Search Count: 1,700
All
Selected
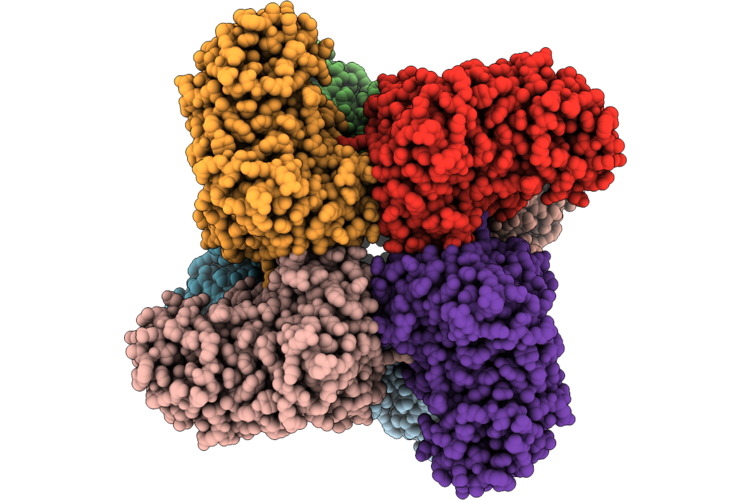 |
Cryo-Electron Microscopic Structure Of A Novel Amidohydrolase Adh3 Triple Mutation
Organism: Stenotrophomonas sp. cw117
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: HYDROLASE Ligands: ZN, 97U |
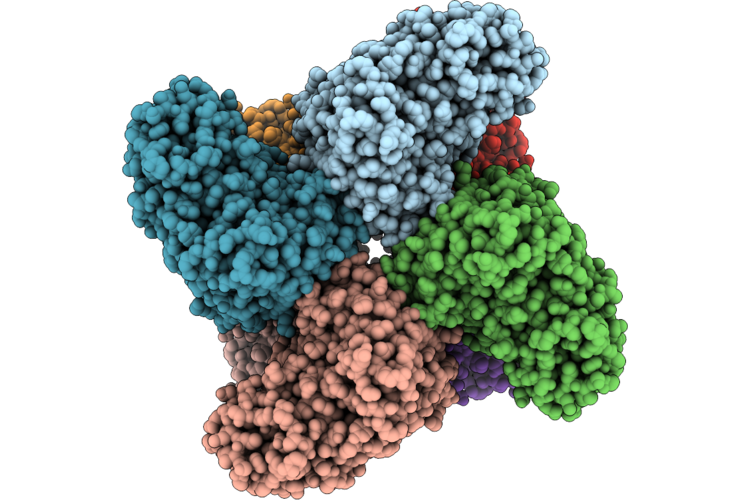 |
Cryo-Em Structure And Rational Engineering Of A Novel Efficient Ochratoxin A-Detoxifying Amidohydrolase
Organism: Pseudoxanthomonas wuyuanensis
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: HYDROLASE Ligands: ZN, 97U |
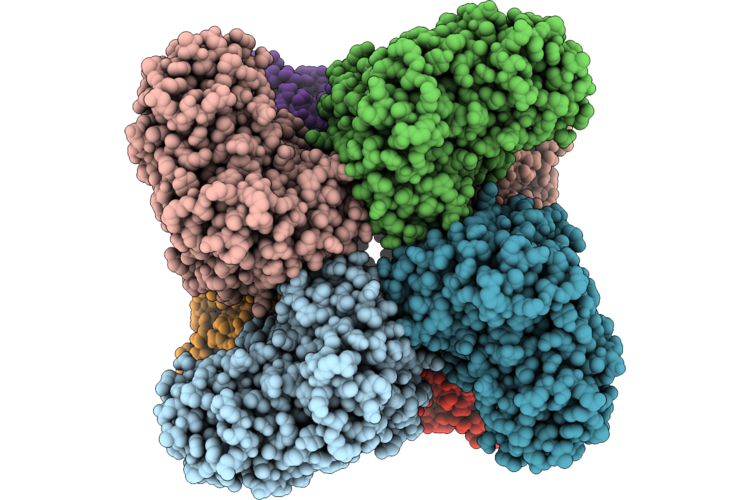 |
Cryo-Electron Microscopic Structure Of A Novel Amidohydrolase With Three Mutations
Organism: Novilysobacter luteus
Method: ELECTRON MICROSCOPY Resolution:2.69 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: ZN, 97U |
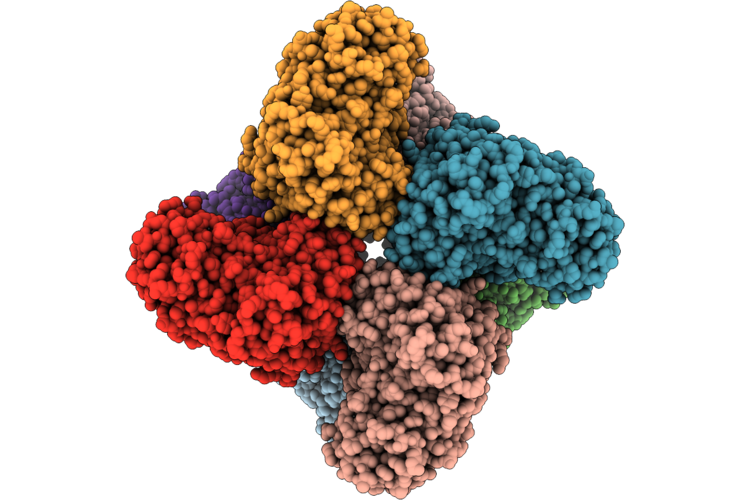 |
Cryo-Electron Microscopic Structure Of A Highly Efficient Ochratoxin Detoxification Enzyme Lladh
