Search Count: 2,513
All
Selected
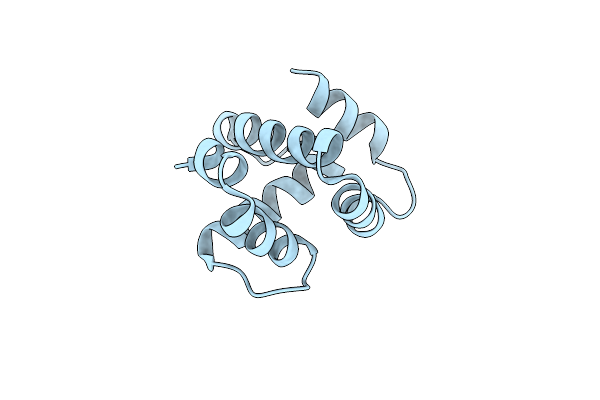 |
Organism: Spodoptera litura
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-01-01 Classification: STRUCTURAL PROTEIN |
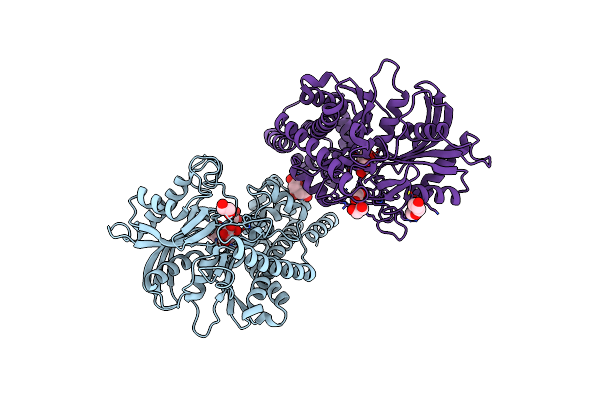 |
Organism: Plasmodium
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2023-08-30 Classification: TRANSFERASE Ligands: GLC, CIT, EDO, PGE |
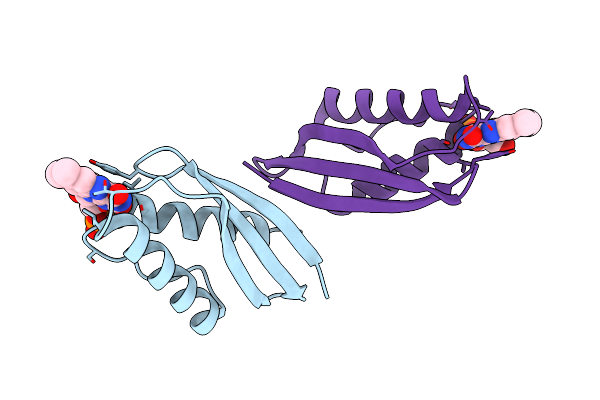 |
Organism: Clostridiaceae bacterium
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2023-07-19 Classification: FLAVOPROTEIN Ligands: FMN |
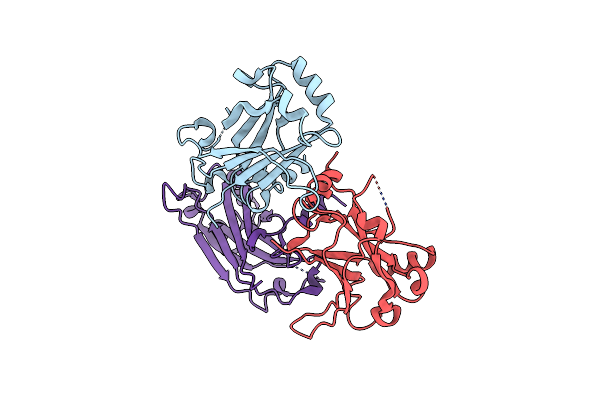 |
Organism: Notothenia coriiceps
Method: X-RAY DIFFRACTION Resolution:2.66 Å Release Date: 2023-07-12 Classification: IMMUNE SYSTEM |
 |
Cytoplasmic Domain Structure Of The Mgte Mg2+ Channel From Clostridiales Bacterium
Organism: Clostridiales bacterium
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2023-04-19 Classification: TRANSPORT PROTEIN Ligands: EPE, MG |
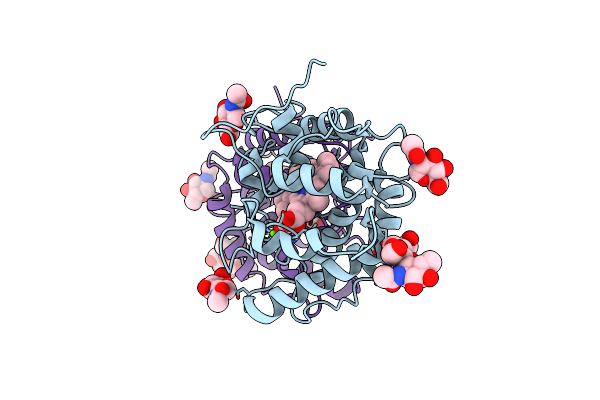 |
Artificial Unspecific Peroxygenase Expressed In Pichia Pastoris At 2.01 Angstrom Resolution
