Search Count: 2,167
All
Selected
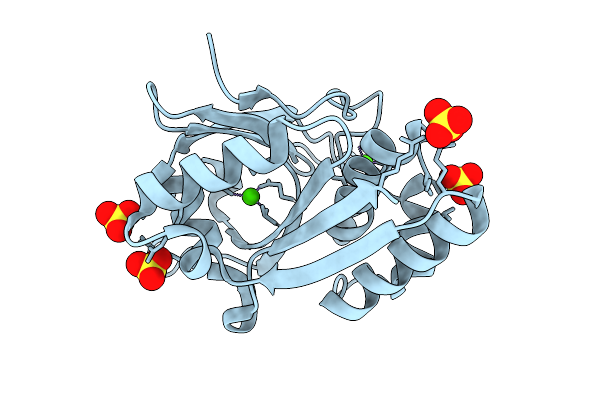 |
Organism: Metagenomes
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-18 Classification: LYASE Ligands: CA, SO4 |
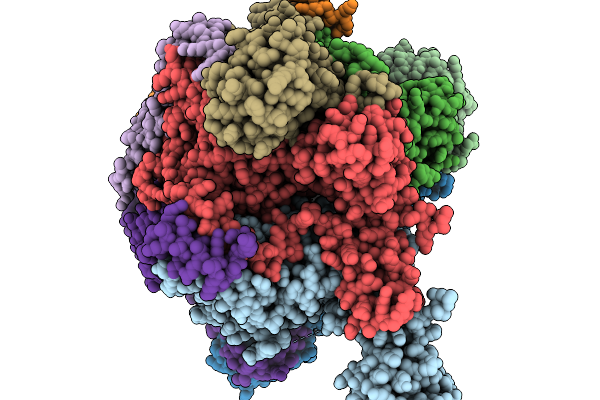 |
Ecoli Dnab Helicase And Phage Lambda Loader P With Adp-Mg In A 6:5 Stoichiometry Ratio
Organism: Escherichia coli, Escherichia phage lambda
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN Ligands: ADP, MG |
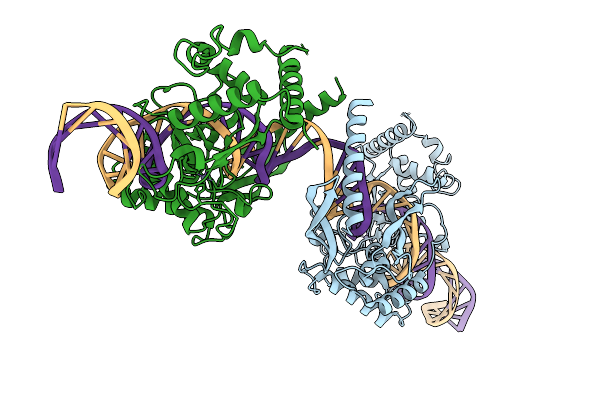 |
Organism: Streptomyces phage phi-c31, Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN Ligands: ZN |
 |
Cryo-Em Structure Of Bacteriophage P22 Gp1-Gp5-Gp4 Complex At 2.76 Angstrom
Organism: Salmonella phage p22
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: VIRAL PROTEIN |
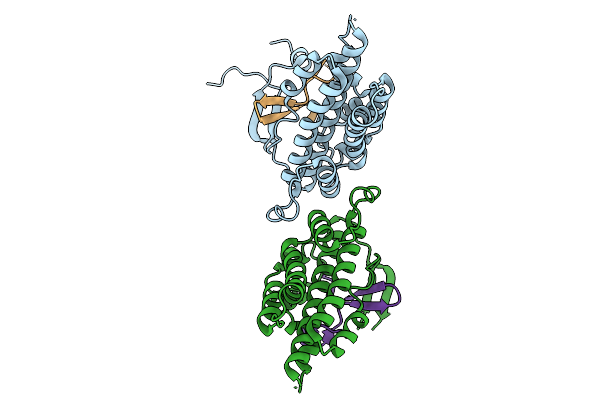 |
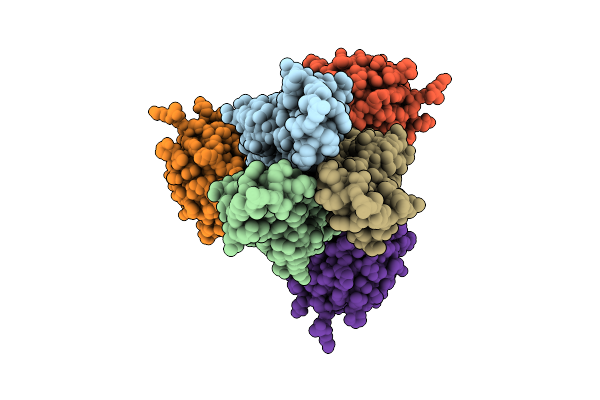
Structure Of S. Typhimurium 14028 Gifsy-1 Prophage Heps Bound To Bacteriophage Lambda J Tail Tip
Organism: Salmonella enterica subsp. enterica serovar typhimurium str. atcc 14028, Escherichia phage lambda
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2026-01-28 Classification: VIRAL PROTEIN |
 |
Organism: Escherichia phage lambda, Escherichia coli o157:h7 str. edl933
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: VIRAL PROTEIN |
