Search Count: 4,710
All
Selected
 |
Organism: Amycolatopsis deserti
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2026-04-15 Classification: HYDROLASE |
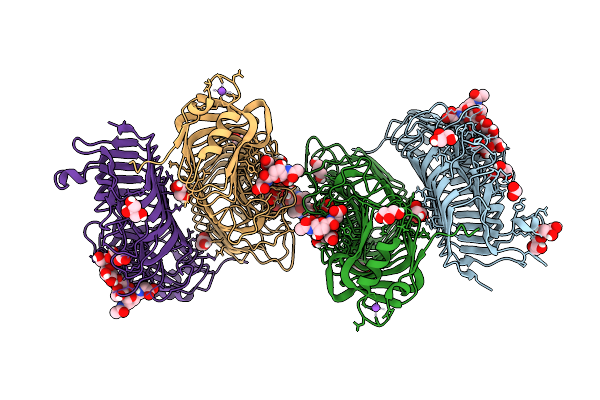 |
Crystal Structure Of The Polysaccharide Lyase Rbmb From Vibrio Cholerae Bound To Vibrio Polysaccharide
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2026-03-11 Classification: LYASE |
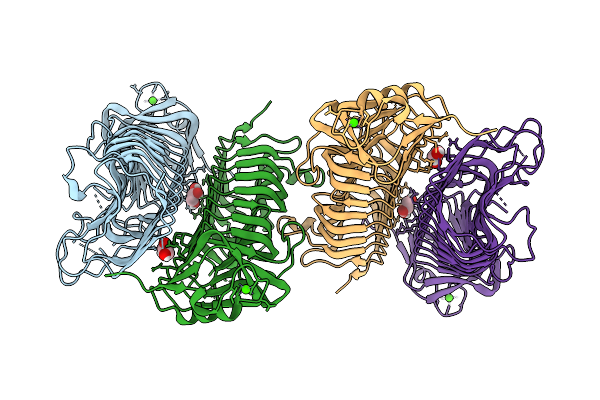 |
Organism: Vibrio cholerae c6706
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-03-11 Classification: LYASE Ligands: GOL, CA |
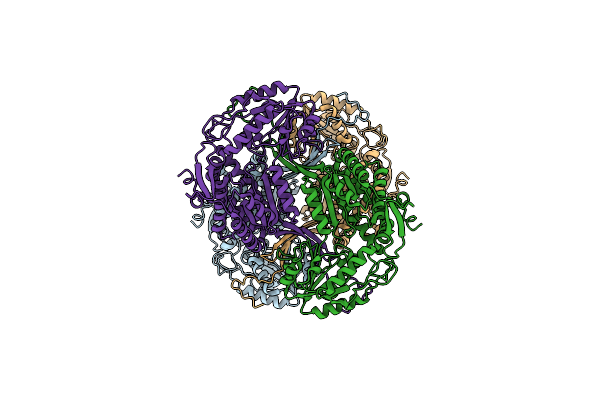 |
Cryoem Structure Of Aldehyde Dehydrogenase From Francisella Tularensis Subsp. Tularensis At 3.03A Resolution
Organism: Francisella tularensis subsp. tularensis
Method: ELECTRON MICROSCOPY Resolution:3.03 Å Release Date: 2026-01-28 Classification: OXIDOREDUCTASE |
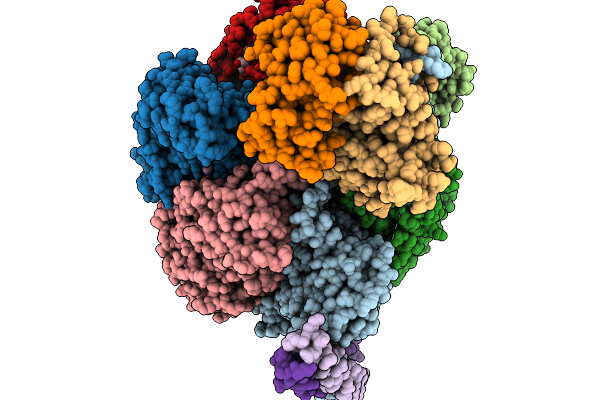 |
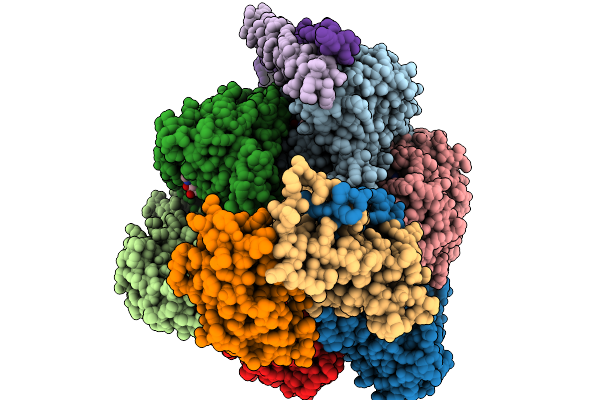
Cryo-Em Structure Of F-Atp Synthase From Mycobacteroides Abscessus (Rotational State 1)
Organism: Mycobacteroides abscessus subsp. abscessus
Method: ELECTRON MICROSCOPY Resolution:2.94 Å Release Date: 2026-01-14 Classification: HYDROLASE Ligands: ATP, MG, ADP |
 |
Cryo-Em Structure Of F-Atp Synthase From Mycobacteroides Abscessus (Rotational State 2)
Organism: Mycobacteroides abscessus subsp. abscessus
Method: ELECTRON MICROSCOPY Resolution:3.41 Å Release Date: 2026-01-14 Classification: HYDROLASE Ligands: ATP, MG, ADP |