Organism: Novilysobacter luteus
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: ZN |
 |
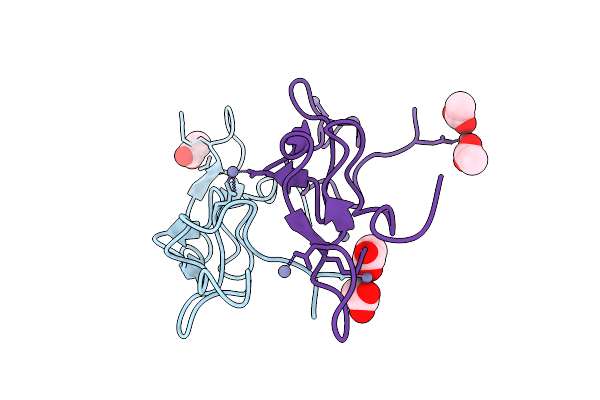
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: A1ES7 |
 |
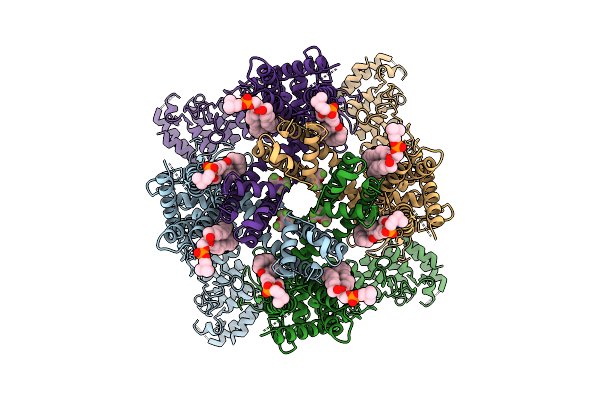
Organism: Protobothrops mucrosquamatus
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-04-08 Classification: BLOOD CLOTTING Ligands: ZN, ACT |
 |
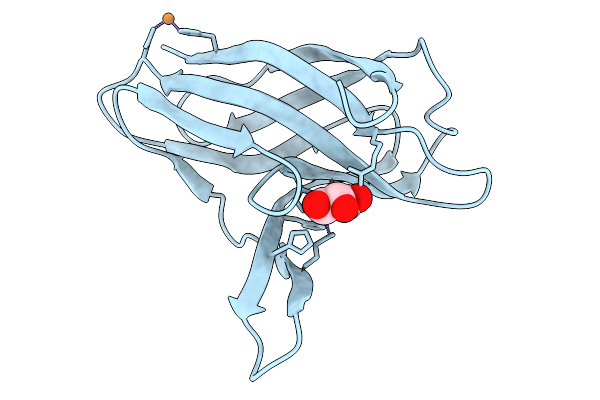
Cryo-Em Structure Of Human Trpv3 In Complex With Sevoflurane Determined In Msp2N2 Nanodisc
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.29 Å Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN Ligands: A1IPI, POV |
 |
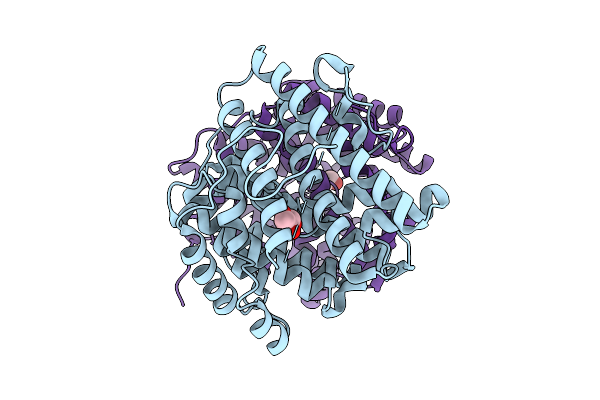
Organism: Neisseria meningitidis serogroup a
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2026-03-25 Classification: METAL BINDING PROTEIN Ligands: CU, GOL |
 |
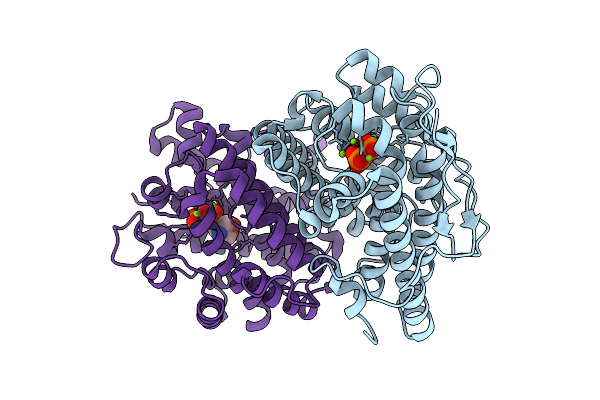
