Search Count: 131
All
Selected
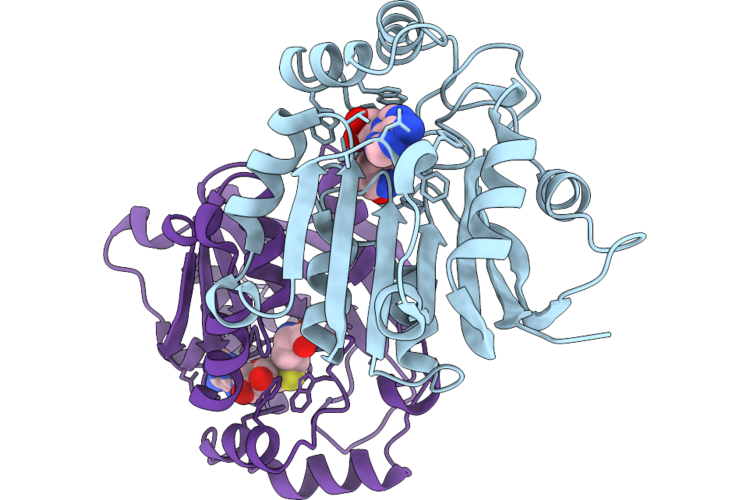 |
Crystal Structure Of Halide Methyltransferase From Moniliophthora Roreri In Complex With Sah
Organism: Moniliophthora roreri
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: SAH |
 |
Crystal Structure Of Halide Methyltransferase From Dichomitus Squalens In Complex With Sah
Organism: Dichomitus squalens
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: SAH, ACT |
 |
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: HYDROLASE |
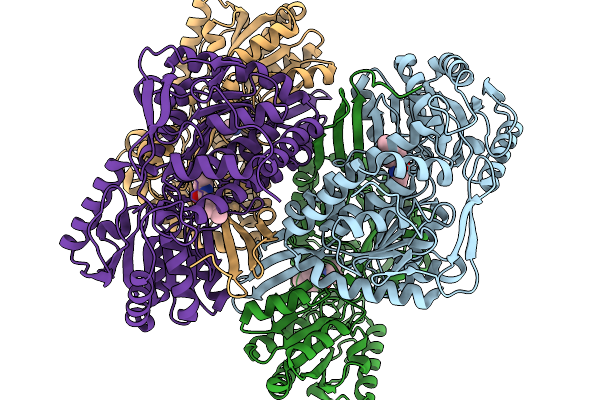 |
Co-Crystal Structure Of Phbdd-L163G With Product N-Isopropyl-4-Methylbenzamide
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: A1EBJ |
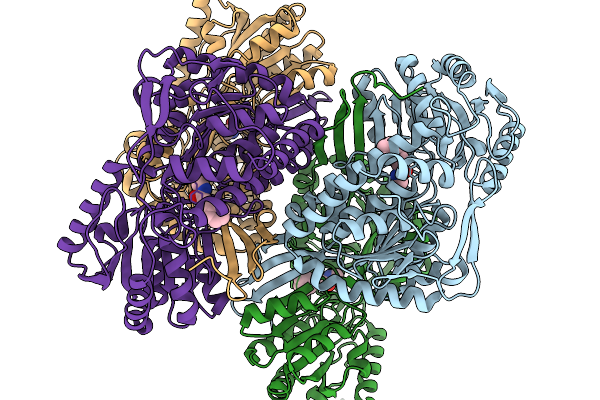 |
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:2.06 Å Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: A1EBQ |
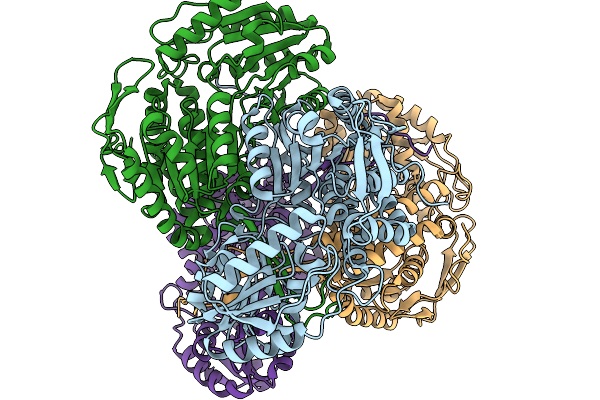 |
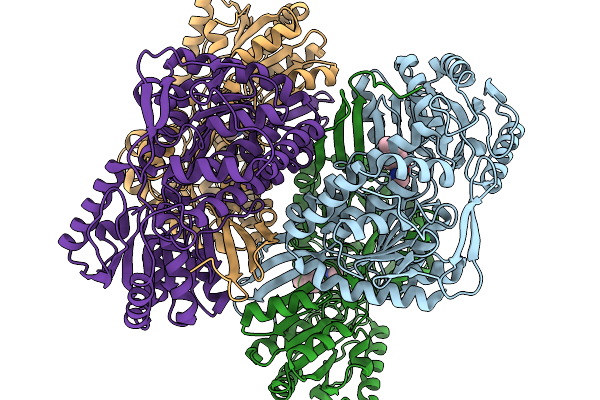
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:1.89 Å Release Date: 2026-01-14 Classification: OXIDOREDUCTASE |
 |
