Search Count: 182
All
Selected
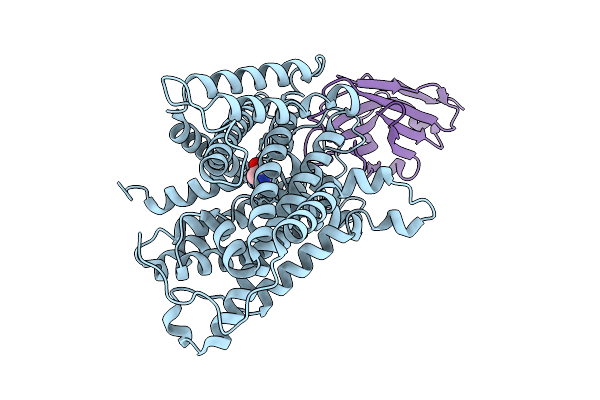 |
Structural Basis Of Specific Lysine Transport By Pseudomonas Aeruginosa Permease Lysp
Organism: Pseudomonas aeruginosa pao1, Lama glama
Method: ELECTRON MICROSCOPY Resolution:3.68 Å Release Date: 2025-10-29 Classification: MEMBRANE PROTEIN Ligands: LYS |
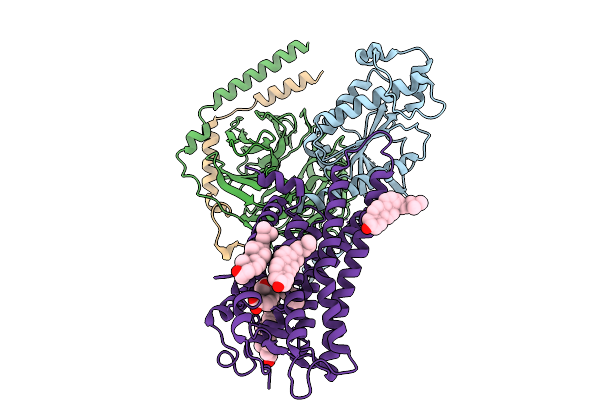 |
Cryo-Em Structure Of Thromboxane A2 Receptor-Minigq Protein Complex Bound To U46619
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-08-27 Classification: MEMBRANE PROTEIN Ligands: PUC, CLR |
 |
Cryo-Em Structure Of Thromboxane A2 Receptor-Minigq Protein Complex Bound To I-Bop
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-08-27 Classification: MEMBRANE PROTEIN Ligands: A1IK0, CLR |
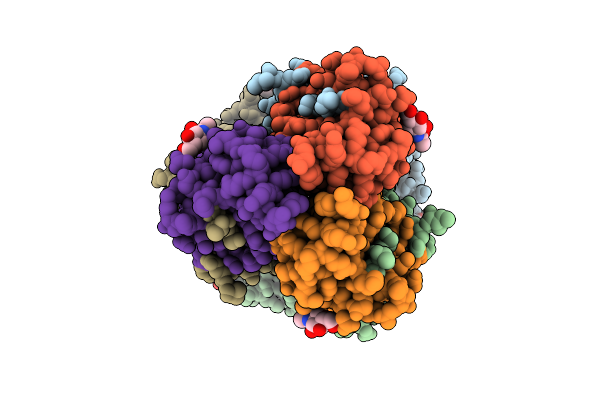 |
Influenza A Virus Group 2 Hemagglutinin (H7, Strain Sh13) In Complex With The Potent Small-Molecule Entry Inhibitor Sa-67
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Release Date: 2025-07-16 Classification: VIRAL PROTEIN/INHIBITOR Ligands: NAG, A1CC2 |
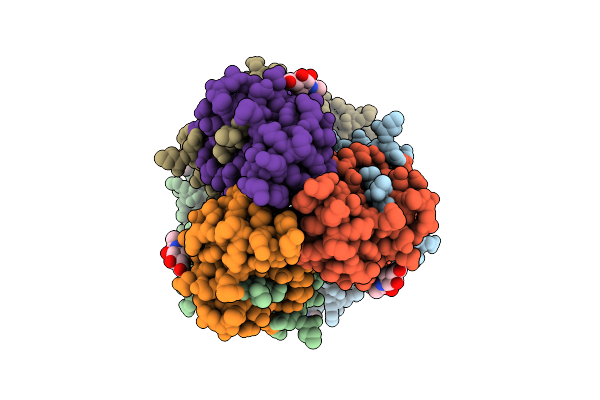 |
Influenza A Virus Group 2 Hemagglutinin (H7, Strain Sh13) In Complex With A Potent Small-Molecule Entry Inhibitor Ing-16-36
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.76 Å Release Date: 2025-07-16 Classification: VIRAL PROTEIN/INHIBITOR Ligands: NAG, A1CC3 |
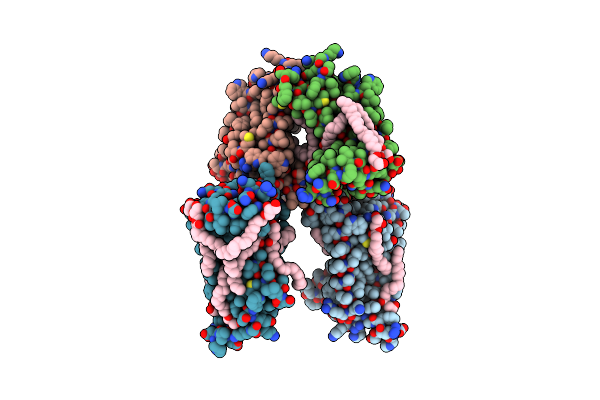 |
Organism: Pseudomonas aeruginosa pao1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2025-03-26 Classification: HYDROLASE Ligands: OLC |
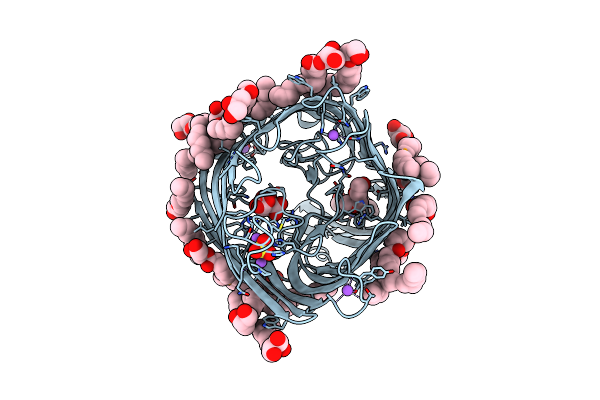 |
Structure Of Alginate Transporter Alge From P. Aeruginosa Pao1 By Using Se-Mag For The The Lipid Cubic Phase Crystallization