Crystal Structure Of Terpeniod Cyclase Spsods From Rhizobacterium Serratia Plymuthica
Organism: Serratia plymuthica 4rx13
Method: X-RAY DIFFRACTION Resolution:2.22 Å Release Date: 2026-03-18 Classification: LYASE Ligands: GOL |
 |
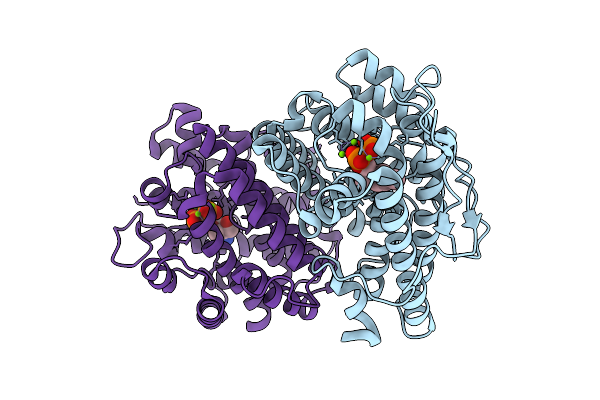
Organism: Serratia plymuthica 4rx13
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2026-03-18 Classification: LYASE Ligands: POP, MG, TRS |
 |
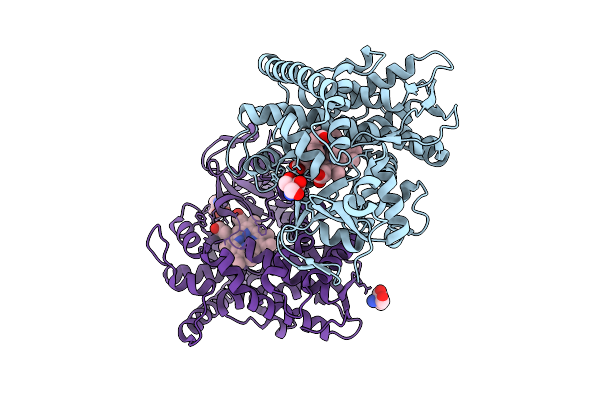
Organism: Serratia plymuthica 4rx13
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-03-18 Classification: LYASE Ligands: GPP, MG, TRS, POP |
 |
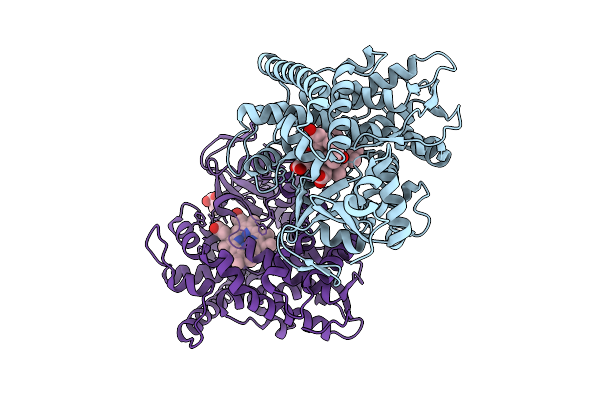
Organism: Priestia megaterium (strain nbrc 15308)
Method: X-RAY DIFFRACTION Resolution:1.57 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: HEM, IMD, PEG, TRS, CL |
 |
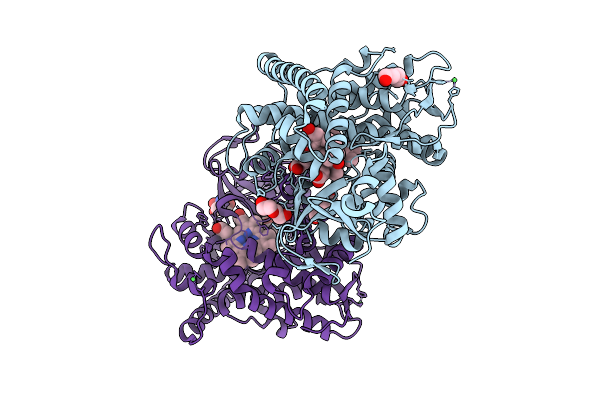
Crystal Structure Of A P450 Bm3 Heme Domain Mutant In Complex With Zearalenone
Organism: Priestia megaterium (strain nbrc 15308)
