Search Count: 749
All
Selected
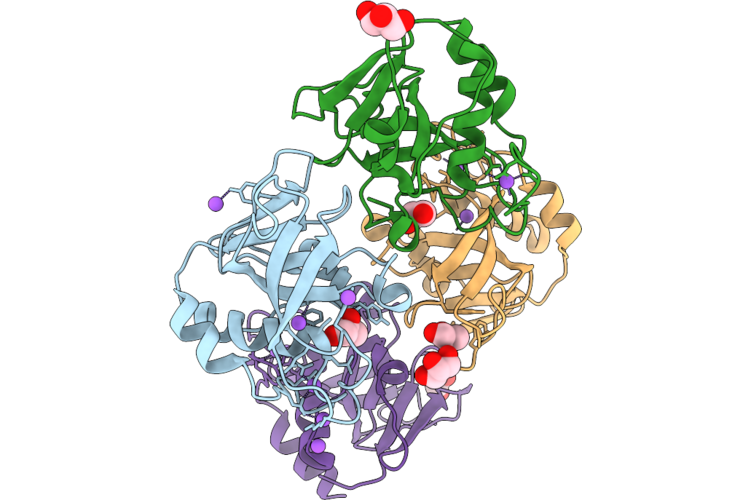 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:1.25 Å Release Date: 2026-05-06 Classification: HYDROLASE |
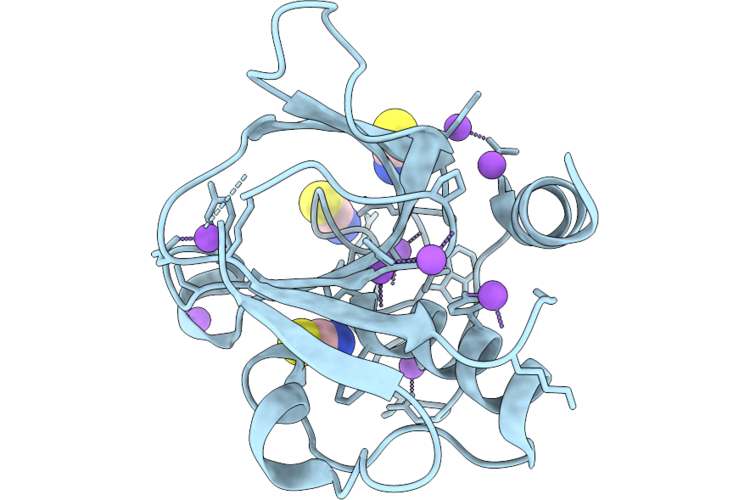 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:1.15 Å Release Date: 2026-05-06 Classification: HYDROLASE |
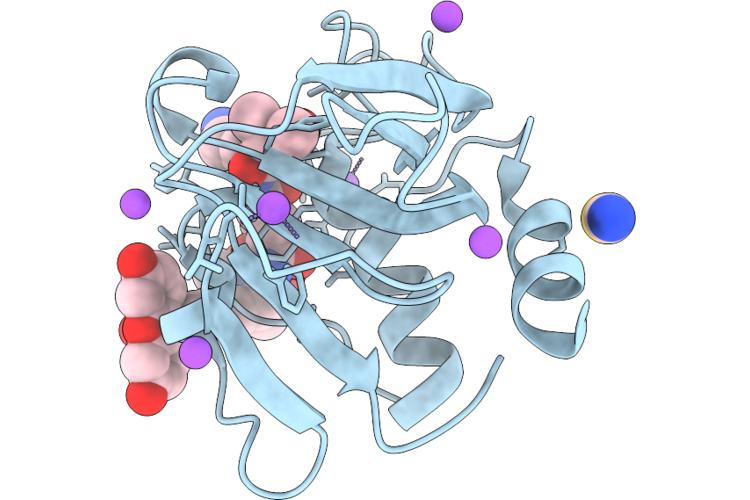 |
Chap Domain Of Staphylococcus Aureus-Specific Lysin L1 Covalently Complexed To Pep1A-Cmk Substrate Mimic
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:1.09 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: A1DEZ, NA, SCN |
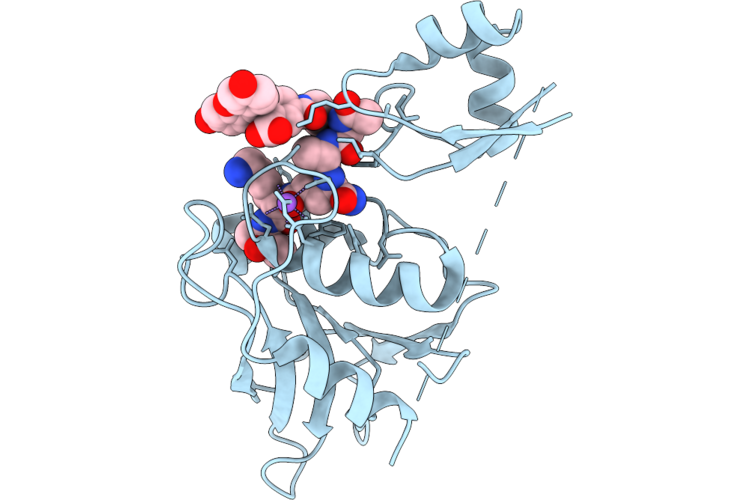 |
Staphylococcus Aureus-Specific Lysin L1-3 (Lysm-Chap) Covalently Complexed To Pep1A-Cmk Substrate Mimic
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.83 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: A1DEZ, NA |
 |
Organism: Bacillus subtilis
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: TOXIN |
 |
Liver Pyruvate Kinase In Complex With A Phthalazine-Based Fluorescent Probe Iv
