Search Count: 77
All
Selected
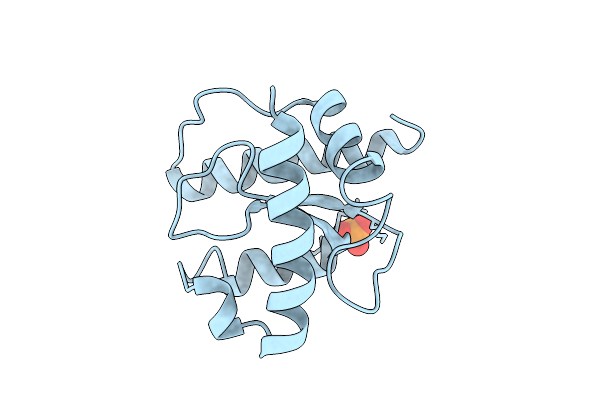 |
X-Ray Structure Of The C-Terminal Domain (Residues 366-485) Of S. Pombe Threonylcarbamoyladenosine Dehydratase At 1.55 A Resolution
Organism: Schizosaccharomyces pombe
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2026-04-01 Classification: RNA BINDING PROTEIN Ligands: PO4 |
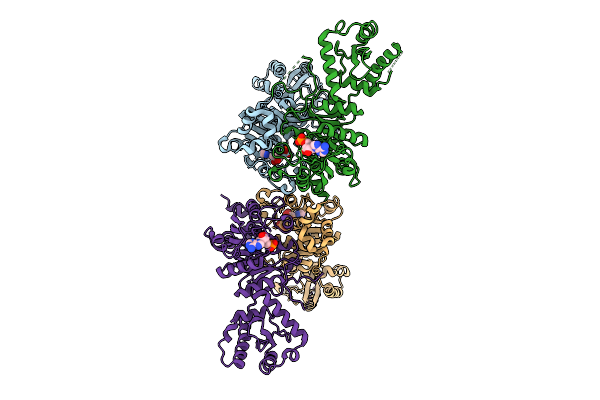 |
X-Ray Structure Of S. Cerevisiae Threonlycarbamoyladenosine Dehydratase 1 (Residues 50-429) In Complex With Amp At 3.7 A Resolution
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:3.70 Å Release Date: 2026-04-01 Classification: RNA BINDING PROTEIN Ligands: AMP, MG |
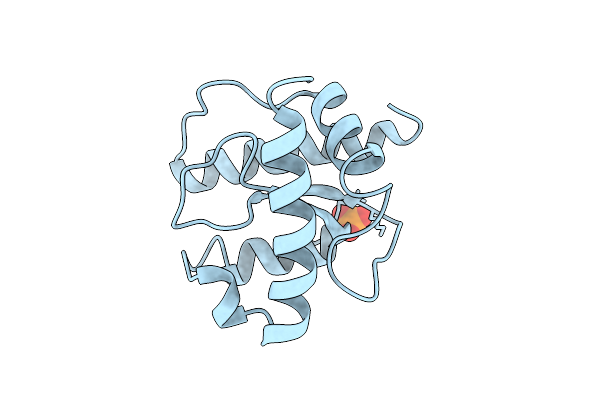 |
X-Ray Structure Of The C-Terminal Domain (Residues 366-485) Of S. Pombe Threonylcarbamoyladenosine Dehydratase
Organism: Schizosaccharomyces pombe
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2025-12-24 Classification: RNA BINDING PROTEIN Ligands: PO4 |
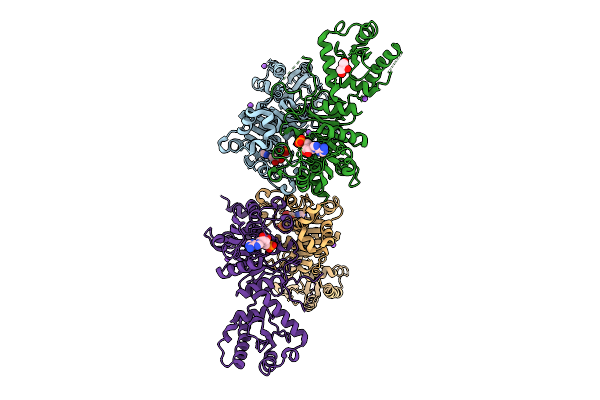 |
X-Ray Structure Of S. Cerevisiae Threonylcarbamoyladenosine Dehydratase 1 (Residues 50-429) In Complex With Amp
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:4.00 Å Release Date: 2025-12-24 Classification: RNA BINDING PROTEIN Ligands: AMP, CL, NA, GOL |
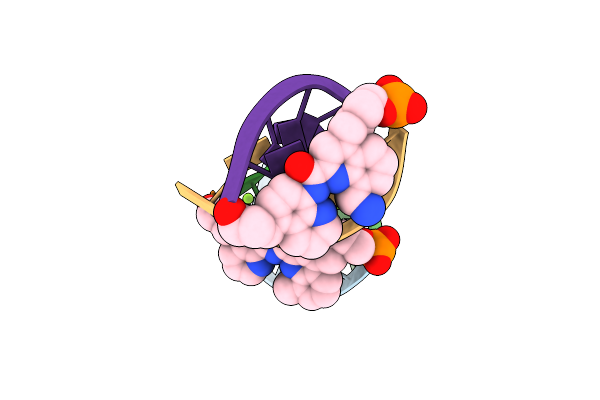 |
Racemic Crystal Structure Of A Synthetic Dna Hairpin With Diquinoline Linker.