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: HEM, ZER, TRS |
 |
Crystal Structure Of A P450 Bm3 Heme Domain Mutant In Complex With Alpha-Zearalanol
Organism: Priestia megaterium (strain nbrc 15308)
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: HEM, 36J, PEG, NI, TRS |
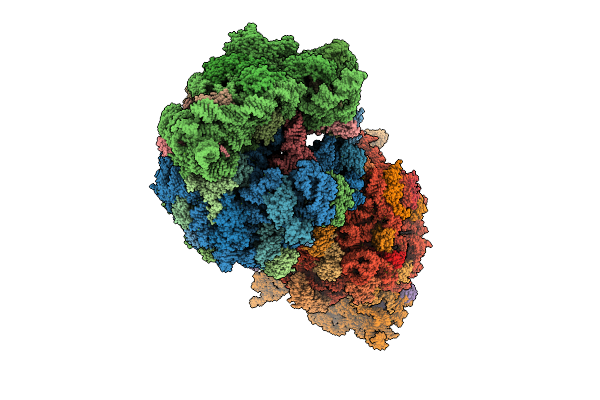 |
Crystal Structure Of The Wild-Type Thermus Thermophilus 70S Ribosome In Complex With Benzoxaborole Derivative Of Azithromycin (Azi-Bb2), Mrna, Aminoacylated A-Site Phe-Trnaphe, Aminoacylated P-Site Fmet-Trnamet, And Deacylated E-Site Trnaphe At 2.45A Resolution
Organism: Escherichia coli, Escherichia phage t4, Thermus thermophilus hb8
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2026-03-11 Classification: RIBOSOME Ligands: MG, K, A1C9N, ZN, SF4 |
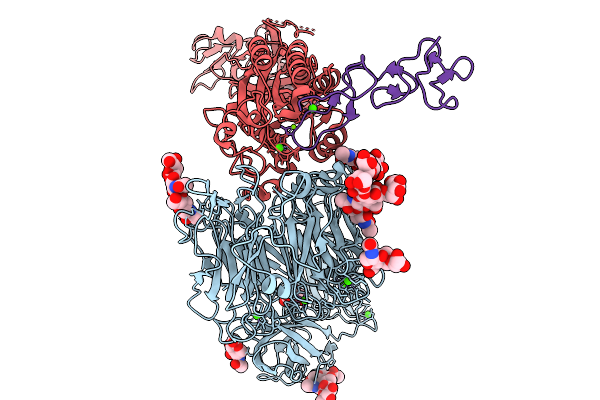 |
Organism: Homo sapiens, Protobothrops mucrosquamatus
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: CA, NAG, MG |