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: MG, A1JEB, OXL, FBP, PEG |
 |
Structure Of Liver Pyruvate Kinase In Complex With Liver Pyruvate Kinase In Complex With Fluorescent Probe I
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.39 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: FBP, OXL, MG, K, A1JFN |
 |
Structure Of Liver Pyruvate Kinase In Complex With Liver Pyruvate Kinase In Complex With Fluorescent Probe Ii
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.27 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: FBP, OXL, MG, K, A1JFT |
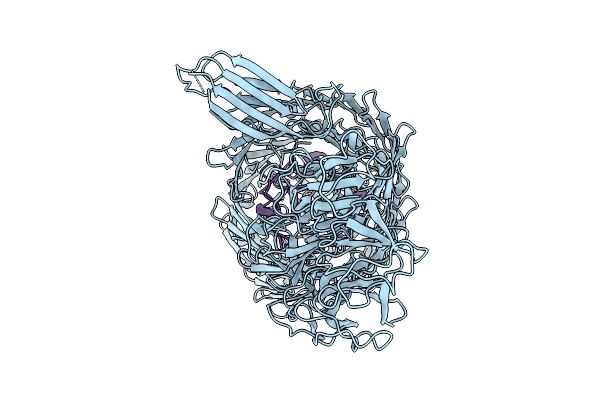 |
Organism: Bacillus inaquosorum
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: CELL ADHESION |
 |
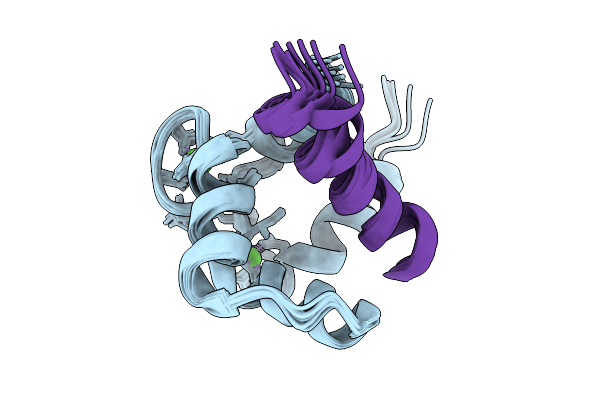
Nmr Structure Of Ca2+/Calmodulin Bound To The Glun1 C0 Domain Of The Nmda Receptor
Organism: Homo sapiens
Method: SOLUTION NMR Release Date: 2026-01-21 Classification: METAL BINDING PROTEIN Ligands: CA |
 |
Nmr Structure Of Ca2+/Calmodulin Bound To The Glun2A C0 Domain Of The Nmda Receptor