 |
Cryo-Em Structure Of F-Atp Synthase From Mycobacteroides Abscessus (Rotational State 3)
Organism: Mycobacteroides abscessus subsp. abscessus
Method: ELECTRON MICROSCOPY Resolution:2.79 Å Release Date: 2026-01-14 Classification: HYDROLASE Ligands: ATP, MG, ADP |
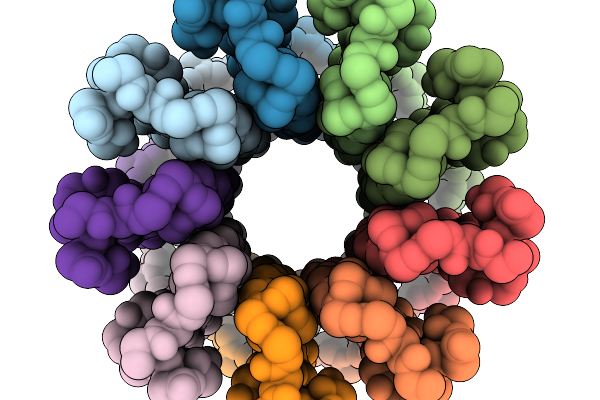 |
Cryo-Em Structure Of F-Atp Synthase C-Ring From Mycobacteroides Abscessus (Backbone)
Organism: Mycobacteroides abscessus subsp. abscessus
Method: ELECTRON MICROSCOPY Resolution:5.61 Å Release Date: 2026-01-14 Classification: HYDROLASE |
 |
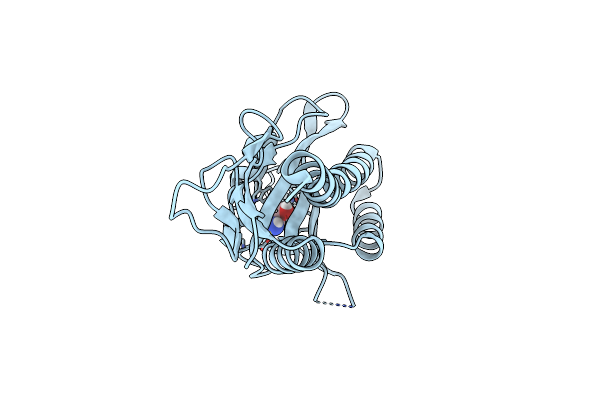
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2025-07-02 Classification: SIGNALING PROTEIN Ligands: CL, ETA |
 |
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2025-07-02 Classification: SIGNALING PROTEIN Ligands: 2A1, CL |
 |
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2025-07-02 Classification: SIGNALING PROTEIN Ligands: SEL, CL |
 |
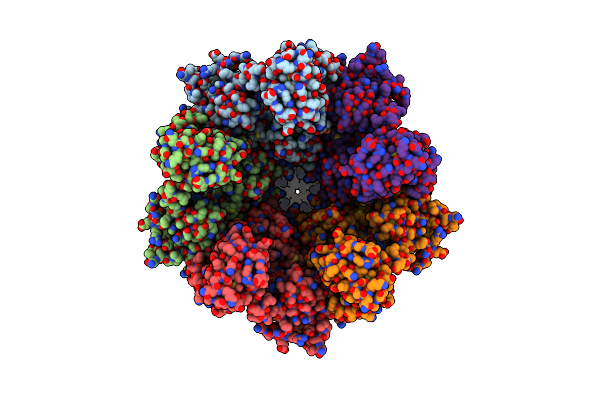
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-07-02 Classification: SIGNALING PROTEIN |
 |
3Dflex Refinement Of The Cryoem Structure Of Declic Nanodisc With 10Mm Edta In Sym-Like State
Organism: Desulfofustis sp. pb-srb1
Method: ELECTRON MICROSCOPY Release Date: 2025-04-09 Classification: TRANSLOCASE |
 |
3Dflex Refinement Of The Cryoem Structure Of Declic Nanodisc With 10Mm Edta In Asym State