Organism: Pseudomonas aeruginosa pao1
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2024-05-15 Classification: TRANSPORT PROTEIN Ligands: FLC, NA, PE5, IRY, C8E, SO4 |
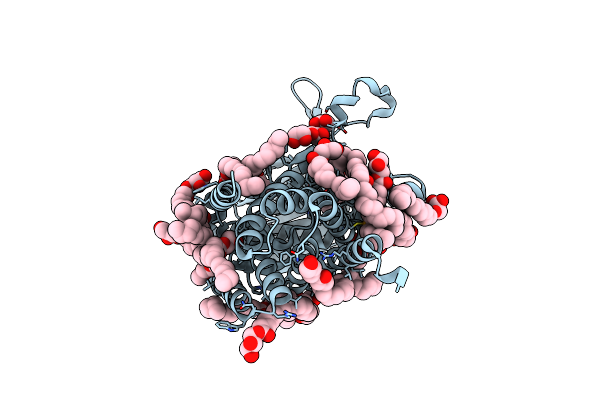 |
Structure Of The Membrane Integral Lipoprotein N-Acyltransferase Lnt From E. Coli By Using Se-Mag For The The Lipid Cubic Phase Crystallization
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2024-05-15 Classification: TRANSFERASE Ligands: IRY, OLC, GOL, PG5 |
 |
In Meso Structure Of The Alginate Exporter, Alge, From Pseudomonas Aeruginosa In 7.10 Monoacylglycerol
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2024-04-03 Classification: MEMBRANE PROTEIN Ligands: CIT, P6G, A1H2K, GOL, A1H52, NA, SO4, CA |
 |
In Meso Structure Of The Adenosine A2A G Protein-Coupled Receptor, A2Ar, In 7.10 Monoacylglycerol
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.37 Å Release Date: 2024-04-03 Classification: MEMBRANE PROTEIN Ligands: ZMA, CLR, A1H2K, A1H52, GOL, NA |
 |
In Meso Structure Of Apolipoprotein N-Acyltransferase, Lnt, From Escherichia Coli In 7.10 Monoacylglycerol
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2024-04-03 Classification: MEMBRANE PROTEIN Ligands: A1H2K, GOL |
 |
In Meso Structure Of The Membrane Integral Lipoprotein N-Acyltransferase Lnt From P. Aeruginosa Covalently Linked With Titc
Organism: Pseudomonas aeruginosa pao1
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2023-07-12 Classification: TRANSFERASE Ligands: OLC, QGT, FLC, GOL |
 |
In Surfo Structure Of The Membrane Integral Lipoprotein N-Acyltransferase Lnt From E. Coli In Complex With Pe