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:2.21 Å Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: A1EC9 |
 |
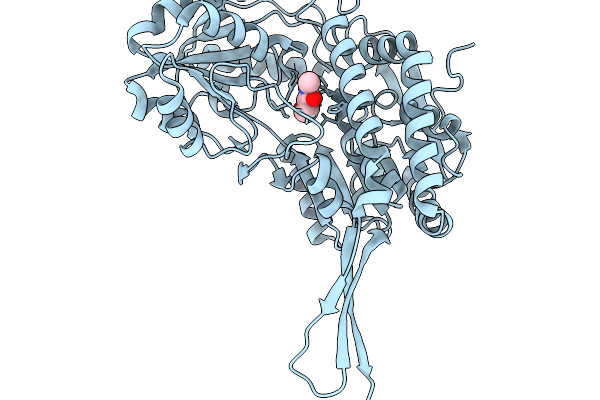
Co-Crystal Structure Of Phbdd-M1-L163G-T234A With N-Cyclopropyl-4-Methylbenzamide
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: A1EDB |
 |
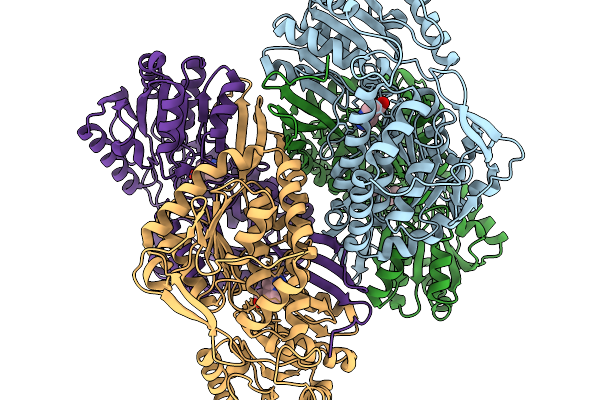
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: A1EDA, SO4 |
 |
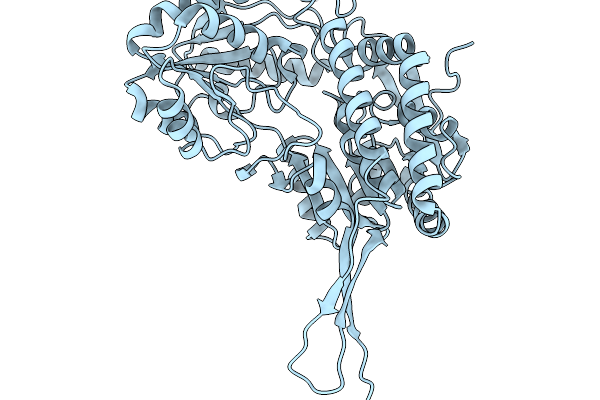
Co-Crystal Structure Of Phbdd-M1 With 1H-Pyrrolo[2,3-B]Pyridine-4-Carbaldehyde
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:2.14 Å Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: A1EDC |
 |
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:3.12 Å Release Date: 2026-01-14 Classification: OXIDOREDUCTASE |
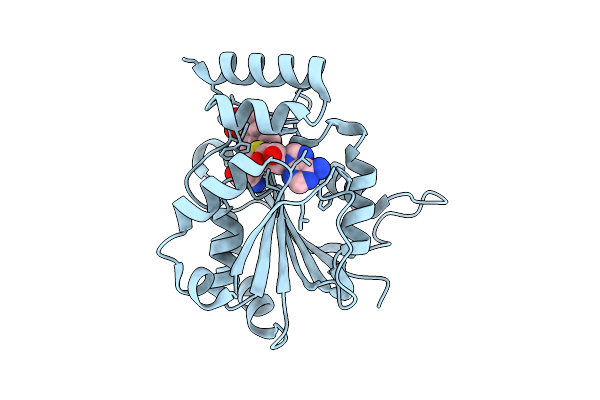 |
Crystal Structure Of Halide Methyltransferase From Tulasnella Calospora In Complex With Sah And Mesitylene Sulfonate
Organism: Tulasnella calospora mut 4182
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-12-24 Classification: TRANSFERASE Ligands: SAH, A1IRH, CL |
 |
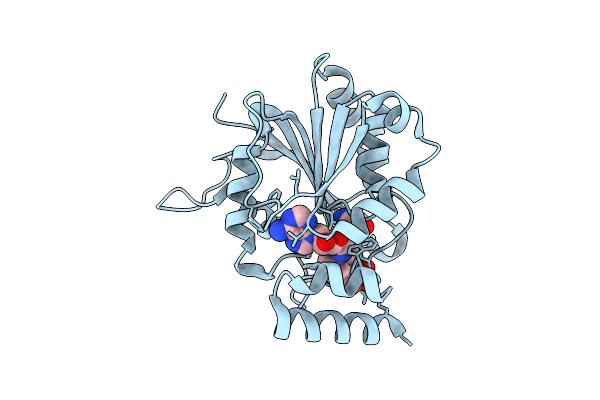
