Search Count: 27
All
Selected
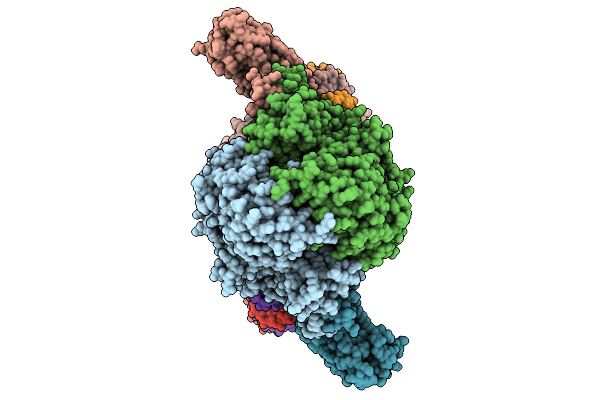 |
Cryoem Structure Of M. Mazei Topoisomerase Vi(A-E342Q)-Minicircle Dna Complex In Cleavage State
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: CA, ANP, K |
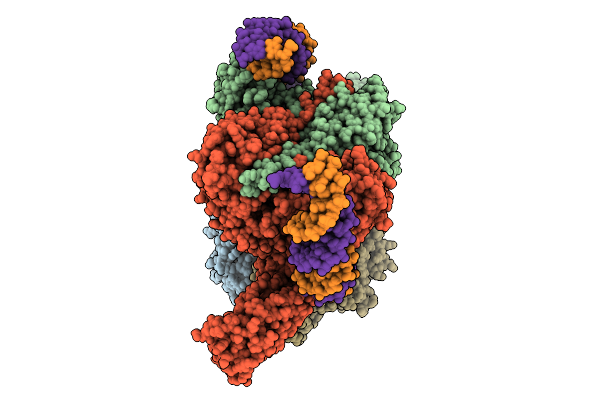 |
Cryoem Structure Of M. Mazei Topoisomerase Vi(A-E342Q)-Minicircle Dna Complex In Asymmetric State
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: ANP, CA, K |
 |
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: ANP, MG, K |
 |
Cryoem Structure Of M. Mazei Topoisomerase Vi-Minicircle Dna Complex In Partially Unfolded Transducer State
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: MG, ANP, K |
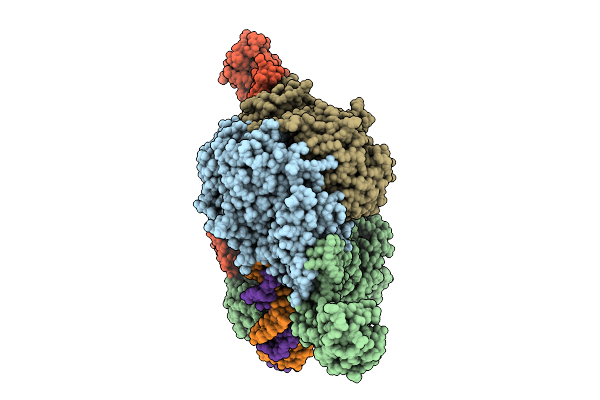 |
Cryoem Structure Of M. Mazei Topoisomerase Vi-Minicircle Dna Complex In Asymmetric State
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: ANP, MG, K |
 |
Organism: Staphylococcus epidermidis
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2023-12-06 Classification: TRANSFERASE Ligands: CL, PO4, UPG, EDO, PG4, PGE |
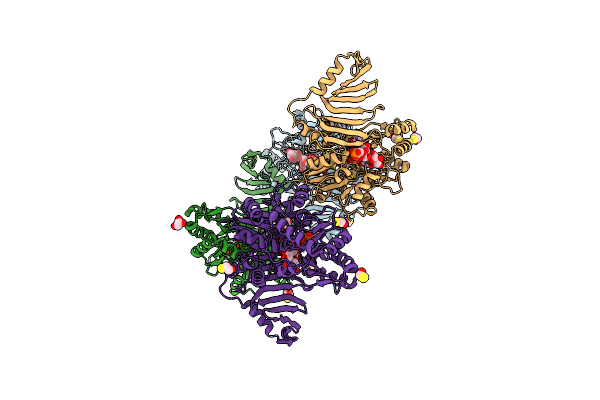 |
Organism: Staphylococcus epidermidis
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2023-12-06 Classification: TRANSFERASE Ligands: BJT, CL, BME, UDP, GOL |
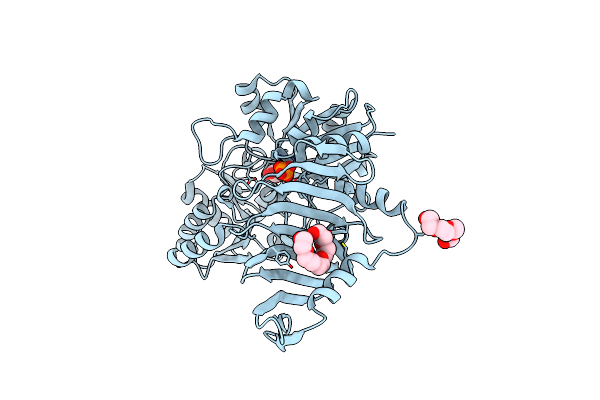 |
Organism: Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Resolution:2.06 Å Release Date: 2023-05-10 Classification: TRANSFERASE Ligands: CL, PO4, GOL, 1PE |
 |
Organism: Staphylococcus epidermidis
Method: X-RAY DIFFRACTION Resolution:2.69 Å Release Date: 2023-05-10 Classification: TRANSFERASE Ligands: CWI, EDO, CL |
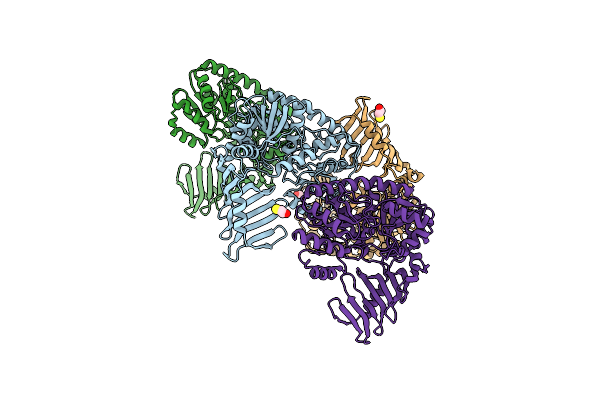 |
Organism: Staphylococcus epidermidis
Method: X-RAY DIFFRACTION Resolution:3.21 Å Release Date: 2023-05-10 Classification: TRANSFERASE Ligands: CL, BME, EDO |
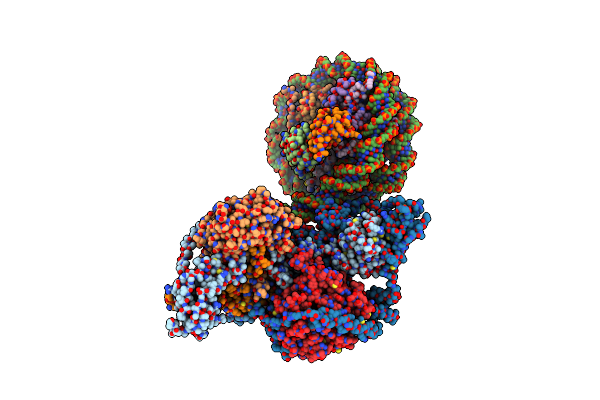 |
