Search Count: 1,141
All
Selected
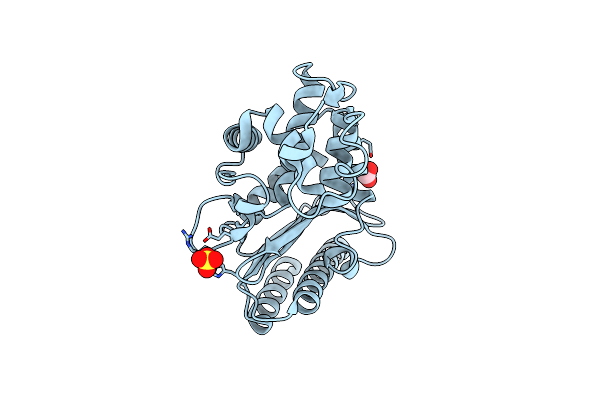 |
High Resolution Structure Of Class A Beta-Lactamase From Bordetella Bronchiseptica Rb50
Organism: Bordetella bronchiseptica rb50
Method: X-RAY DIFFRACTION Resolution:1.05 Å Release Date: 2024-06-12 Classification: HYDROLASE Ligands: FMT, SO4 |
 |
Structure Of Class A Beta-Lactamase From Bordetella Bronchiseptica Rb50 In A Complex With Avibactam
Organism: Bordetella bronchiseptica rb50
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2024-06-12 Classification: HYDROLASE Ligands: NXL, SIN, FMT, EDO |
 |
Structure Of Class A Beta-Lactamase From Bordetella Bronchiseptica Rb50 In A Complex With Clavulonate
Organism: Bordetella bronchiseptica rb50
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2024-06-12 Classification: HYDROLASE Ligands: MLA, TEM, CL |
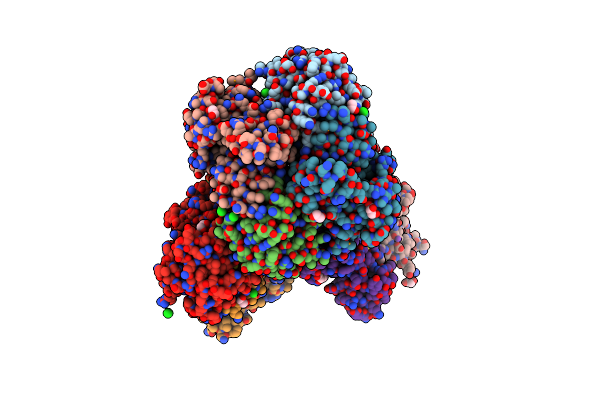 |
Crystal Structure Of Putataive Short-Chain Dehydrogenase/Reductase (Fabg) From Klebsiella Pneumoniae Subsp. Pneumoniae Ntuh-K2044 In Complex With Nadh
Organism: Klebsiella pneumoniae subsp. pneumoniae ntuh-k2044
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2022-03-02 Classification: BIOSYNTHETIC PROTEIN Ligands: K, CL, NAI, GOL, EDO |
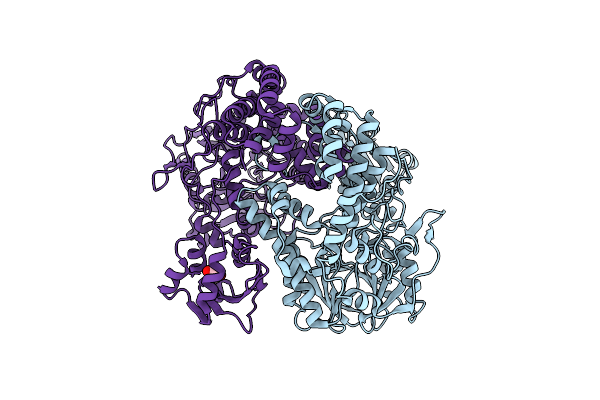 |
Organism: Klebsiella pneumoniae subsp. pneumoniae ntuh-k2044
Method: X-RAY DIFFRACTION Resolution:2.69 Å Release Date: 2022-02-02 Classification: HYDROLASE Ligands: EDO |
 |
Crystal Structure Of The Peptidyl-Prolyl Cis-Trans Isomerase (Ppib) From Streptococcus Pyogenes.
