Search Count: 14
All
Selected
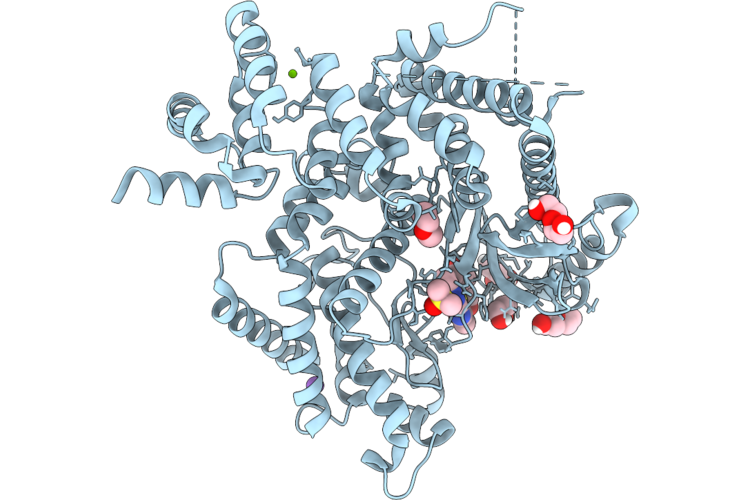 |
The Structure Of Human Vacuolar Protein Sorting 34 Catalytic Domain Bound To Rd-Ii-83
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: A1C15, GOL, PEG, DMS, MG, CL, K |
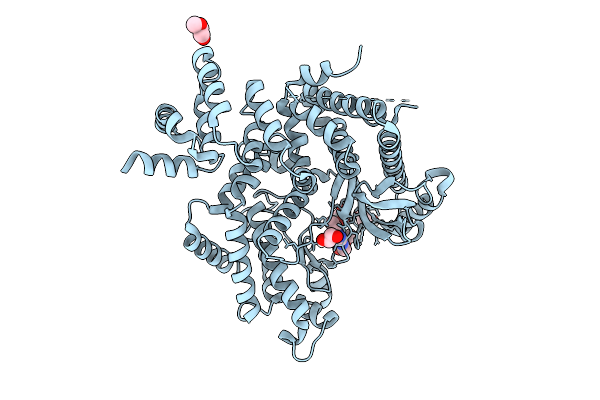 |
The Structure Of Human Vacuolar Protein Sorting 34 Catalytic Domain Bound To Rd-I-137
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.06 Å Release Date: 2026-04-15 Classification: TRANSFERASE Ligands: A1CD6, GOL, PEG, CL |
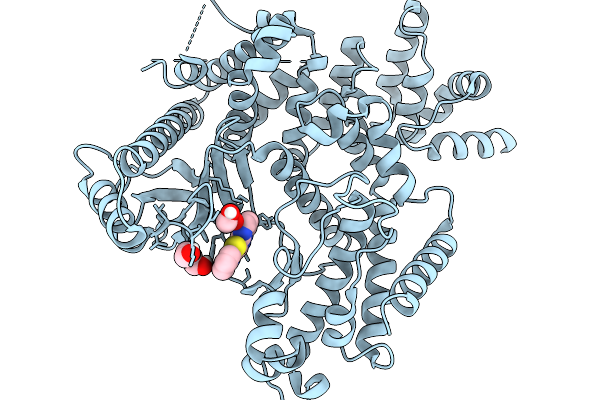 |
The Structure Of Human Vacuolar Protein Sorting 34 Catalytic Domain Bound To Rd-Ii-123
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.41 Å Release Date: 2026-03-25 Classification: TRANSFERASE Ligands: A1DAY, EDO, PEG, CL |
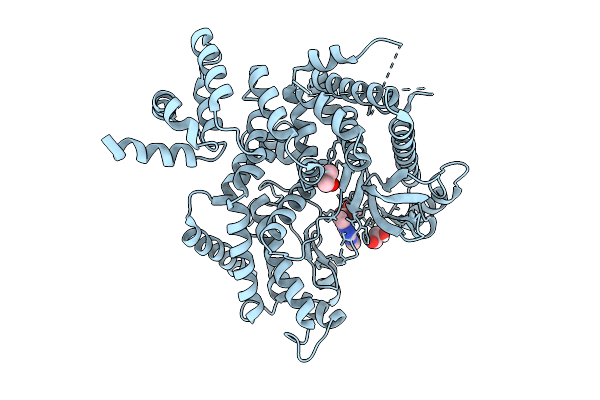 |
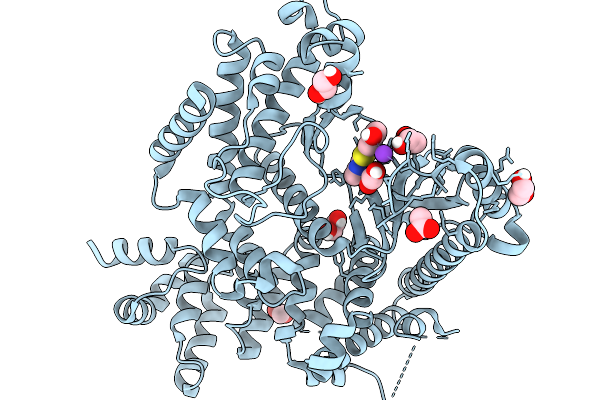
The Structure Of Human Vacuolar Protein Sorting 34 Catalytic Domain Bound To Rd-Ii-81
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.69 Å Release Date: 2026-03-18 Classification: TRANSFERASE Ligands: GOL, A1C9V, DMS |
 |
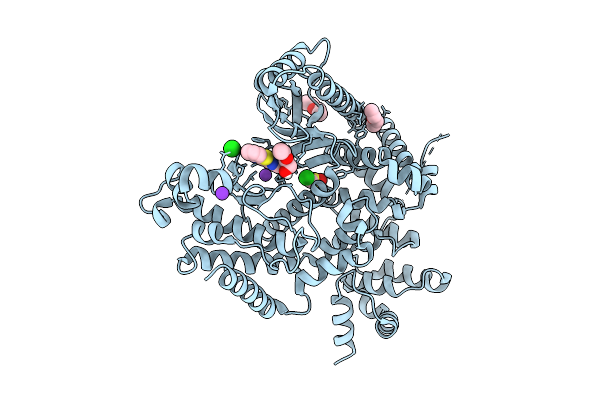
The Structure Of Human Vacuolar Protein Sorting 34 Catalytic Domain Bound To Rd-I-86
