Search Count: 15
All
Selected
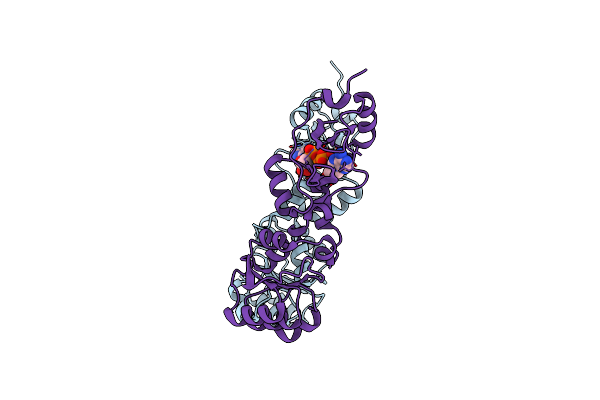 |
Crystal Structure Of The Cbs And Drtgg Domains Of The Regulatory Region Of Clostridium Perfringens Pyrophosphatase Complexed With Activator, Diadenosine Tetraphosphate
Organism: Clostridium perfringens
Method: X-RAY DIFFRACTION Resolution:2.27 Å Release Date: 2010-04-21 Classification: HYDROLASE Ligands: B4P |
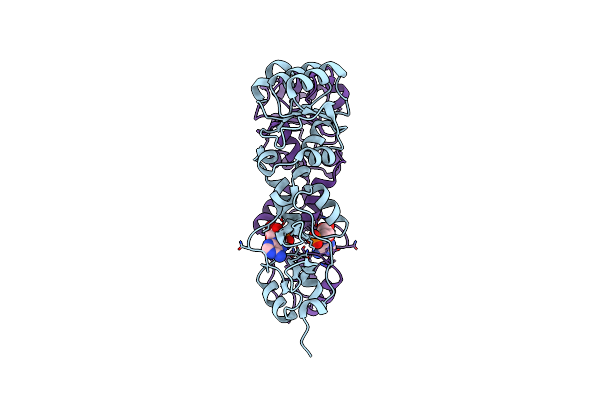 |
Crystal Structure Of The Cbs And Drtgg Domains Of The Regulatory Region Of Clostridium Perfringens Pyrophosphatase Complexed With The Inhibitor, Amp
Organism: Clostridium perfringens
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2010-04-21 Classification: HYDROLASE Ligands: AMP |
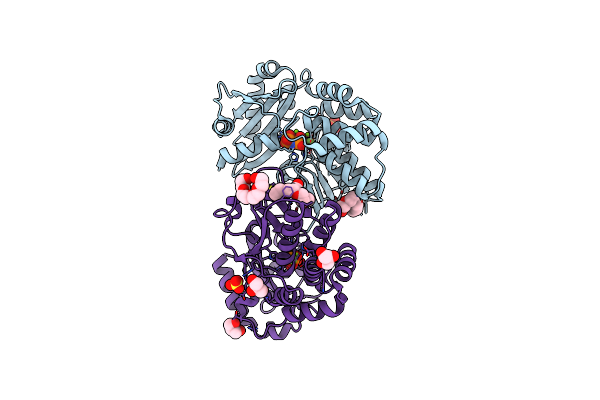 |
Crystal Structure Of Family Ii Inorganic Pyrophosphatase In Complex With Pnp
Organism: Bacillus subtilis
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2006-11-14 Classification: HYDROLASE Ligands: MG, F, SO4, CL, PG4, 2PN, 1PE, GOL |
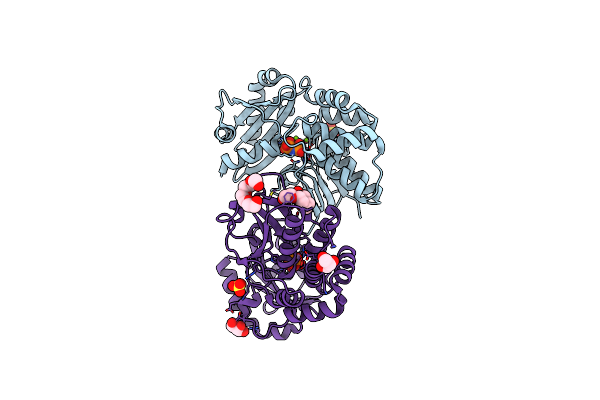 |
