Search Count: 5,429
All
Selected
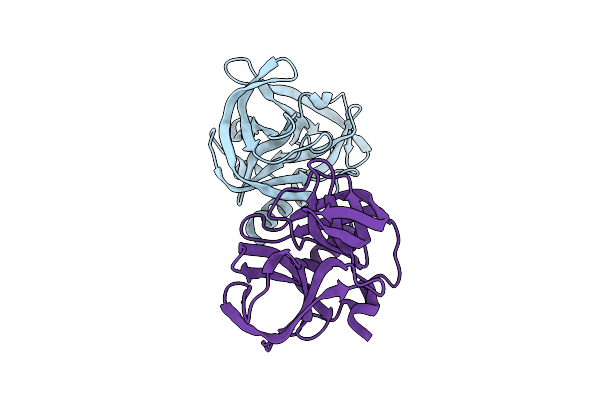 |
Crystal Structure Of Enterovirus D68 3C Protease Determined Via Sulfur Phasing
Organism: Enterovirus d68
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-04-01 Classification: VIRAL PROTEIN Ligands: CL |
 |
Crystal Structure Of Zika Virus Ns2B-Ns3 Protease Determined Via Sulfur Phasing
Organism: Zika virus
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2026-04-01 Classification: VIRAL PROTEIN |
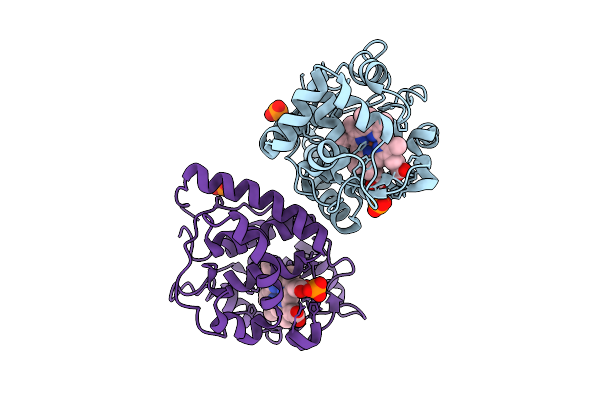 |
Organism: Glycine max
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: HEM, PO4 |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: RIBOSOME Ligands: MG, NA, ZN |
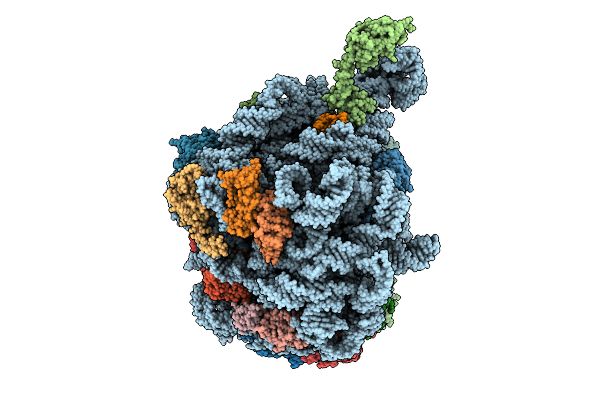 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: RIBOSOME Ligands: MG, NA, ZN |
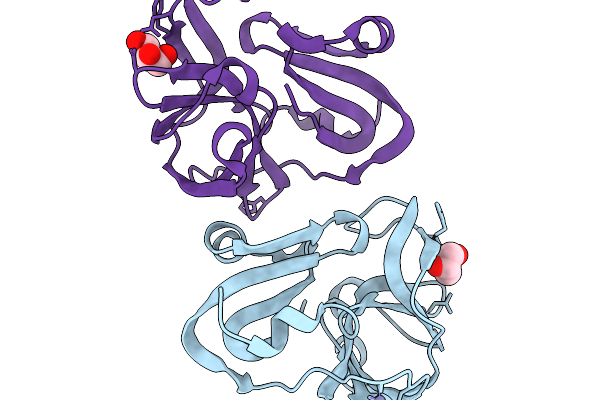 |
Crystal Structure Of Coxsackievirus A16 (G-10) 2A Protease Determined Via Sulfur Phasing
Organism: Coxsackievirus a16
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-03-25 Classification: VIRAL PROTEIN Ligands: GOL, ZN |
 |
Organism: Trypanosoma cruzi
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-25 Classification: LYASE Ligands: SO4 |
 |
Organism: Streptomyces fagopyri
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2026-03-18 Classification: BIOSYNTHETIC PROTEIN |
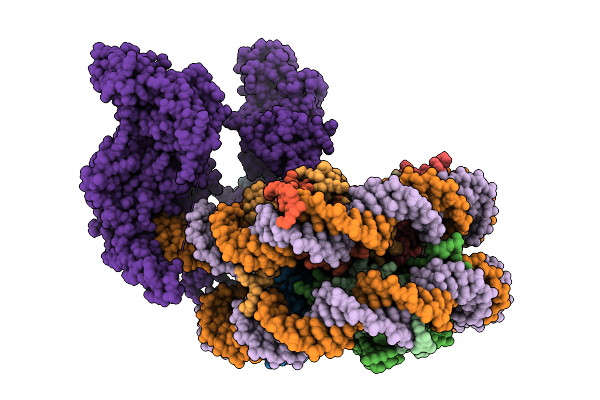 |
Uhrf1 Bound To A Mononucleosome In Its Pre-Active State, With The Ring Domain Bound To The Sra Domain.
