Search Count: 1,532
All
Selected
 |
Organism: Prevotella
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-01-28 Classification: HYDROLASE Ligands: GOL |
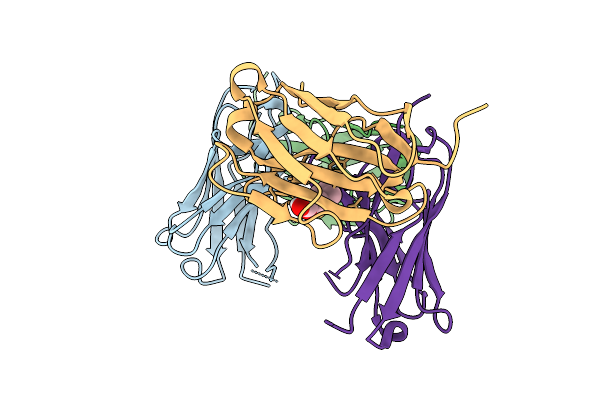 |
Organism: Camelus dromedarius
Method: X-RAY DIFFRACTION Resolution:2.86 Å Release Date: 2026-01-28 Classification: IMMUNE SYSTEM Ligands: PG4 |
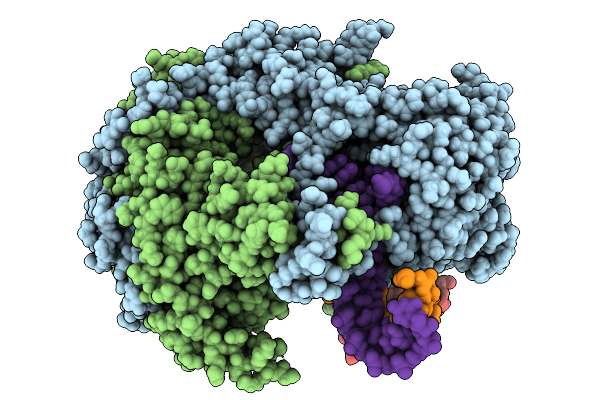 |
Organism: Influenza a virus (strain a/goose/guangdong/1/1996 h5n1 genotype gs/gd)
Method: ELECTRON MICROSCOPY Resolution:2.98 Å Release Date: 2026-01-21 Classification: VIRAL PROTEIN/RNA |
 |
Crystal Structure Of S. Aureus Protein A Bound To A Camelid Single-Domain Antibody
Organism: Staphylococcus aureus (strain nctc 8325 / ps 47), Camelus dromedarius
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2025-11-12 Classification: IMMUNE SYSTEM |
 |
Organism: Homo sapiens, Camelus dromedarius
Method: ELECTRON MICROSCOPY Resolution:2.71 Å Release Date: 2025-07-23 Classification: MEMBRANE PROTEIN Ligands: NAG, ZN, CA, CL |
 |
Organism: Homo sapiens, Camelus dromedarius
Method: ELECTRON MICROSCOPY Resolution:2.66 Å Release Date: 2025-07-16 Classification: MEMBRANE PROTEIN Ligands: NAG, ZN, CA, CL |
 |
Organism: Homo sapiens, Camelus dromedarius
Method: ELECTRON MICROSCOPY Resolution:2.46 Å Release Date: 2025-07-16 Classification: MEMBRANE PROTEIN Ligands: NAG, ZN, CA, CL |
 |
Organism: Homo sapiens, Camelus dromedarius
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2025-07-16 Classification: MEMBRANE PROTEIN Ligands: NAG, ZN, CA, CL |