Crystal Structure Of Basillus Subtilis Family Ii Inorganic Pyrophosphatase Mutant, H98Q, In Complex With Pnp
Organism: Bacillus subtilis
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2006-11-07 Classification: HYDROLASE Ligands: FE, MG, MN, 2PN, SO4, PG4, GOL |
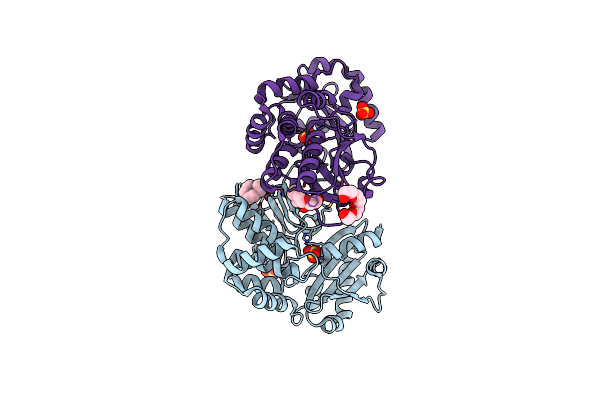 |
Organism: Bacillus subtilis
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2004-11-23 Classification: HYDROLASE Ligands: SO4, PG4 |
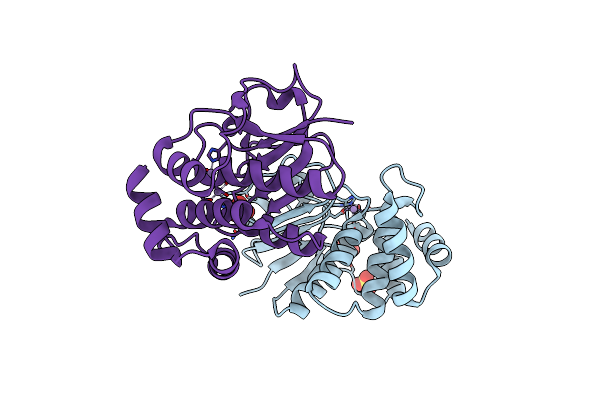 |
Crystal Structure Of The N-Terminal Core Of Bacillus Subtilis Inorganic Pyrophosphatase
Organism: Bacillus subtilis
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2004-11-23 Classification: HYDROLASE Ligands: MN, SO4 |
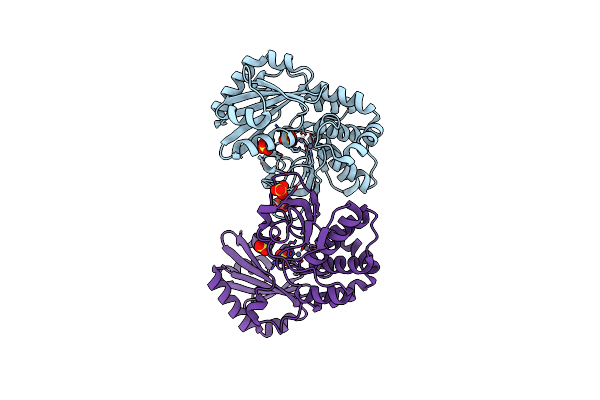 |
Organism: Streptococcus gordonii
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2004-11-23 Classification: HYDROLASE Ligands: ZN, SO4, CL |
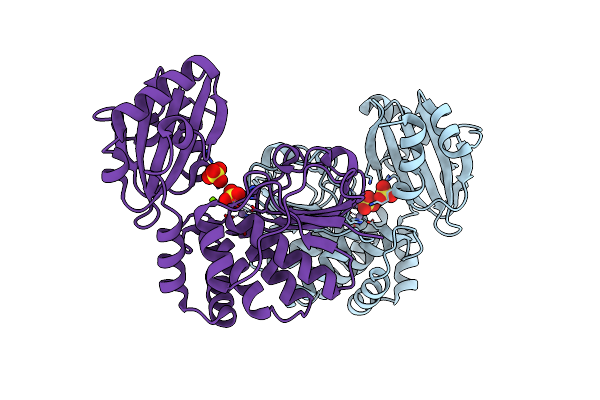 |
Organism: Streptococcus mutans
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2001-06-06 Classification: HYDROLASE Ligands: MN, MG, SO4 |
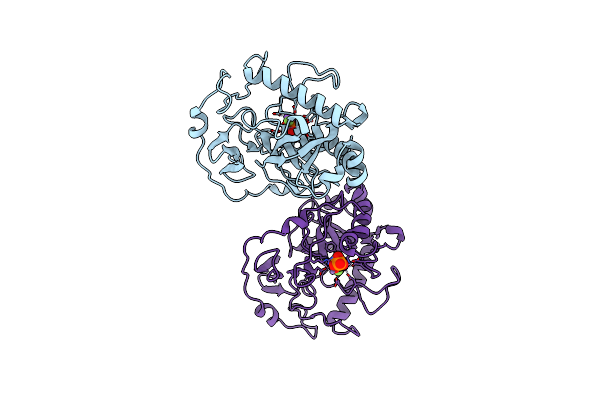 |