Crystal Structure Of Halide Methyltransferase From Tulasnella Calospora In Complex With Sah
Organism: Tulasnella calospora mut 4182
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-12-24 Classification: TRANSFERASE Ligands: SAH, CL |
 |
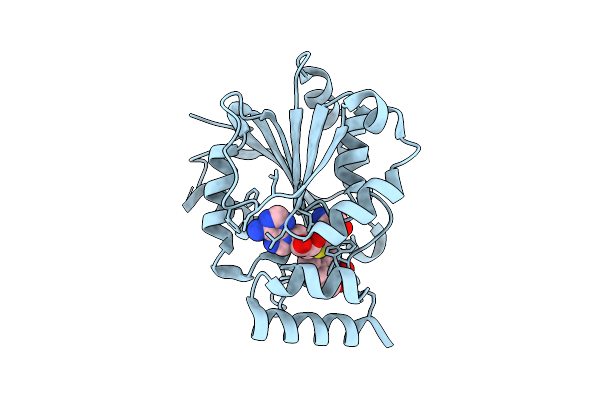
Crystal Structure Of Halide Methyltransferase From Tulasnella Calospora In Complex With Sinefungin And Mesitylene Sulfonate
Organism: Tulasnella calospora mut 4182
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2025-12-24 Classification: TRANSFERASE Ligands: SFG, A1IRH, CL |
 |
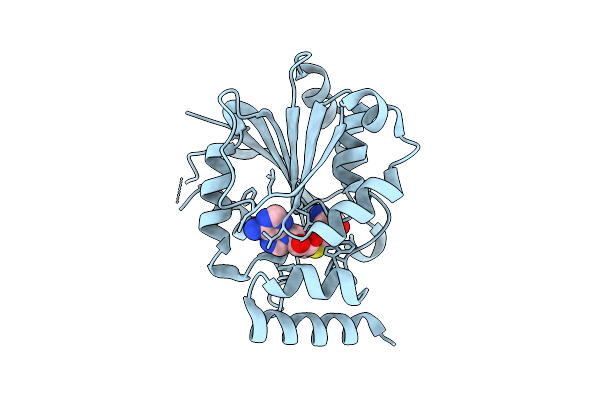
Crystal Structure Of Halide Methyltransferase From Tulasnella Calospora In Complex With Sah And Mesitylene Sulfonate At Room Temperature
Organism: Tulasnella calospora mut 4182
Method: X-RAY DIFFRACTION Resolution:2.41 Å Release Date: 2025-12-24 Classification: TRANSFERASE Ligands: SAH, A1IRH |
 |
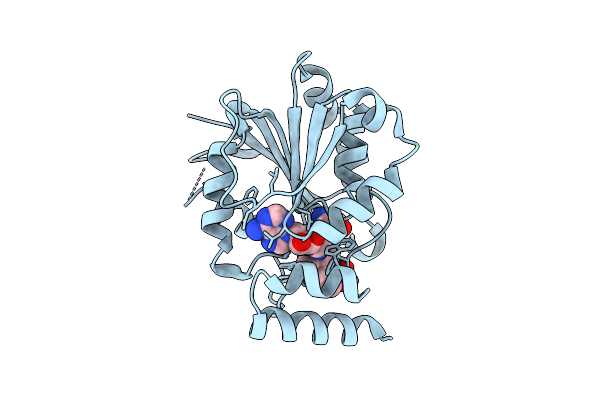
Crystal Structure Of Halide Methyltransferase From Tulasnella Calospora In Complex With Sah At Room Temperature
Organism: Tulasnella calospora mut 4182
