Search Count: 331
All
Selected
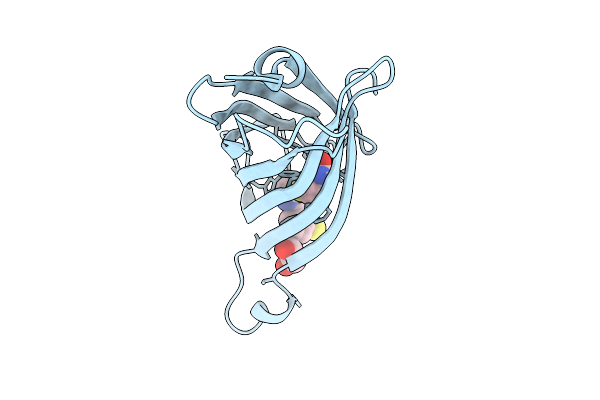 |
Streptavidin K121M With A Thiophenol Cofactor As Artificial Hydrogen Atom Transferase
Organism: Streptomyces avidinii
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: A1I8C, EDO |
 |
Apo Multidrug Resistance-Associated Protein 2 In Complex With Amp-Pnp In Rest State
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: MEMBRANE PROTEIN Ligands: MG, ANP |
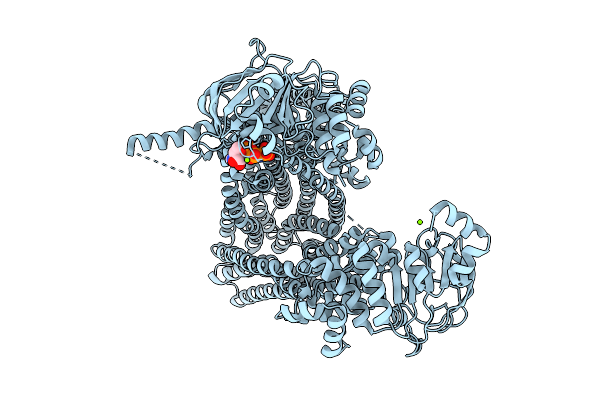 |
Multidrug Resistance-Associated Protein 2 In Complex With Amp-Pnp In Active State
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: MG, ANP |
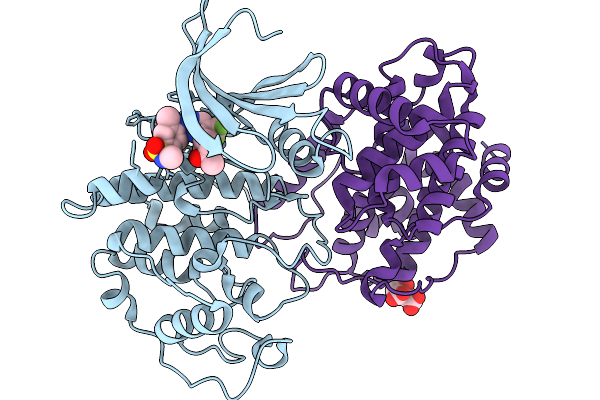 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2026-03-04 Classification: CELL CYCLE Ligands: A1B7H, CIT |
 |
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: POV, AV0, ERG |
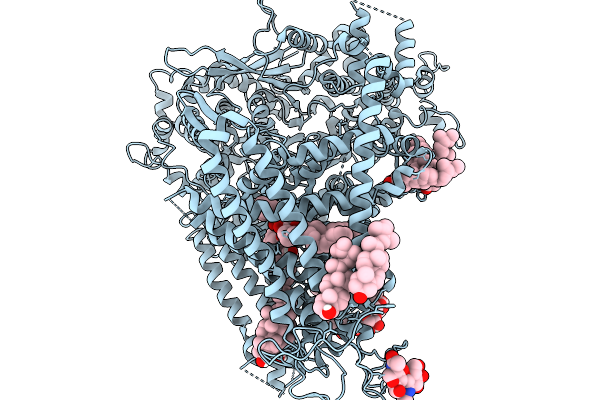 |
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: ERG, A1E06, POV |
 |