Organism: Xenopus laevis, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:4.10 Å Release Date: 2026-03-11 Classification: DNA BINDING PROTEIN Ligands: ZN |
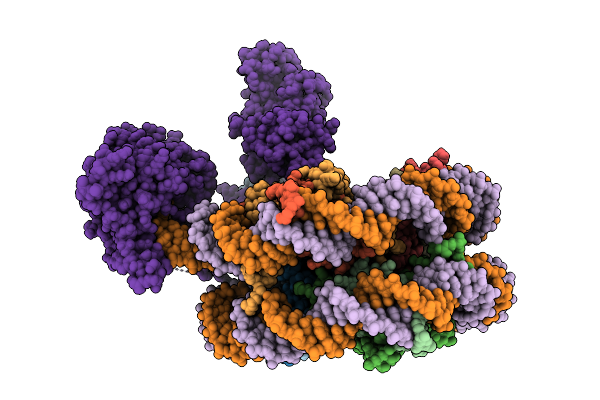 |
The Activated State Of Human Uhrf1 Bound To A Mononucleosome, With The Finger Loop Ordered And Linker 4 Disordered.
Organism: Xenopus laevis, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.70 Å Release Date: 2026-03-11 Classification: DNA BINDING PROTEIN Ligands: ZN |
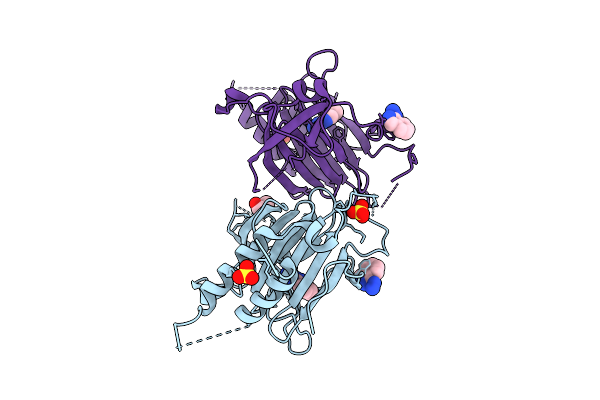 |
Crystal Structure Of Ferric Human Ado C18S/C239S Variant In Complex With Hydralazine At 1.98 Angstrom Resolution
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.98 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: FE, HLZ, SO4, GOL |
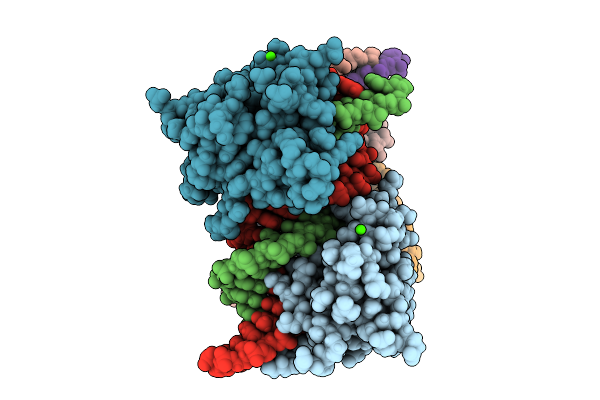 |
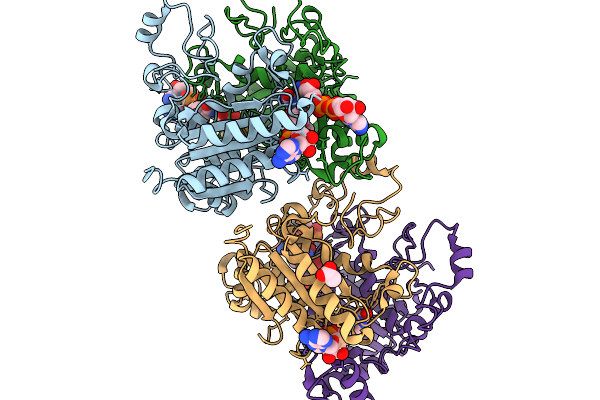
Investigating The Binding Mechanism Of Interferon Regulatory Factor 4 To Dna In The Context Of Multiple Myeloma
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-03-11 Classification: DNA BINDING PROTEIN Ligands: CA |
 |
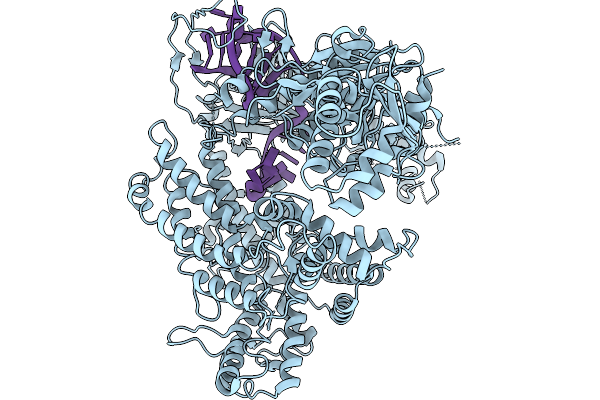
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: NAD, FAD, ACT |
 |
Organism: Francisella tularensis subsp. novicida u112, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.26 Å Release Date: 2026-02-11 Classification: DNA BINDING PROTEIN |