Organism: Streptococcus pyogenes mgas5005
Method: X-RAY DIFFRACTION Resolution:1.28 Å Release Date: 2021-12-01 Classification: ISOMERASE |
 |
Crystal Structure Of Peptidyl-Prolyl Cis-Trans Isomerasefrom (Ppib) Streptococcus Pneumoniae R6
Organism: Streptococcus pneumoniae (strain atcc baa-255 / r6)
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2021-12-01 Classification: ISOMERASE Ligands: CL, EDO, MES |
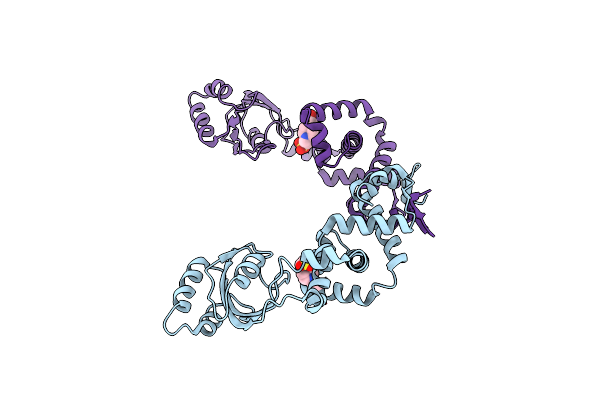 |
Crystal Structure Of Peptidylprolyl Isomerase Prsa From Streptococcus Mutans.
Organism: Streptococcus mutans
Method: X-RAY DIFFRACTION Resolution:3.15 Å Release Date: 2021-12-01 Classification: ISOMERASE Ligands: EPE, CL |
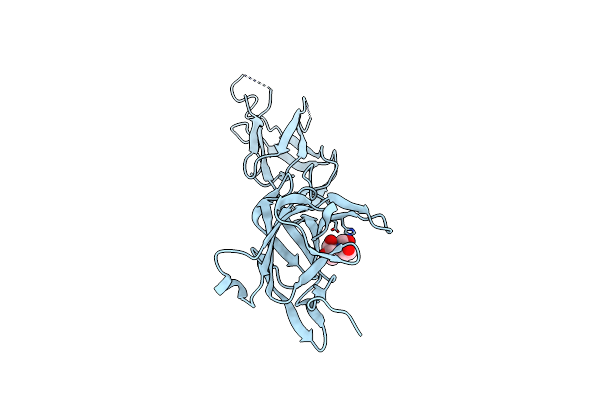 |
Crystal Structure Of Srtb-Anchored Collagen-Binding Adhesin Fragment (Residues 206-565) From Clostridioides Difficile Strain 630
Organism: Clostridioides difficile
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2021-10-20 Classification: CELL ADHESION Ligands: BTB |
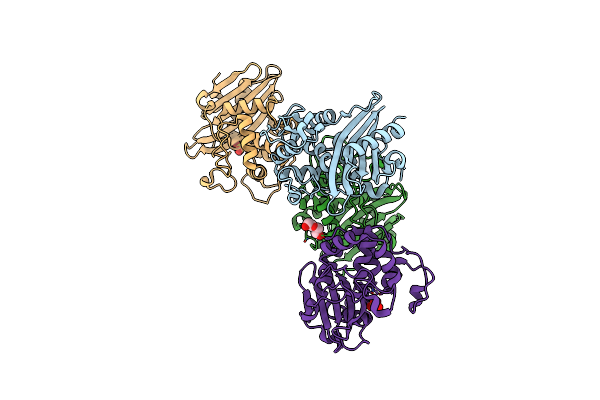 |
Crystal Structure Of C79A Mutant Of Class D Beta-Lactamase From Clostridium Difficile 630
Organism: Clostridioides difficile
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2021-08-11 Classification: HYDROLASE Ligands: PEG, SO4 |
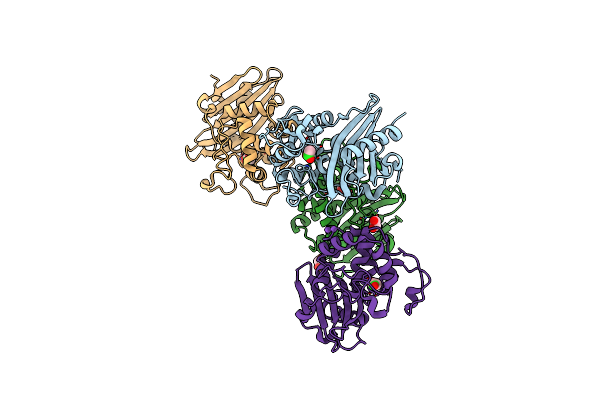 |
Crystal Structure Of K83A Mutant Of Class D Beta-Lactamase From Clostridium Difficile 630
Organism: Clostridioides difficile
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2021-08-11 Classification: HYDROLASE Ligands: CL, ACT, EDO, NA |
 |
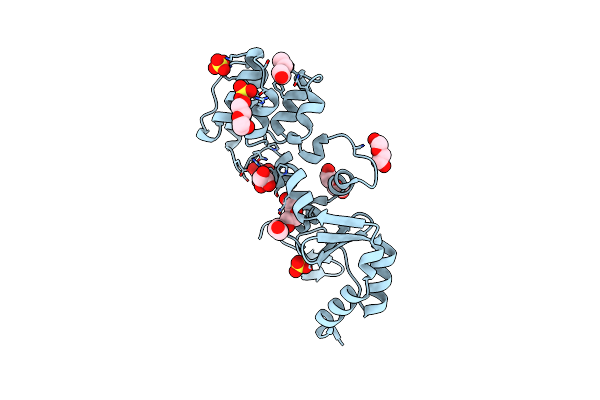
Crystal Structure Of The Peptidoglycan Binding Domain Of The Outer Membrane Protein (Ompa) From Klebsiella Pneumoniae With Bound D-Alanine
Organism: Klebsiella pneumoniae subsp. pneumoniae
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2021-07-28 Classification: PEPTIDE BINDING PROTEIN Ligands: DAL, CL |
 |
The Structure Of A Sensor Domain Of A Histidine Kinase (Vxra) From Vibrio Cholerae O1 Biovar Eltor Str. N16961, N239 Deletion Mutant
Organism: Vibrio cholerae serotype o1 (strain atcc 39315 / el tor inaba n16961)
