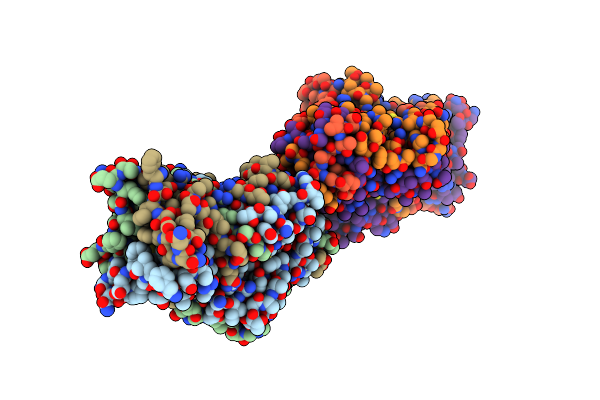
Structure Of The C-Terminal Beta Helix Domain Of The Bdellovibrio Bacteriovorus Bd3182 Fibre
Organism: Bdellovibrio bacteriovorus hd100
Method: X-RAY DIFFRACTION Resolution:1.12 Å Release Date: 2023-10-25 Classification: CELL ADHESION |
 |
Structure Of The C-Terminal Beta Helix Domain Of The Bdellovibrio Bacteriovorus Bd3182 Fibre
Organism: Bdellovibrio bacteriovorus hd100
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-10-25 Classification: CELL ADHESION Ligands: EDO |
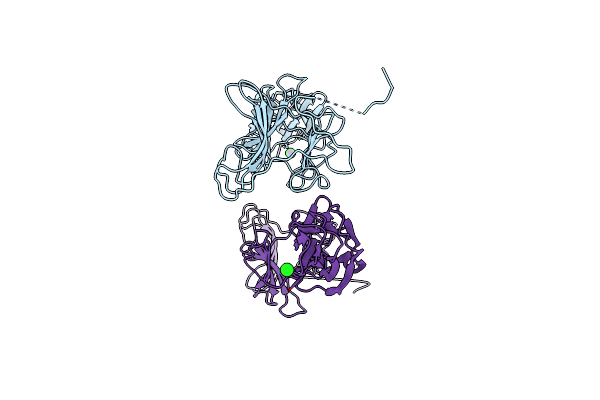 |
Structure Of The C-Terminal Domains Of The Bdellovibrio Bacteriovorus Bd2133 Fibre
Organism: Bdellovibrio bacteriovorus hd100
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2023-10-25 Classification: CELL ADHESION Ligands: CL, NA |
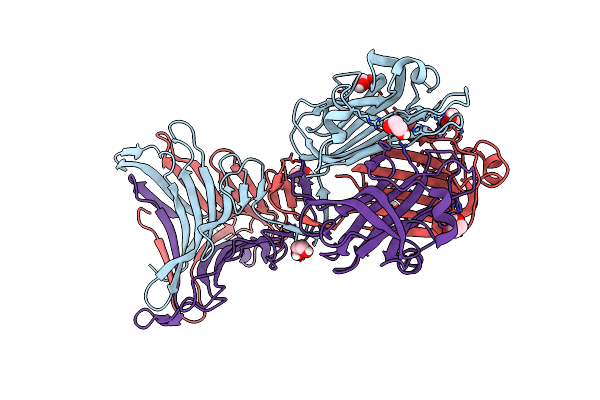 |
Crystal Structure Of Bdellovibrio Bacteriovorus Bd2439 Fibre C-Terminal Domains With Ethylene Glycol
Organism: Bdellovibrio bacteriovorus hd100
Method: X-RAY DIFFRACTION Resolution:1.53 Å Release Date: 2023-10-25 Classification: CELL ADHESION Ligands: EDO |
 |
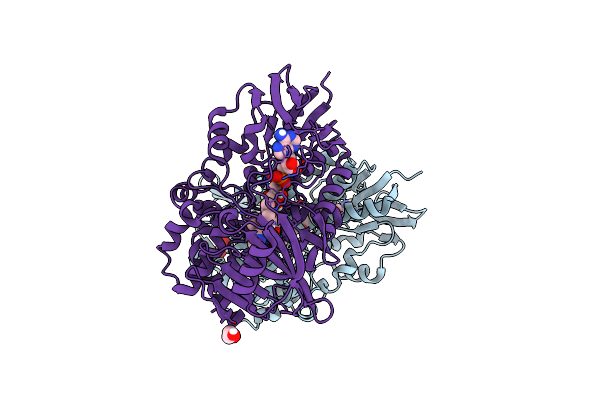
The Structure Of Nica2 Variant F104L/A107T/S146I/G317D/H368R/L449V/N462S From Pseudomonas Putida
Organism: Pseudomonas putida s16
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2023-07-26 Classification: FLAVOPROTEIN Ligands: FDA |
 |
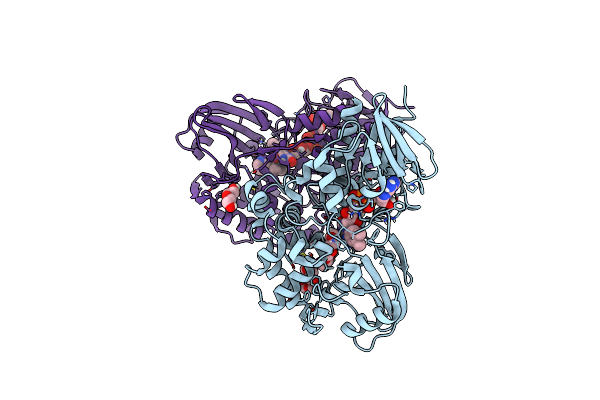
The Structure Of Nica2 Variant F104L/A107T/S146I/G317D/H368R/L449V/N462S In Complex With N-Methylmyosmine
Organism: Pseudomonas putida s16
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2023-07-26 Classification: FLAVOPROTEIN Ligands: NCT, FDA, EDO |
 |
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2023-07-26 Classification: FLAVOPROTEIN, OXIDOREDUCTASE Ligands: FDA, PG4, PEG, TXU |
 |
Crystal Structure Of Myeloperoxidase Subform C (Mpo) Complex With Compound-29 A.K.A 7-(1,2-Diphenylethyl)-1H-[1,2,3]Triazolo[4,5-B]Pyridin-5-Amine
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.23 Å Release Date: 2020-10-14 Classification: OXIDOREDUCTASE Ligands: CL, CA, NAG, HEC, UEY, MAN |