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-02-19 Classification: DNA Ligands: JY9, MG |
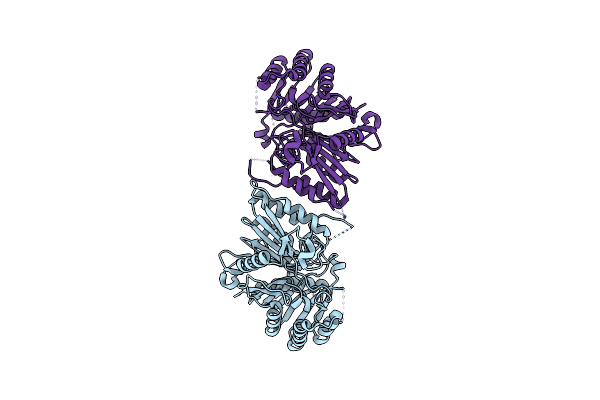 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-05-15 Classification: IMMUNE SYSTEM |
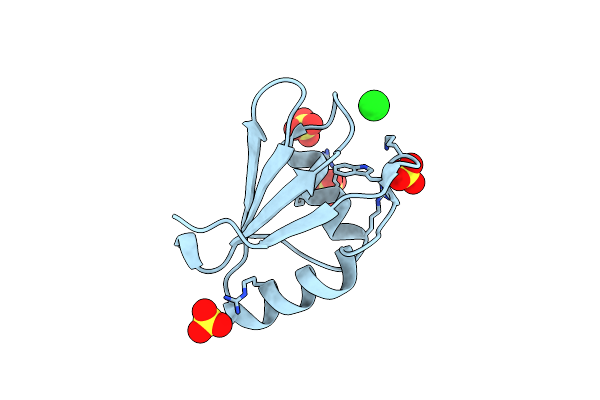 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.02 Å Release Date: 2022-01-19 Classification: TRANSFERASE Ligands: SO4, CL, NA |
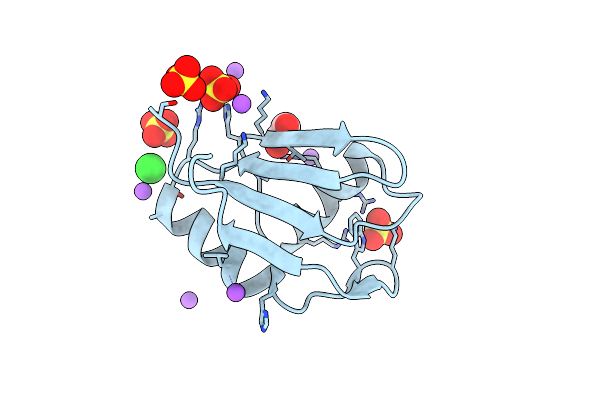 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.23 Å Release Date: 2022-01-19 Classification: TRANSFERASE Ligands: SO4, NA, EDO, CL |
 |
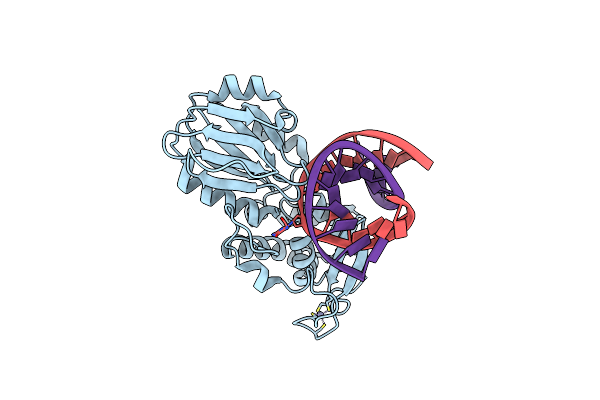
Crystal Structure Of The Complex Between The Lactococcus Lactis Fpg Mutant G226P And A Fapy-Dg Containing Dna
Organism: Lactococcus lactis subsp. cremoris, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2019-07-31 Classification: HYDROLASE Ligands: ZN, GOL |
 |
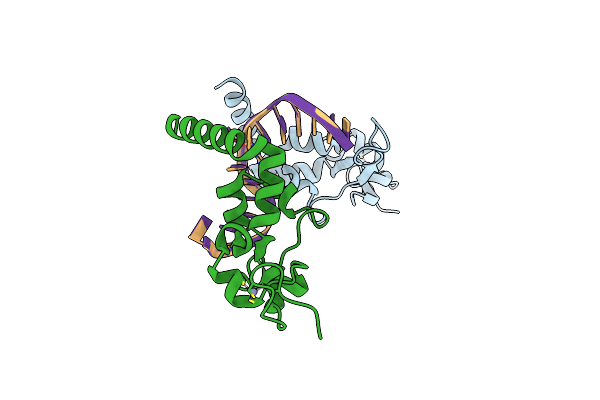
Crystal Structure Of The Complex Between The Lactococcus Lactis Fpg Mutant T221P And A Fapy-Dg Containing Dna
Organism: Lactococcus lactis subsp. cremoris, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2019-02-06 Classification: HYDROLASE Ligands: ZN |
 |