Method: X-RAY DIFFRACTION Resolution:1.98 Å Release Date: 2021-01-27 Classification: SIGNALING PROTEIN Ligands: GOL, PEG, PG4, SO4 |
 |
The Structure Of A Sensor Domain Of A Histidine Kinase (Vxra) From Vibrio Cholerae O1 Biovar Eltor Str. N16961, 2Nd Form
Organism: Vibrio cholerae serotype o1 (strain atcc 39315 / el tor inaba n16961)
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2020-10-14 Classification: SIGNALING PROTEIN Ligands: GOL, ACT, CL, PEG, SRT |
 |
The Structure Of A Sensor Domain Of A Histidine Kinase (Vxra) From Vibrio Cholerae O1 Biovar Eltor Str. N16961, N239-T240 Deletion Mutant
Organism: Vibrio cholerae serotype o1 (strain atcc 39315 / el tor inaba n16961)
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2020-10-14 Classification: SIGNALING PROTEIN Ligands: EDO, SO4, MG |
 |
The Structure Of A Sensor Domain Of A Histidine Kinase (Vxra) From Vibrio Cholerae O1 Biovar Eltor Str. N16961, D238-T240 Deletion Mutant
Organism: Vibrio cholerae serotype o1 (strain atcc 39315 / el tor inaba n16961)
Method: X-RAY DIFFRACTION Resolution:1.98 Å Release Date: 2020-10-14 Classification: SIGNALING PROTEIN Ligands: GOL, EDO |
 |
The Crystal Structure Of A Functional Uncharacterized Protein Kp1_0663 From Klebsiella Pneumoniae Subsp. Pneumoniae Ntuh-K2044
Organism: Klebsiella pneumoniae subsp. pneumoniae ntuh-k2044
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2020-06-03 Classification: UNKNOWN FUNCTION |
 |
Crystal Structure Of Wild Type Class D Beta-Lactamase From Clostridium Difficile 630
Organism: Clostridioides difficile (strain 630)
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2020-05-27 Classification: HYDROLASE Ligands: PEG, NA |
 |
1.52 Angstrom Resolution Crystal Structure Of Transcriptional Regulator Hdfr From Klebsiella Pneumoniae
Organism: Klebsiella pneumoniae subsp. pneumoniae
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2020-05-06 Classification: TRANSCRIPTION Ligands: CL |
 |
2.70 Angstrom Resolution Crystal Structure Of Uracil Phosphoribosyl Transferase From Klebsiella Pneumoniae
Organism: Klebsiella pneumoniae subsp. pneumoniae
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2020-05-06 Classification: TRANSFERASE Ligands: BDF, SO4, CL |