Organism: Marasmius rotula
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2023-03-01 Classification: OXIDOREDUCTASE Ligands: HEM, NAG, MG, CL |
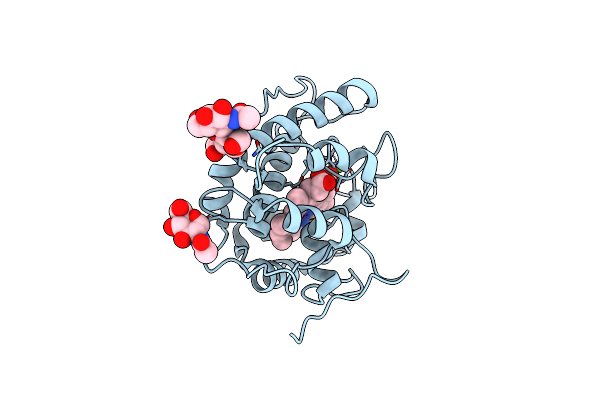 |
Artificial Unspecific Peroxygenase Expressed In Pichia Pastoris At 1.21 Angstrom Resolution
Organism: Marasmius rotula
Method: X-RAY DIFFRACTION Resolution:1.21 Å Release Date: 2023-03-01 Classification: OXIDOREDUCTASE Ligands: HEM, NAG, MG |
 |
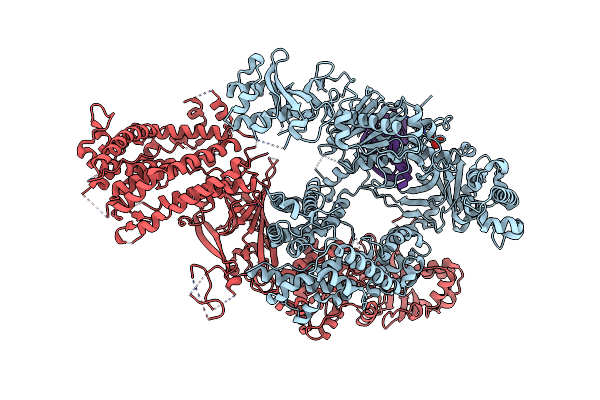
Artificial Unspecific Peroxygenase Expressed In Escherichia Coli At 2.09 Angstrom Resolution
Organism: Marasmius rotula
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2023-03-01 Classification: OXIDOREDUCTASE Ligands: HEM, MG |
 |
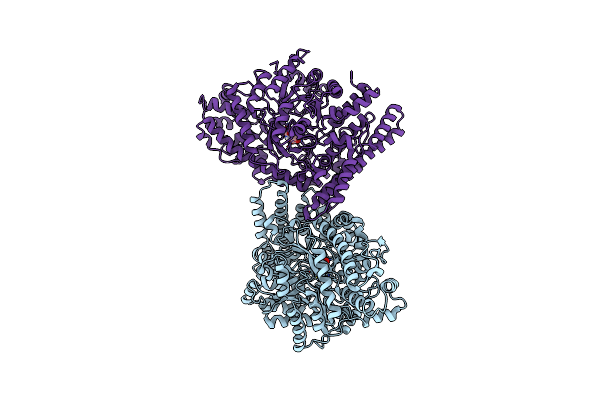
Organism: Lachnospiraceae bacterium
Method: X-RAY DIFFRACTION Resolution:3.10 Å Release Date: 2023-02-08 Classification: DNA BINDING PROTEIN Ligands: SO4, MG |
 |
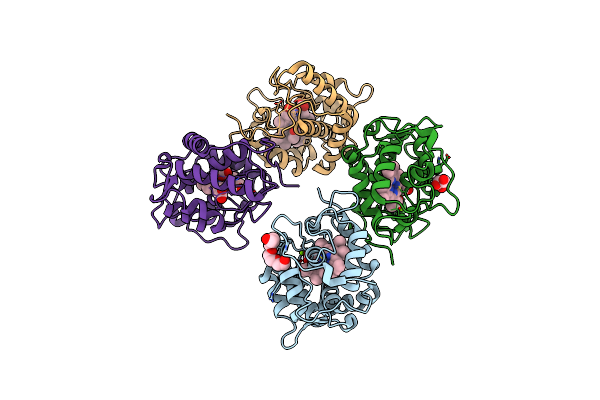
Anaerobic Hydroxyproline Degradation Involving C-N Cleavage By A Glycyl Radical Enzyme
Organism: Clostridiales bacterium