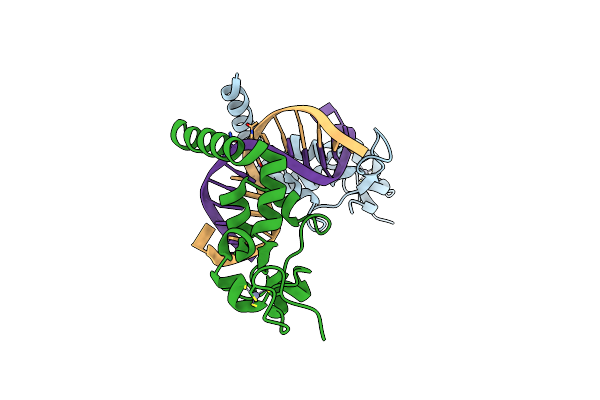
Structure Of The Rad14 Dna-Binding Domain In Complex With C8-Aminofluorene- Guanine Containing Dna
Organism: Saccharomyces cerevisiae s288c, Saccharomyces
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2017-05-10 Classification: DNA BINDING PROTEIN Ligands: ZN |
 |
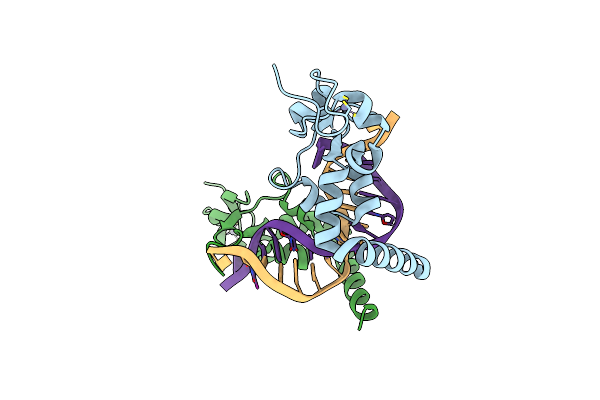
Structure Of The Rad14 Dna-Binding Domain In Complex With N2-Acetylaminonaphtyl- Guanine Containing Dna
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c), Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2017-05-10 Classification: DNA BINDING PROTEIN Ligands: ZN |
 |
Structural Insights Into The Recognition Of Cisplatin And Aaf-Dg Lesions By Rad14 (Xpa)
Organism: Saccharomyces cerevisiae, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2015-07-01 Classification: DNA BINDING PROTEIN Ligands: ZN |
 |
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2015-06-24 Classification: REPLICATION Ligands: ZN, CPT |
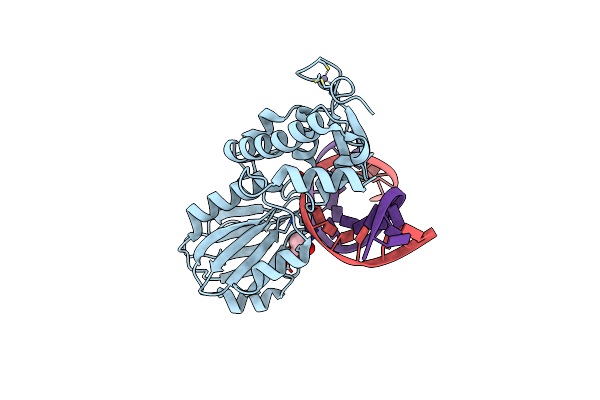 |
Crystal Structure Of A Complex Between R247G Llfpg Mutant And A Thf Containing Dna
Organism: Lactococcus lactis subsp. cremoris, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2015-04-01 Classification: Hydrolase/DNA Ligands: ZN, GOL |