Organism: Desulfofustis sp. pb-srb1
Method: ELECTRON MICROSCOPY Release Date: 2025-04-09 Classification: TRANSLOCASE |
 |
Non-Uniform Refinement Of The Cryoem Structure Of Declic Nanodisc With 10Mm Calcium
Organism: Desulfofustis sp. pb-srb1
Method: ELECTRON MICROSCOPY Release Date: 2025-04-09 Classification: TRANSLOCASE Ligands: CA, OCT, D12, D10, C14, PX6 |
 |
Non-Uniform Refinement Of The Cryoem Structure Of Declic Nanodisc With 10Mm Edta In Asym State
Organism: Desulfofustis sp. pb-srb1
Method: ELECTRON MICROSCOPY Release Date: 2025-04-09 Classification: TRANSLOCASE Ligands: OCT, D10, C14 |
 |
3Dflex Refinement Of The Cryoem Structure Of Declic Nanodisc With 10Mm Calcium
Organism: Desulfofustis sp. pb-srb1
Method: ELECTRON MICROSCOPY Release Date: 2025-04-09 Classification: TRANSLOCASE Ligands: CA |
 |
Non-Uniform Refinement Of The Cryoem Structure Of Declic Nanodisc With 10Mm Edta In Sym-Like State
Organism: Desulfofustis sp. pb-srb1
Method: ELECTRON MICROSCOPY Release Date: 2025-04-09 Classification: TRANSLOCASE Ligands: OCT, D10, C14 |
 |
Organism: Human herpesvirus 8 strain gk18
Method: ELECTRON MICROSCOPY Release Date: 2025-02-19 Classification: VIRAL PROTEIN |
 |
Closed Conformation Of The Pentameric Ligand-Gated Ion Channel, Declic At Ph 5 With 10 Mm Ca2+
Organism: Desulfofustis sp. pb-srb1
Method: ELECTRON MICROSCOPY Release Date: 2024-09-11 Classification: MEMBRANE PROTEIN Ligands: CA |