 |
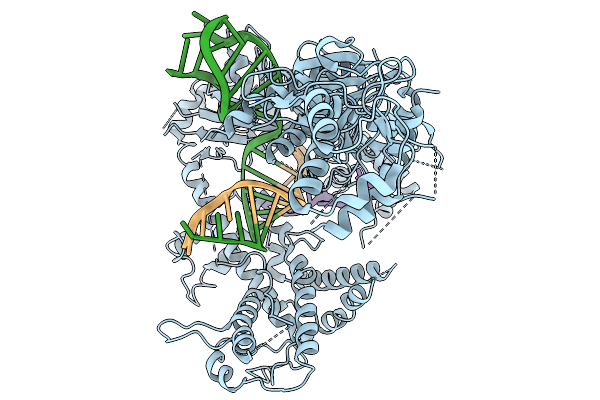
Organism: Francisella tularensis subsp. novicida u112, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:4.00 Å Release Date: 2026-02-11 Classification: DNA BINDING PROTEIN |
 |
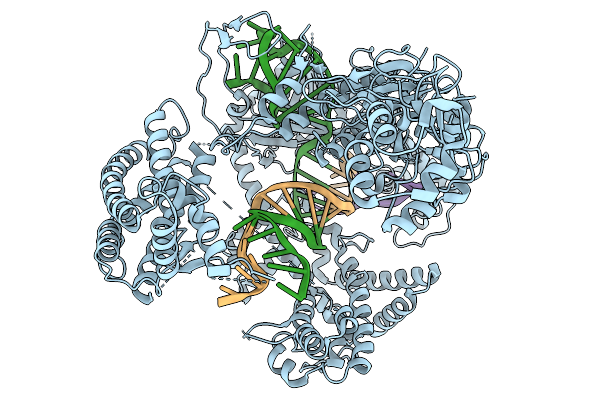
Organism: Francisella tularensis subsp. novicida u112, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.68 Å Release Date: 2026-02-11 Classification: DNA BINDING PROTEIN |
 |
Organism: Francisella tularensis subsp. novicida, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.63 Å Release Date: 2026-02-11 Classification: DNA BINDING PROTEIN |
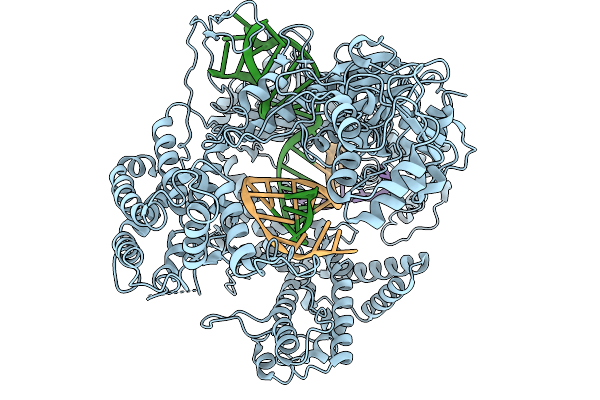 |
Organism: Francisella tularensis subsp. novicida, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.21 Å Release Date: 2026-02-11 Classification: DNA BINDING PROTEIN |
 |
Native Structure Of Full-Length Pesticidal Protein Cry1Ca18 At Ph9, From Crystals Formed In Vivo
Organism: Bacillus thuringiensis
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-02-11 Classification: TOXIN Ligands: CA, ZN |
 |
Organism: Dioscoreophyllum cumminsii
Method: X-RAY DIFFRACTION Resolution:2.87 Å Release Date: 2026-02-04 Classification: PLANT PROTEIN |