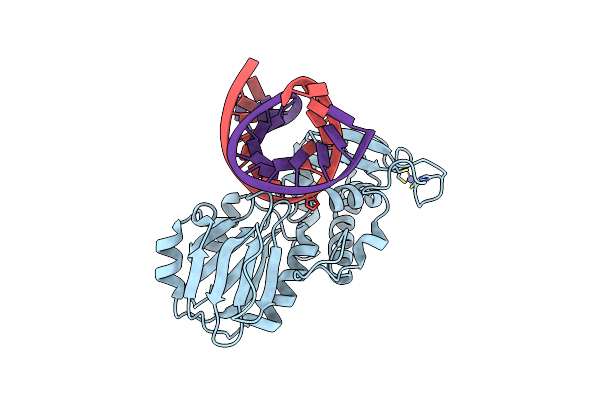 |
Crystal Structure Of A Complex Between A C248Gh Llfpg Mutant And A Thf Containing Dna
Organism: Lactococcus lactis subsp. cremoris, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2015-04-01 Classification: Hydrolase/DNA Ligands: ZN |
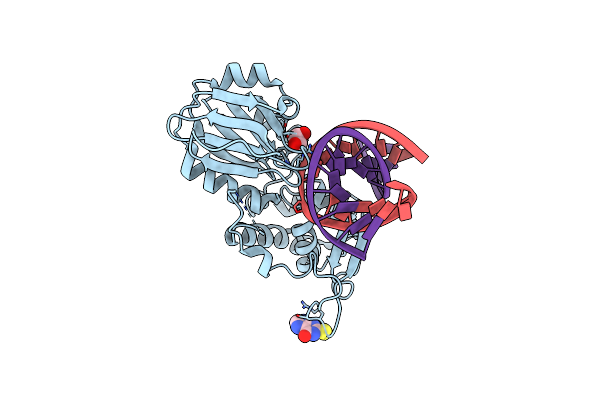 |
Crystal Structure Of A Complex Between An Inhibited Llfpg And A Thf Containing Dna
Organism: Lactococcus lactis subsp. cremoris, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2015-04-01 Classification: Hydrolase/DNA Ligands: 2ON, GOL |
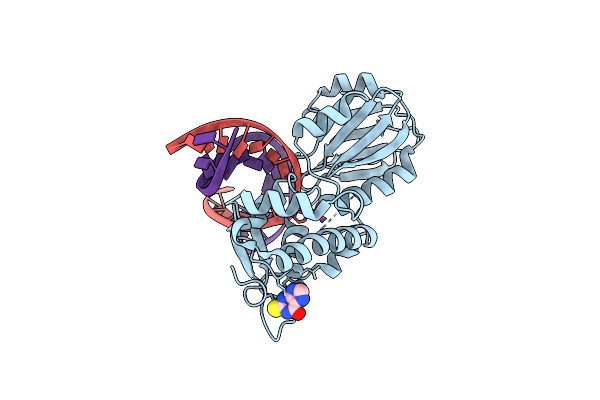 |
Crystal Structure Of A Complex Between An Inhibited Llfpg And A N7-Benzyl-Fapy-Dg Containing Dna
Organism: Lactococcus lactis subsp. cremoris, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2015-04-01 Classification: Hydrolase/DNA Ligands: 2ON |
 |
Organism: Geobacillus stearothermophilus, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2014-12-03 Classification: TRANSFERASE/DNA Ligands: SO4 |
 |
Organism: Geobacillus thermodenitrificans ng80-2
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2014-10-22 Classification: LYASE Ligands: EEM, SO4, EDO, SF4 |