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2026-03-04 Classification: TRANSFERASE Ligands: A1BYW, ACT, EDO, PEG, CL, K |
 |
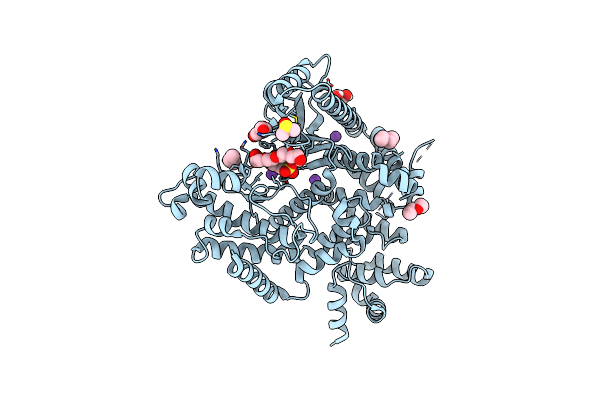
The Structure Of Human Vacuolar Protein Sorting 34 Catalytic Domain Bound To Rd-I-53
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.16 Å Release Date: 2025-09-17 Classification: TRANSFERASE Ligands: A1A6F, ETX, ETZ, DMS, K, CL |
 |
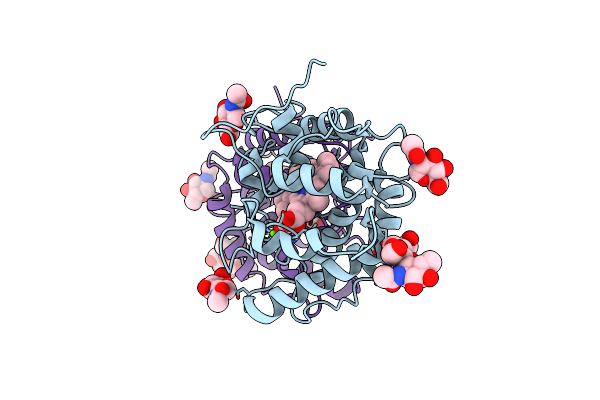
The Structure Of Human Vacuolar Protein Sorting 34 Catalytic Domain Bound To Mes
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.36 Å Release Date: 2024-11-06 Classification: TRANSFERASE Ligands: MES, PG4, DMS, PEG, GOL, CL, K |
 |
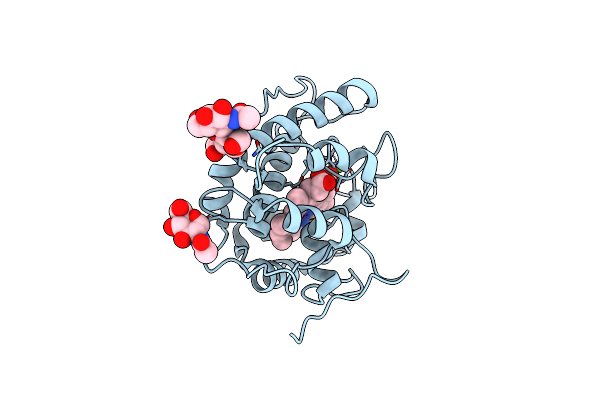
Artificial Unspecific Peroxygenase Expressed In Pichia Pastoris At 2.01 Angstrom Resolution
Organism: Marasmius rotula
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2023-03-01 Classification: OXIDOREDUCTASE Ligands: HEM, NAG, MG, CL |
 |
Artificial Unspecific Peroxygenase Expressed In Pichia Pastoris At 1.21 Angstrom Resolution
Organism: Marasmius rotula
Method: X-RAY DIFFRACTION Resolution:1.21 Å Release Date: 2023-03-01 Classification: OXIDOREDUCTASE Ligands: HEM, NAG, MG |