Organism: Homo sapiens
Method: SOLUTION NMR Release Date: 2026-01-21 Classification: METAL BINDING PROTEIN Ligands: CA |
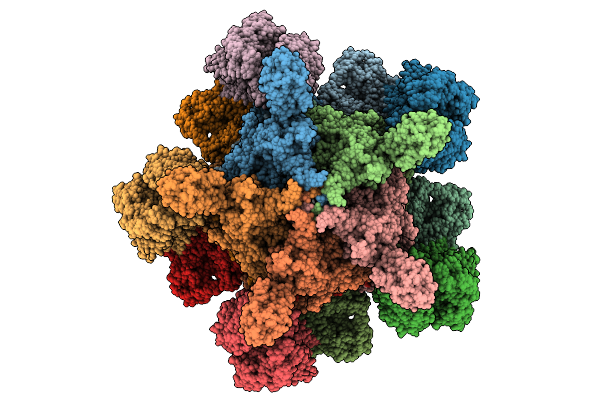 |
Organism: Yersinia entomophaga
Method: ELECTRON MICROSCOPY Release Date: 2025-12-31 Classification: TOXIN |
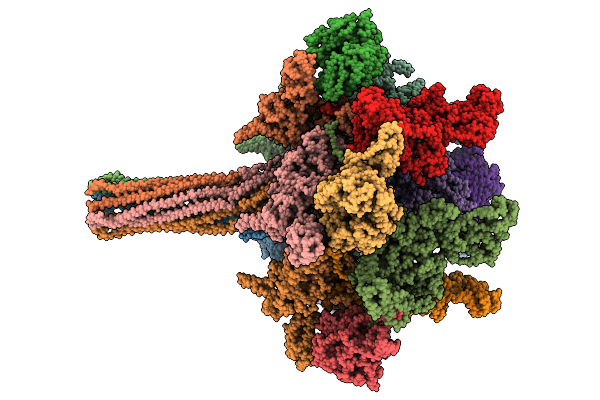 |
Organism: Yersinia entomophaga
Method: ELECTRON MICROSCOPY Release Date: 2025-12-31 Classification: TOXIN |
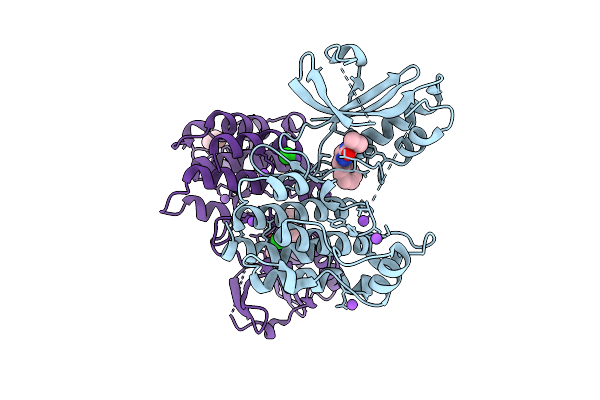 |
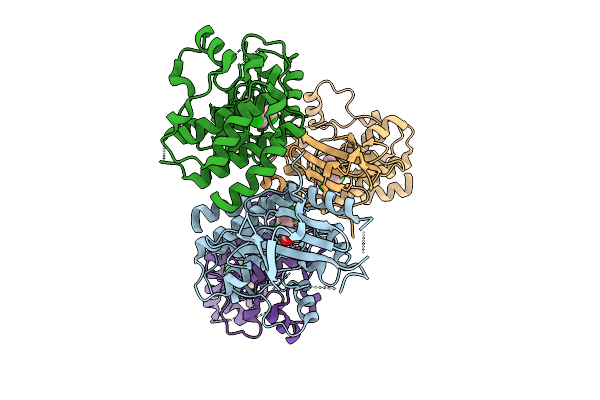
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2025-11-12 Classification: TRANSFERASE/Inhibitor Ligands: A1CNU, BR, CL, NA, TME |
 |
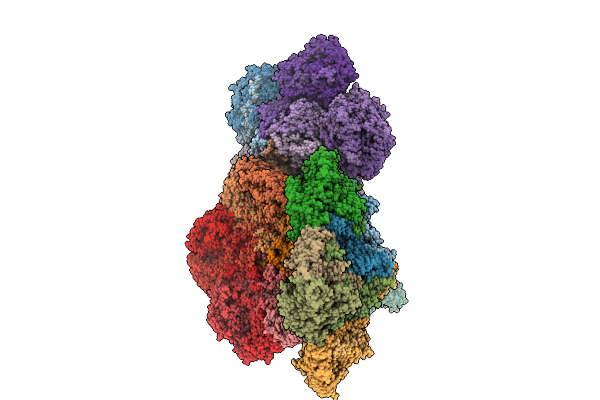
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.49 Å Release Date: 2025-11-12 Classification: Transferase/Inhibitor Ligands: A1CNV |
 |
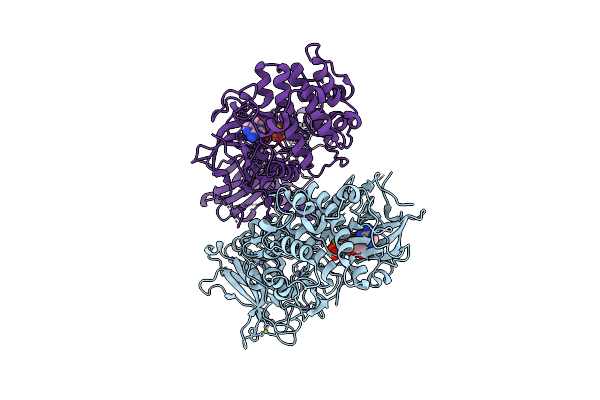
Organism: Coxsackievirus b1
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2025-10-08 Classification: VIRUS LIKE PARTICLE Ligands: MYR |
 |