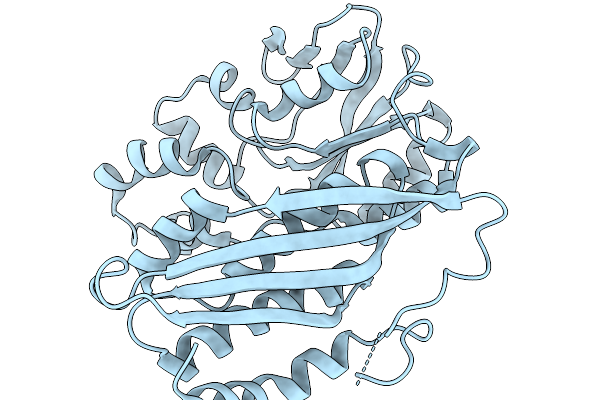 |
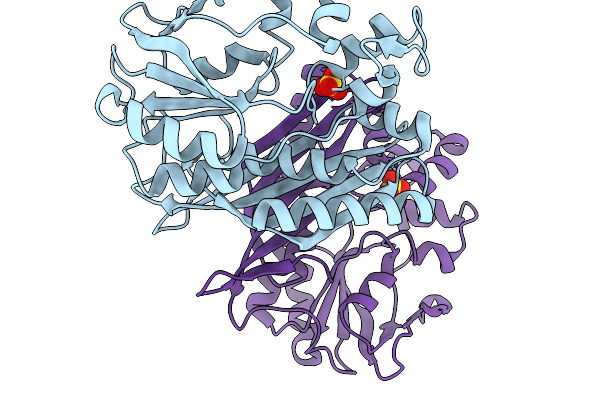
Crystal Structure Of Chimeric Air Synthetase Constructed From E. Coli (Residues 1-168) And Pyrococcus Abyssi (Residues 163-334)
Organism: Escherichia coli, Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-02-11 Classification: LIGASE |
 |
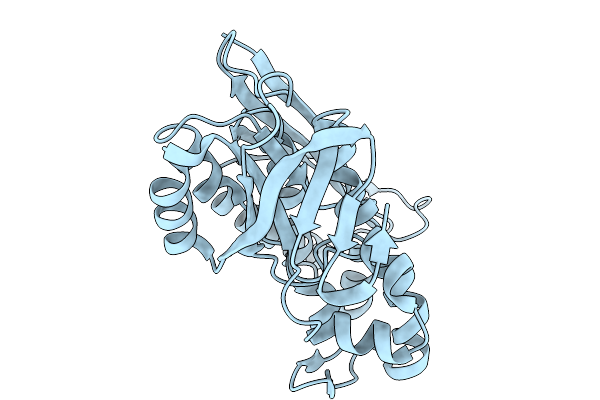
Crystal Structure Of Pyrococcus Abyssi Air Synthetase With An N-Terminal 43-Residues Deletion
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-02-11 Classification: LIGASE Ligands: SO4 |
 |
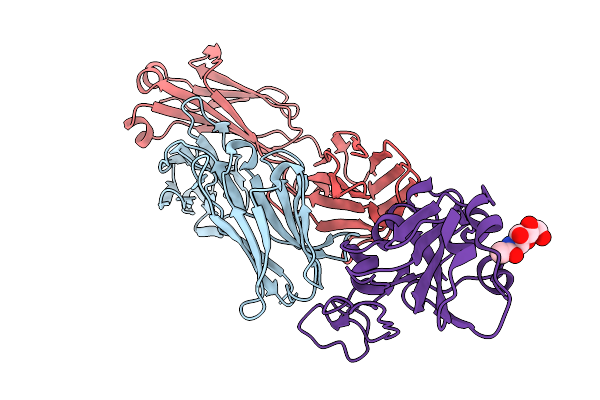
Crystal Structure Of Pyrococcus Abyssi Air Synthetase With An N-Terminal 54-Residues Deletion
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-02-11 Classification: LIGASE |
 |
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.39 Å Release Date: 2026-01-07 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |