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2022-06-01 Classification: LYASE Ligands: UY7 |
 |
Organism: Marasmius rotula
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2022-05-18 Classification: OXIDOREDUCTASE Ligands: PEG, HEM, MG, ACT, GOL |
 |
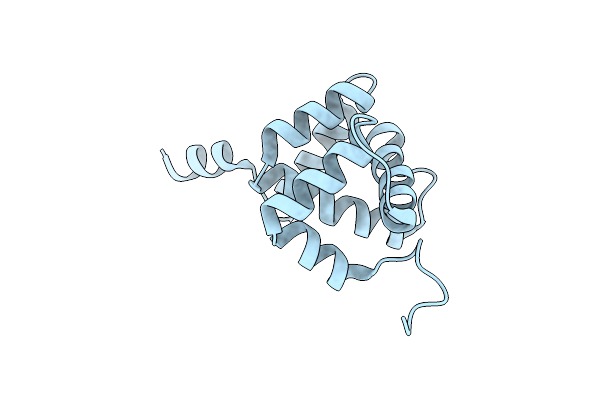
Organism: Gloeocapsa sp. pcc 7513
Method: X-RAY DIFFRACTION Resolution:2.17 Å Release Date: 2022-04-13 Classification: PHOTOSYNTHESIS Ligands: 45D |
 |
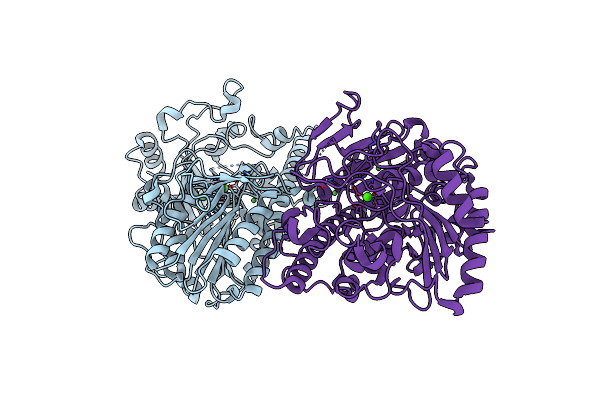
Organism: Spodoptera litura
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2022-03-02 Classification: STRUCTURAL PROTEIN |
 |
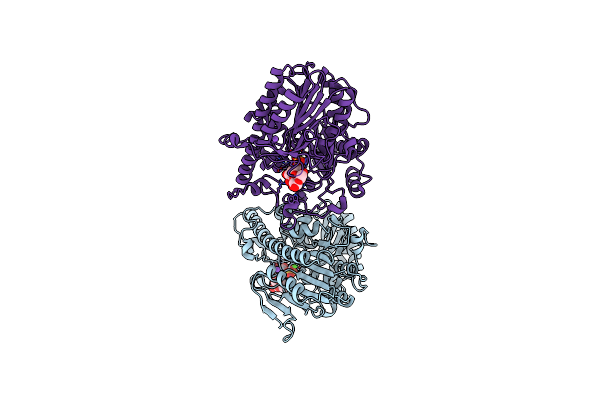
Structure Of The Exo-L-Galactose-6-Sulfatase Bus1_11 From Bacteroides Uniformis
Organism: Bacteroides uniformis
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2022-02-09 Classification: HYDROLASE Ligands: CA, NI |
 |
Structure Of Exo-L-Galactose-6-Sulfatase Bus1_11 From Bacteroides Uniformis In Complex With Neoporphyrabiose
Organism: Bacteroides uniformis
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2022-02-09 Classification: HYDROLASE Ligands: CA, NA |