 |
Organism: Homo sapiens, Camelus dromedarius
Method: ELECTRON MICROSCOPY Resolution:2.63 Å Release Date: 2025-07-16 Classification: MEMBRANE PROTEIN |
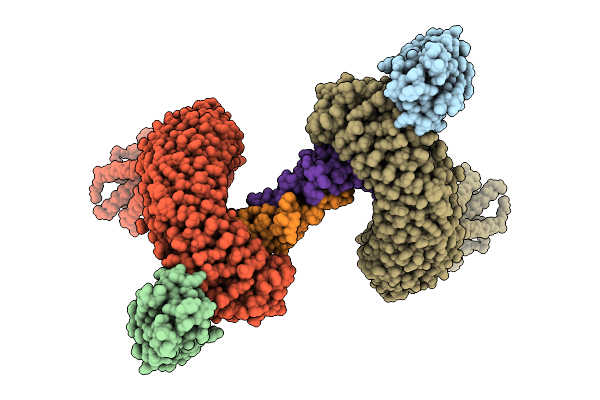 |
Organism: Camelus dromedarius, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-16 Classification: MEMBRANE PROTEIN |
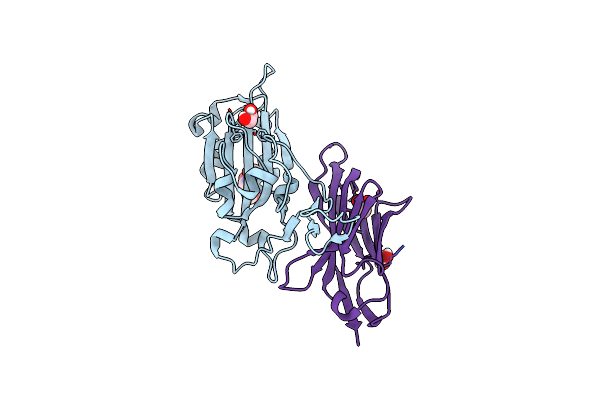 |
The Crystal Structure Of The Sars-Cov-2 Receptor Binding Domain In Complex With The Neutralizing Nanobody 4.
Organism: Severe acute respiratory syndrome coronavirus 2, Camelus dromedarius
Method: X-RAY DIFFRACTION Resolution:1.21 Å Release Date: 2025-06-11 Classification: ANTIVIRAL PROTEIN Ligands: NAG, EDO, ACT |
 |
Crystal Structure Of The Decarboxylase Kdc4427 From Enterobacter Sp. Cgmcc 5087
Organism: Enterobacter sp. cgmcc 5087
Method: X-RAY DIFFRACTION Resolution:1.94 Å Release Date: 2025-05-21 Classification: LYASE Ligands: TPP, MG |
 |
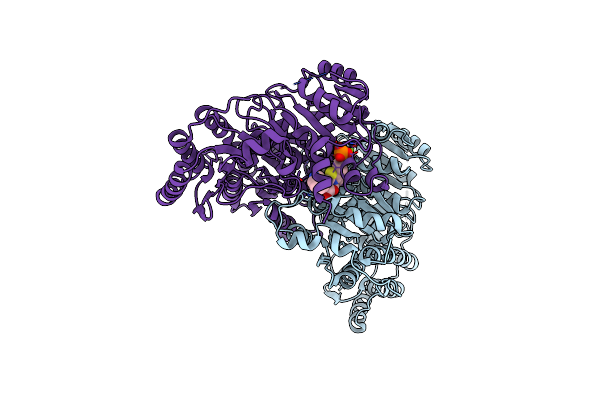
Crystal Structure Of The Decarboxylase Kdc4427 Mutant E468L From Enterobacter Sp. Cgmcc 5087
Organism: Enterobacter sp. cgmcc 5087
Method: X-RAY DIFFRACTION Resolution:2.32 Å Release Date: 2025-05-21 Classification: LYASE Ligands: TPP, MG |
 |
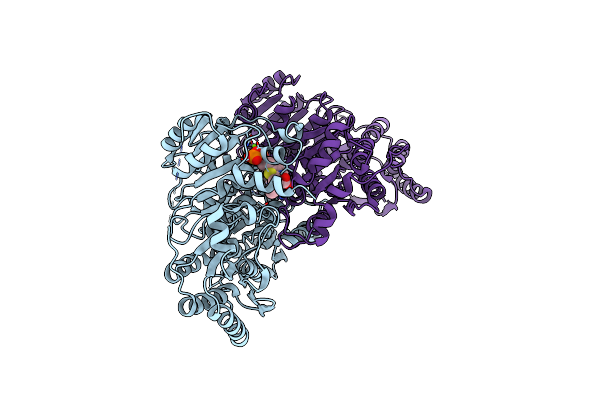
Crystal Structure Of The Decarboxylase Kdc4427 Mutant E468L In Complex With Indole-3-Pyruvic Acid
Organism: Enterobacter sp. cgmcc 5087
Method: X-RAY DIFFRACTION Resolution:2.16 Å Release Date: 2025-05-21 Classification: LYASE Ligands: TPP, MG, 3IO |