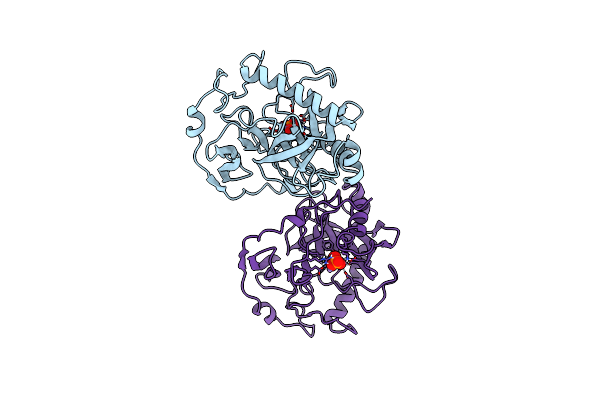
Fluoride-Inhibited Substrate Complex Of Saccharomyces Cerevisiae Inorganic Pyrophosphatase
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2001-03-19 Classification: PHOSPHORYL TRANSFER Ligands: MN, POP, F, NA, PO4 |
 |
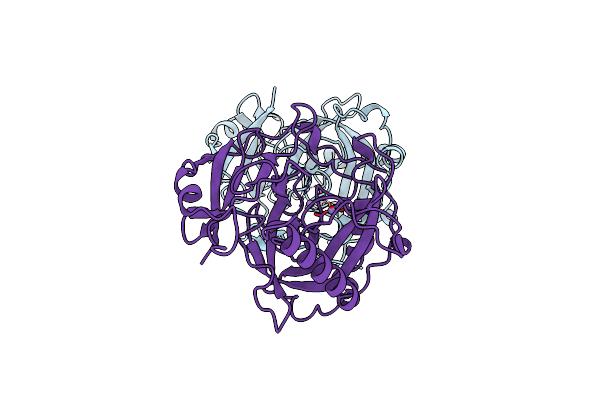
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:1.15 Å Release Date: 2001-03-19 Classification: HYDROLASE Ligands: MN, PO4 |
 |
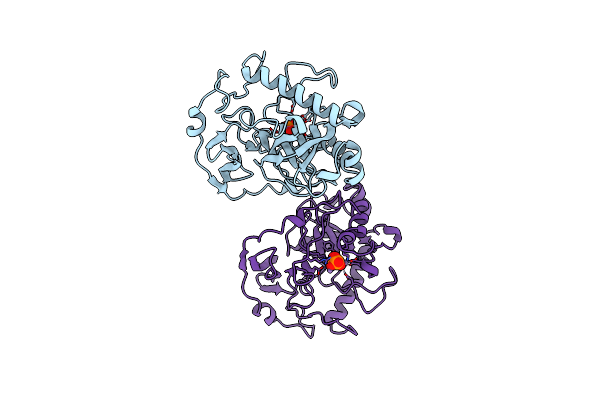
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 1997-11-19 Classification: HYDROLASE Ligands: MN |
 |
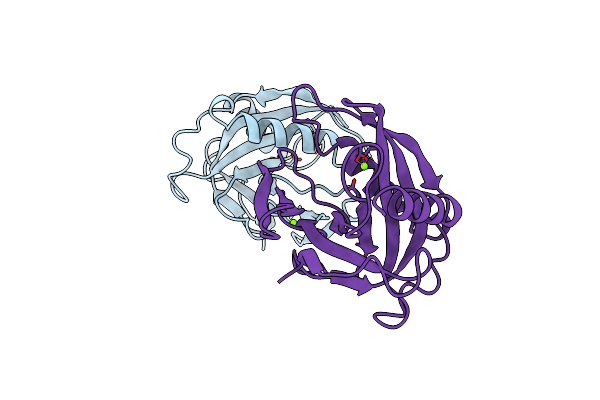
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 1997-11-19 Classification: HYDROLASE Ligands: MN, PO4 |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 1997-08-20 Classification: HYDROLASE Ligands: MG |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 1996-11-08 Classification: INORGANIC PYROPHOSPHATASE |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 1996-11-08 Classification: INORGANIC PYROPHOSPHATASE |