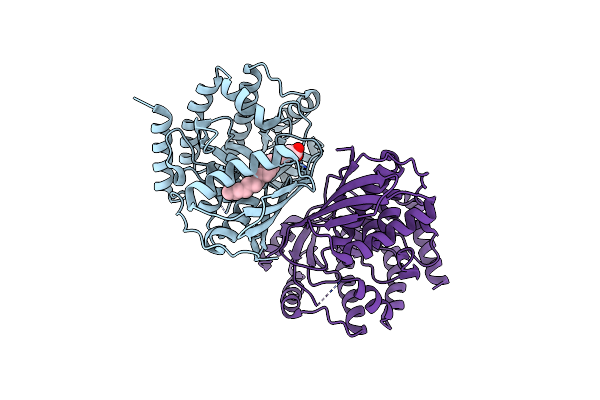 |
Crystal Structure Of Mevalonate 3,5-Bisphosphate Decarboxylase From Picrophilus Torridus
Organism: Picrophilus torridus dsm 9790
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2021-12-22 Classification: LYASE Ligands: OLA |
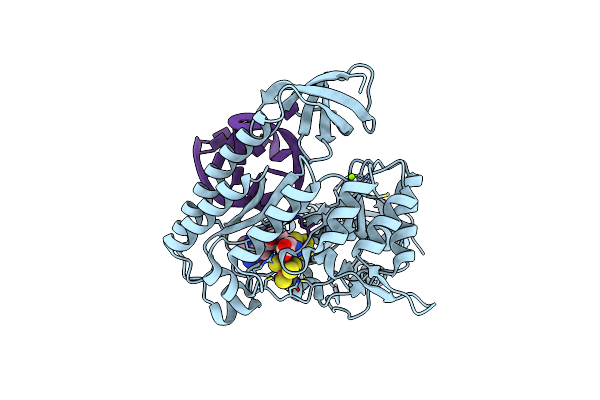 |
Organism: Bacteroides uniformis
Method: X-RAY DIFFRACTION Resolution:2.24 Å Release Date: 2021-09-15 Classification: TRANSFERASE/RNA Ligands: F3S, SF4, SAM, MG |
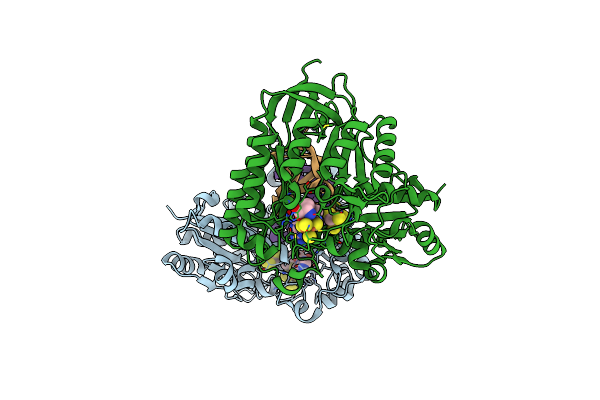 |
Organism: Bacteroides uniformis
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2021-09-15 Classification: TRANSFERASE/RNA Ligands: ZKP, MET, 5AD, SF4, MG |
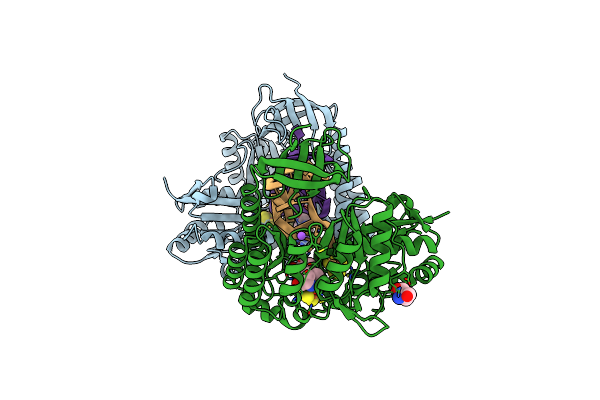 |
Organism: Bacteroides uniformis
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2021-09-15 Classification: TRANSFERASE/RNA Ligands: F3S, SF4, 5AD, MET, NA, TRS |
 |
Organism: Bacteroides uniformis
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2021-09-15 Classification: TRANSFERASE/RNA Ligands: PGE, PEG, F3S, SF4, SAH, CL |