 |
Crystal Structure Of The Decarboxylase Kdc4427 Mutant E468L In Complex With Phenylpyruvic Acid
Organism: Enterobacter sp. cgmcc 5087
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2025-05-21 Classification: LYASE Ligands: TPP, MG, PPY |
 |
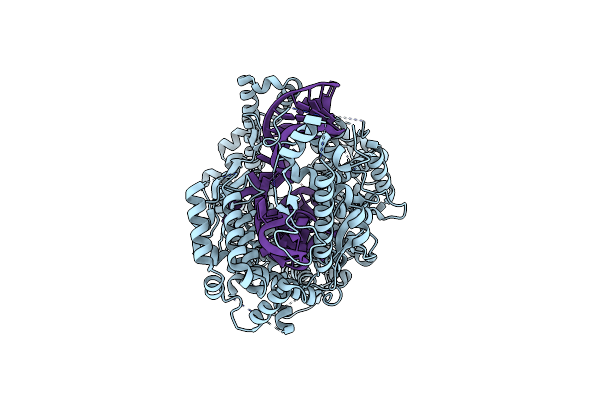
Crystal Structure Of The Decarboxylase Kdc4427 In Complex With Phenylpyruvic Acid Intermediate
Organism: Enterobacter sp. cgmcc 5087
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2025-05-21 Classification: LYASE Ligands: MG, A1L14 |
 |
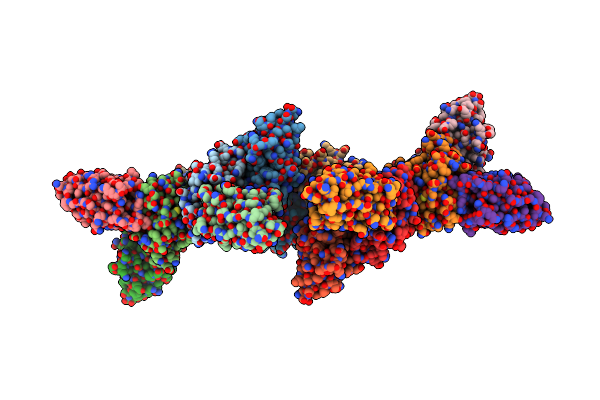
Organism: Listeria seeligeri serovar 1/2b str. slcc3954
Method: ELECTRON MICROSCOPY Release Date: 2025-04-23 Classification: RNA BINDING PROTEIN/RNA |
 |
Organism: Homo sapiens, Vicugna pacos, Camelus dromedarius
Method: X-RAY DIFFRACTION Resolution:3.94 Å Release Date: 2025-02-12 Classification: OXIDOREDUCTASE/IMMUNE SYSTEM Ligands: ZN |
 |
Crystal Structure Of The C. Difficile Toxin A Crops Domain Fragment 2592-2710 Bound To H5.2 Nanobody
Organism: Camelus dromedarius, Clostridioides difficile 342
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2024-12-18 Classification: TOXIN/IMMUNE SYSTEM Ligands: EDO, PO4 |
 |
Crystal Structure Of The C. Difficile Toxin A Crops Domain Fragment 2639-2707 Bound To C4.2 Nanobody
Organism: Camelus dromedarius, Clostridioides difficile
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2024-12-18 Classification: TOXIN/IMMUNE SYSTEM Ligands: EDO |