Organism: Homo sapiens, Xenopus laevis
Method: ELECTRON MICROSCOPY Release Date: 2021-02-03 Classification: GENE REGULATION/DNA Ligands: MG, SAH, ZN |
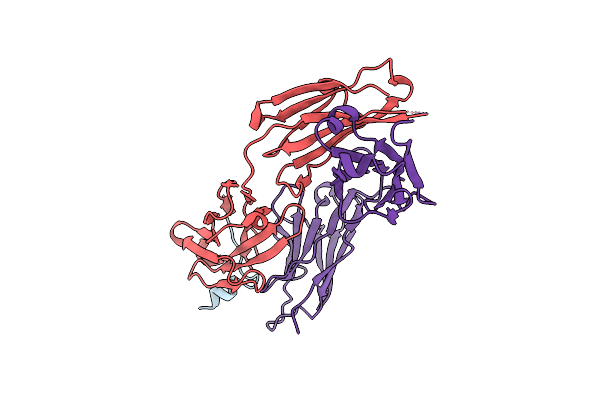 |
Organism: Homo sapiens, Human immunodeficiency virus 1
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2020-02-12 Classification: IMMUNE SYSTEM/VIRAL PROTEIN |
 |
Organism: Homo sapiens, Human immunodeficiency virus 1
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2020-02-12 Classification: IMMUNE SYSTEM/Viral Protein Ligands: CL |
 |
Organism: Homo sapiens, Human immunodeficiency virus 1
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2020-02-12 Classification: IMMUNE SYSTEM/Viral Protein |
 |
Organism: Chaetomium thermophilum (strain dsm 1495 / cbs 144.50 / imi 039719)
Method: ELECTRON MICROSCOPY Release Date: 2019-10-30 Classification: TRANSFERASE Ligands: AGS, MG |
 |
Organism: Chaetomium thermophilum (strain dsm 1495 / cbs 144.50 / imi 039719)
Method: ELECTRON MICROSCOPY Release Date: 2019-10-30 Classification: TRANSFERASE Ligands: AGS, MG |
 |
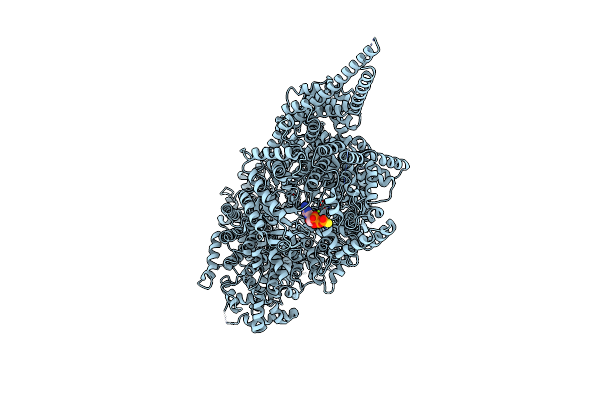
Organism: Chaetomium thermophilum (strain dsm 1495 / cbs 144.50 / imi 039719)
Method: ELECTRON MICROSCOPY Release Date: 2019-10-30 Classification: TRANSFERASE Ligands: AGS, MG |
 |
Organism: Chaetomium thermophilum (strain dsm 1495 / cbs 144.50 / imi 039719)
Method: ELECTRON MICROSCOPY Release Date: 2019-10-30 Classification: TRANSFERASE Ligands: AGS, MG |
 |
Organism: Escherichia coli o157:h7, Escherichia coli o6:k15:h31 (strain 536 / upec)
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2016-07-27 Classification: TOXIN |
 |
Cdia-Ct From Uropathogenic Escherichia Coli In Complex With Cognate Immunity Protein And Cysk
Organism: Escherichia coli o157:h7, Escherichia coli o6:k15:h31 (strain 536 / upec)
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2016-07-27 Classification: TOXIN |