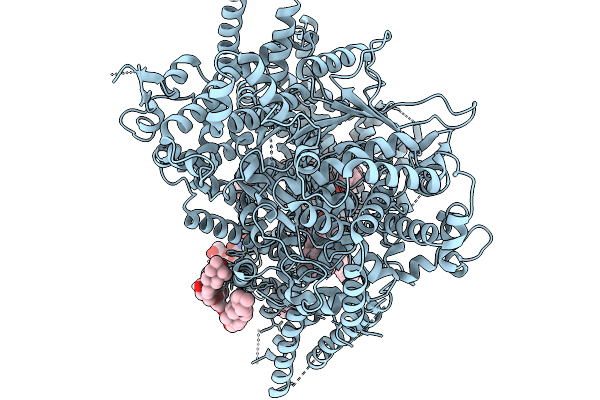
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c)
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: AV0, ERG, POV |
 |
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: ERG, A1E06, POV |
 |
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: ERG, A1E06, POV |
 |
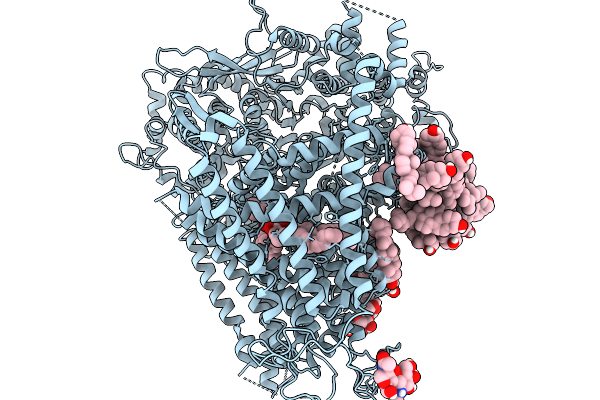
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c)
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN |
 |
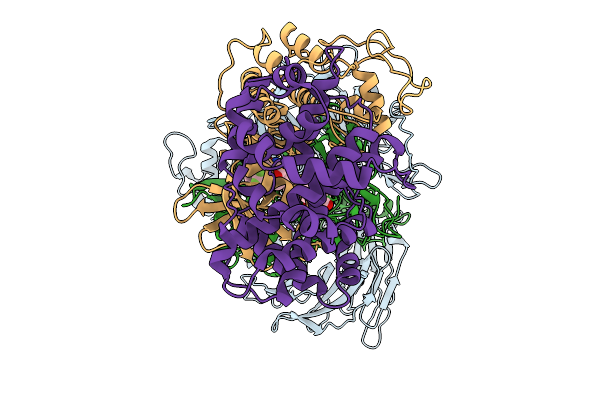
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: ERG, A1E06, POV |
 |
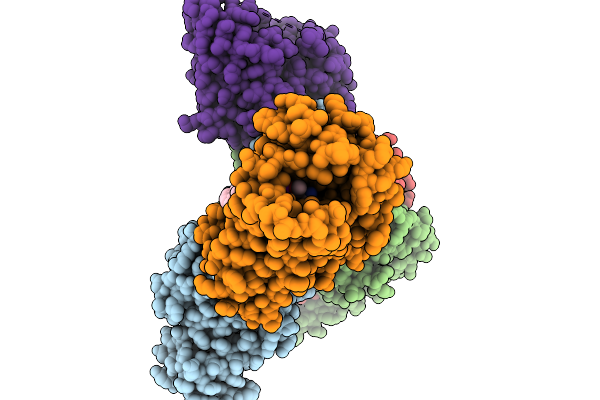
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: CELL CYCLE Ligands: A1B7G, ZN |
 |
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2026-01-14 Classification: MEMBRANE PROTEIN Ligands: CLR, A1EK5 |
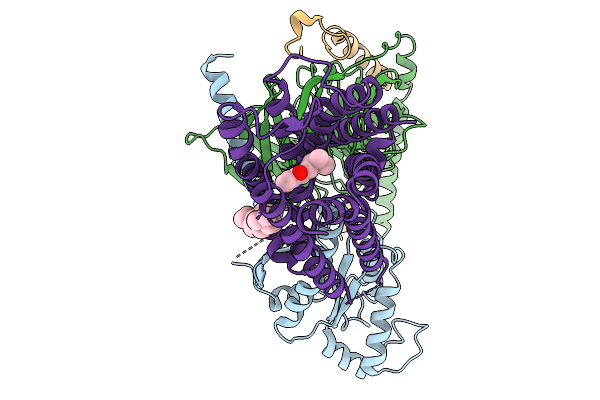 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.62 Å Release Date: 2026-01-14 Classification: MEMBRANE PROTEIN Ligands: 91Q, CLR |