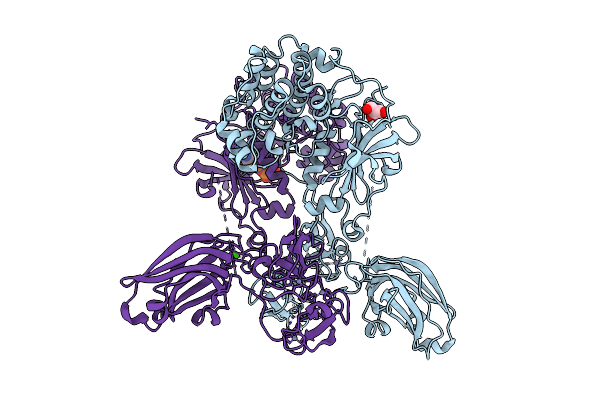
Structure Of Full-Length Human Protein Kinase C Beta 2 (Pkcbii) In The Inactive Conformation
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.32 Å Release Date: 2025-08-20 Classification: SIGNALING PROTEIN Ligands: ZN, ANP, MG |
 |
Structure Of Full-Length Human Protein Kinase C Beta 1 (Pkcbi) In The Active And Inactive Conformation Soaked In Manganese Chloride
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.95 Å Release Date: 2025-08-20 Classification: SIGNALING PROTEIN,TRANSFERASE Ligands: GOL, ZN, ANP, MN, CA |
 |
Structure Of Full-Length Human Protein Kinase C Beta 1 (Pkcbi) In The Active Conformation
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2025-08-20 Classification: SIGNALING PROTEIN,TRANSFERASE Ligands: GOL, ZN |
 |
Structure Of Full-Length Human Protein Kinase C Beta 1 (Pkcbi) In The Active And Inactive Conformation
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.68 Å Release Date: 2025-08-20 Classification: SIGNALING PROTEIN, TRANSFERASE Ligands: ZN, ADN |