Method: X-RAY DIFFRACTION Resolution:2.36 Å Release Date: 2025-12-24 Classification: TRANSFERASE Ligands: SAH |
 |
Crystal Structure Of Halide Methyltransferase From Tulasnella Calospora In Complex With Sinefungin And Mesitylene Sulfonate At Room Temperature
Organism: Tulasnella calospora mut 4182
Method: X-RAY DIFFRACTION Resolution:2.56 Å Release Date: 2025-12-24 Classification: TRANSFERASE Ligands: SFG, A1IRH |
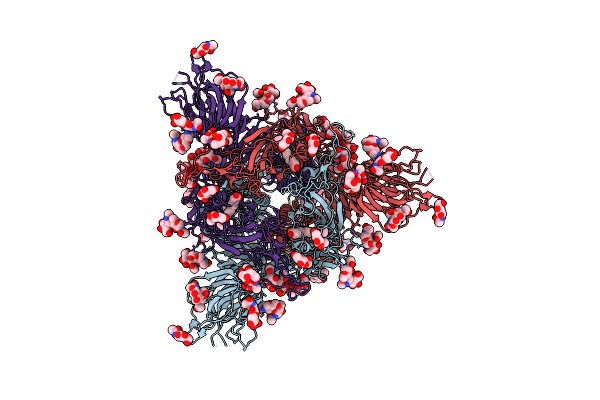 |
Cryo-Em Structure Of The Spike Glycoprotein From Bat Sars-Like Coronavirus (Bat Sl-Cov) Wiv1 In Locked State
Organism: Bat sars-like coronavirus wiv1
Method: ELECTRON MICROSCOPY Release Date: 2025-05-28 Classification: VIRAL PROTEIN Ligands: EIC, NAG |
 |
Organism: Drosophila melanogaster
Method: ELECTRON MICROSCOPY Release Date: 2025-03-05 Classification: VIRUS LIKE PARTICLE |
 |
Cryo-Em Structure Of Single-Layered Nucleoprotein-Rna Helical Assembly From Marburg Virus, Trimeric Repeat Unit
Organism: Marburg virus - musoke, kenya, 1980
Method: ELECTRON MICROSCOPY Release Date: 2024-10-09 Classification: VIRAL PROTEIN |