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2023-07-12 Classification: TRANSFERASE Ligands: 6OU, PG5, PG6, TRS, ACT, GOL, LMT, CL |
 |
In Surfo Structure Of The Membrane Integral Lipoprotein N-Acyltransferase Lnt From E. Coli In Complex With Titc And Lyso-Pe
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:2.62 Å Release Date: 2023-07-12 Classification: TRANSFERASE Ligands: PG5, GOL, 6OU, QGT, PG6, TRS, ACT, LMT, CL |
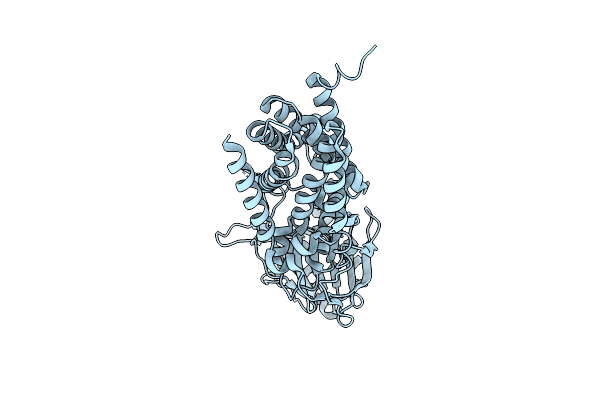 |
Cryo-Em Structure Of Apolipoprotein N-Acyltransferase Lnt From E. Coli (Apo Form)
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2023-07-12 Classification: TRANSFERASE |
 |
Cryo-Em Structure Of Apolipoprotein N-Acyltransferase Lnt From E. Coli In Complex With Pe
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2023-07-12 Classification: TRANSFERASE Ligands: PTY |
 |
Cryo-Em Structure Of Apolipoprotein N-Acyltransferase Lnt From E. Coli In Complex With Pe (C387S Mutant)
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2023-07-12 Classification: TRANSFERASE Ligands: PTY |
 |
Cryo-Em Structure Of Apolipoprotein N-Acyltransferase Lnt From E. Coli In Complex With Lyso-Pe
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2023-07-12 Classification: TRANSFERASE Ligands: OU9 |
 |
Cryo-Em Structure Apolipoprotein N-Acyltransferase Lnt From E.Coli In Complex With Fp3
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2023-07-12 Classification: TRANSFERASE Ligands: OJF |
 |
Cryo-Em Structure Of Apolipoprotein N-Acyltransferase Lnt From E. Coli In Complex With Pam3
Organism: Escherichia coli k-12, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2023-07-12 Classification: TRANSFERASE Ligands: IG7 |