 |
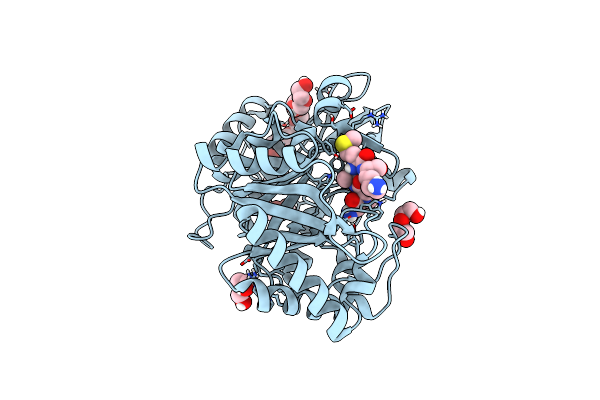
Artificial Unspecific Peroxygenase Expressed In Escherichia Coli At 2.09 Angstrom Resolution
Organism: Marasmius rotula
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2023-03-01 Classification: OXIDOREDUCTASE Ligands: HEM, MG |
 |
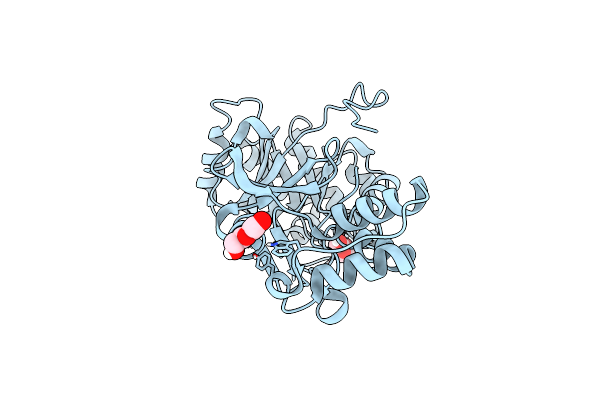
Crystal Structure Of The Chitinase Domain Of The Spore Coat Protein Cote From Clostridium Difficile
Organism: Peptoclostridium difficile (strain 630), Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2020-07-22 Classification: STRUCTURAL PROTEIN Ligands: 1PE, PEG |
 |
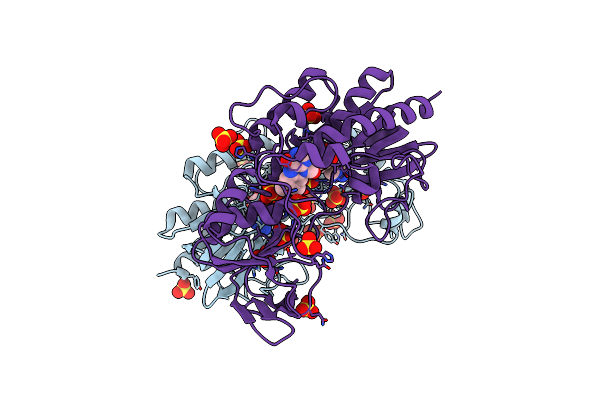
Crystal Structure Of The Chitinase Domain Of The Spore Coat Protein Cote From Clostridium Difficile
Organism: Clostridioides difficile (strain 630)
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2020-06-24 Classification: STRUCTURAL PROTEIN Ligands: PEG |
 |
Organism: Stenotrophomonas maltophilia
Method: X-RAY DIFFRACTION Resolution:2.72 Å Release Date: 2012-04-25 Classification: OXIDOREDUCTASE Ligands: FAD, SO4 |
 |
Organism: Limulus polyphemus
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2000-06-28 Classification: SUGAR BINDING PROTEIN |

