Search Count: 176
All
Selected
 |
Organism: Adeno-associated virus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: VIRUS |
 |
Cryo-Em Structure Of J601-1B2 Fab In Complex With Hiv-1 Bg505 Ds-Sosip Env Trimer
Organism: Human immunodeficiency virus 1, Macaca mulatta
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG, MAN |
 |
Cryo-Em Structure Of J601-A6 Fab In Complex With Hiv-1 Bg505 Ds-Sosip Env Trimer
Organism: Human immunodeficiency virus 1, Macaca mulatta
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Cryo-Em Structure Of K001-A1 Fab In Complex With Hiv-1 459C-Opt Rns Ds-Sosip Env Trimer
Organism: Macaca mulatta, Human immunodeficiency virus 1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Organism: Human immunodeficiency virus 1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Organism: Human immunodeficiency virus 1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2025-11-12 Classification: LIPID BINDING PROTEIN Ligands: A1EN5 |
 |
Cryo-Em Structure Of Neutralizing Murine Antibody Ws.Hsv-1.24 In Complex With Hsv-1 Glycoprotein B Trimer Gb-Ecto.516P.531E
Organism: Human alphaherpesvirus 1, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Cryo-Em Structure Of Neutralizing Human Antibody D48 In Complex With Hsv-1 Glycoprotein B Trimer Gb-Ecto.516P.531E.Ds
Organism: Human alphaherpesvirus 1, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: VIRAL PROTEIN Ligands: NAG |
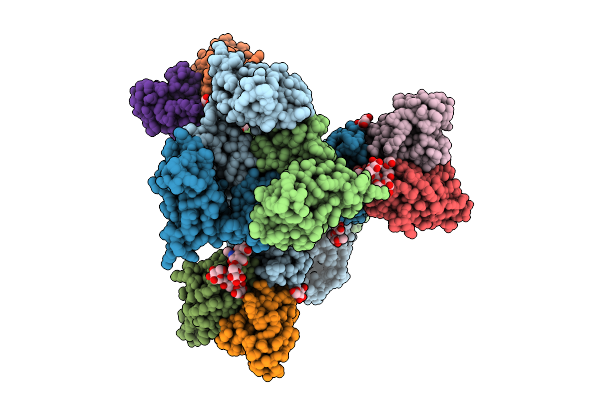 |
Cryo-Em Structure Of Neutralizing Human Antibody D48 In Complex With Hsv-1 Glycoprotein B Trimer Gb-Ecto.516P
Organism: Human alphaherpesvirus 1, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: VIRAL PROTEIN Ligands: NAG |
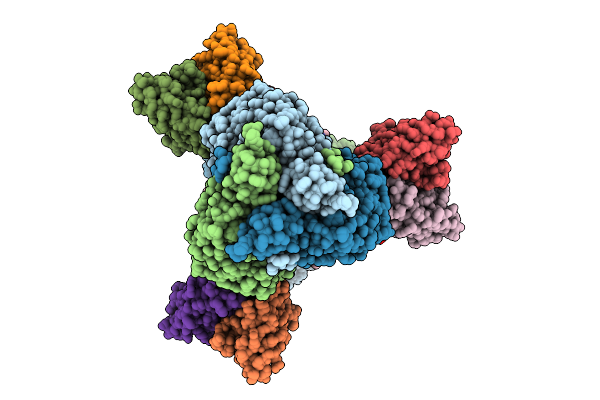 |
Cryo-Em Structure Of Neutralizing Human Antibody D48 In Complex With Hsv-1 Glycoprotein B Trimer Gb-Ecto.516P.531E
Organism: Human alphaherpesvirus 1, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: VIRAL PROTEIN Ligands: NAG |
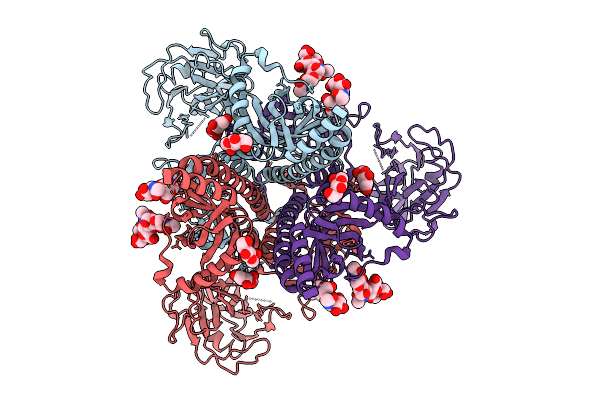 |