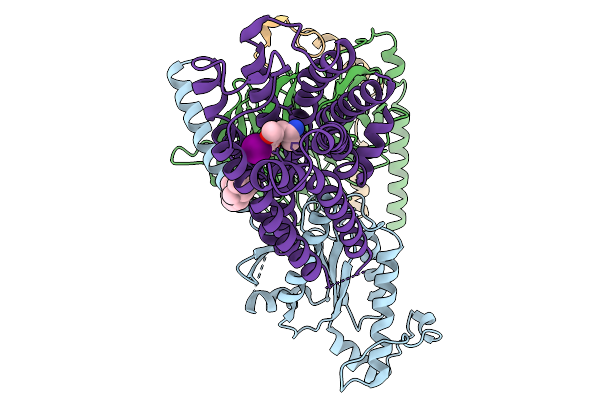 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.76 Å Release Date: 2026-01-14 Classification: MEMBRANE PROTEIN Ligands: A1EK5, CLR |
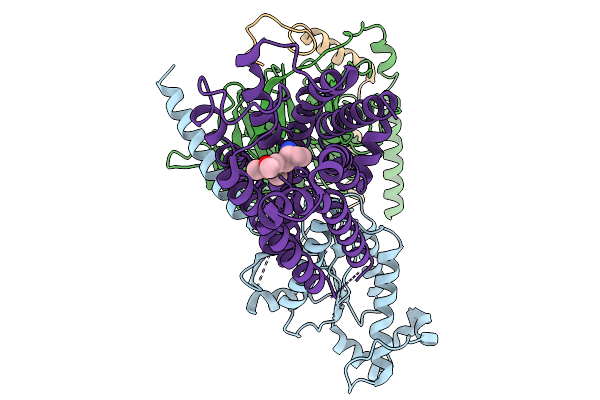 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.27 Å Release Date: 2026-01-14 Classification: MEMBRANE PROTEIN Ligands: A1EK8 |
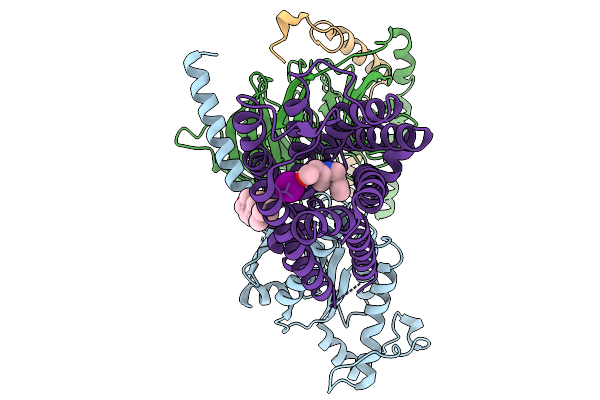 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.98 Å Release Date: 2026-01-14 Classification: MEMBRANE PROTEIN Ligands: CLR, A1ELA |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-12-17 Classification: DE NOVO PROTEIN Ligands: 308 |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-12-17 Classification: DE NOVO PROTEIN Ligands: UNX, 308 |
 |
Crystal Structure Of De Novo Designed Amantadine Induced Heterodimer Daid23.4
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2025-12-17 Classification: DE NOVO PROTEIN Ligands: 308 |