Cryo-Em Structure Of Gb-Ecto.516P.531E.Ds, A Prefusion-Stabilized Hsv-1 Glycoprotein B Extracellular Domain
Organism: Human alphaherpesvirus 1
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: VIRAL PROTEIN Ligands: NAG |
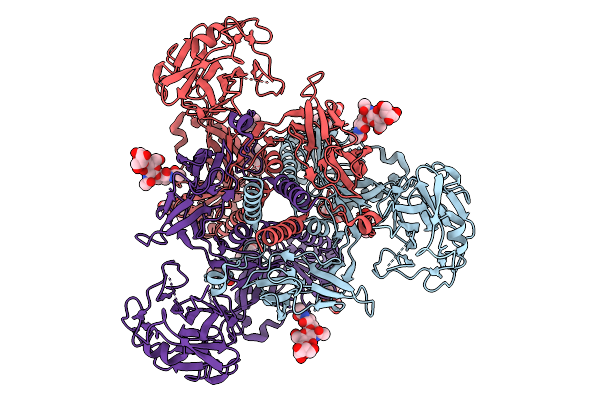 |
Cryo-Em Structure Of Gb-Ecto.516P.531E, A Prefusion-Stabilized Hsv-1 Glycoprotein B Extracellular Domain
Organism: Human alphaherpesvirus 1
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Cryo-Em Structure Of Gb-Ecto.516P, An Hsv-1 Glycoprotein B Extracellular Domain
Organism: Human alphaherpesvirus 1
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Structural Basis Of The Bifunctionality Of M. Salinexigens Zyf650T Glucosylglycerol Phosphorylase In Glucosylglycerol Catabolism
Organism: Marinobacter salinexigens
Method: X-RAY DIFFRACTION Resolution:2.72 Å Release Date: 2025-09-10 Classification: STRUCTURAL PROTEIN Ligands: GLC, GOL, NA, G1P |
 |
Structural Basis Of The Bifunctionality Of M. Salinexigens Zyf650T Glucosylglycerol Phosphorylase In Glucosylglycerol Catabolism
Organism: Marinobacter salinexigens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-09-10 Classification: STRUCTURAL PROTEIN Ligands: GOL, GLC, PEG |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-09-03 Classification: TRANSPORT PROTEIN Ligands: URE, PLM |
 |
Structural Basis Of The Bifunctionality Of M. Salinexigens Zyf650T Glucosylglycerol Phosphorylase In Glucosylglycerol Catabolism
Organism: Marinobacter salinexigens
Method: X-RAY DIFFRACTION Resolution:2.74 Å Release Date: 2025-09-03 Classification: STRUCTURAL PROTEIN Ligands: PEG, TRS, PO4, SO4 |
 |
Structural Basis Of Elongation Factor Tu In Regulating Photoinhibition In Synechococcus Sp. Pcc 7942.
Organism: Synechococcus elongatus (strain atcc 33912 / pcc 7942 / fachb-805)
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2025-03-19 Classification: PHOTOSYNTHESIS Ligands: GOL, GDP, BME, MG |
 |
Organism: Macaca mulatta, Simian immunodeficiency virus
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2025-01-15 Classification: IMMUNE SYSTEM/Viral Protein Ligands: PG4, GOL |