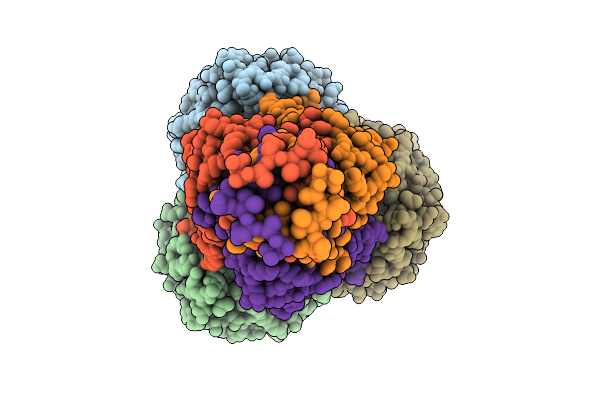 |
Open State Of A8 Gpj 713 Central Tail Fiber With Ompc G17 From E. Coli Edl933
Organism: Escherichia phage lambda, Escherichia coli o157:h7 str. edl933
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: VIRAL PROTEIN |
 |
Cryo-Em Structure Of E.Coli Transcription Initiation Complex With Escherichia Phage Mu Late Transcription Activator C
Organism: Escherichia coli (strain k12), Escherichia phage mu
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: TRANSCRIPTION/DNA |
 |
Cryo-Em Structure Of E.Coli Transcription Initiation Complex With Escherichia Phage Mu Middle Transcription Activator Mor
Organism: Escherichia coli (strain k12), Escherichia phage mu
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: TRANSCRIPTION/DNA |
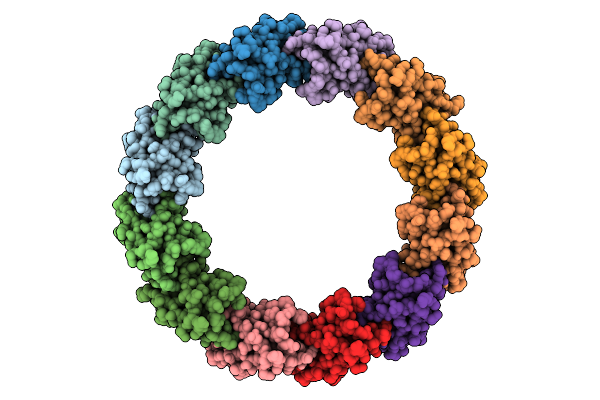 |
Organism: Escherichia phage lambda
Method: ELECTRON MICROSCOPY Release Date: 2025-11-26 Classification: VIRAL PROTEIN |
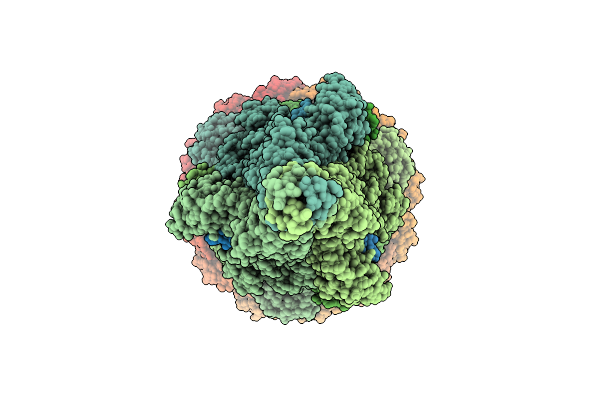 |
Organism: Escherichia phage lambda
Method: ELECTRON MICROSCOPY Release Date: 2025-11-05 Classification: VIRAL PROTEIN Ligands: SF4 |
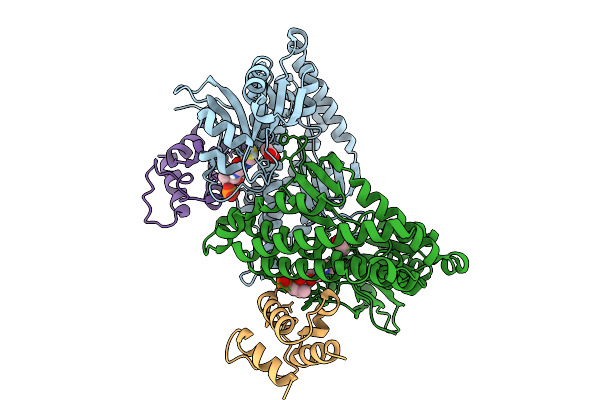 |
Organism: Streptomyces caelestis
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2025-10-01 Classification: BIOSYNTHETIC PROTEIN Ligands: ACT, PNS |
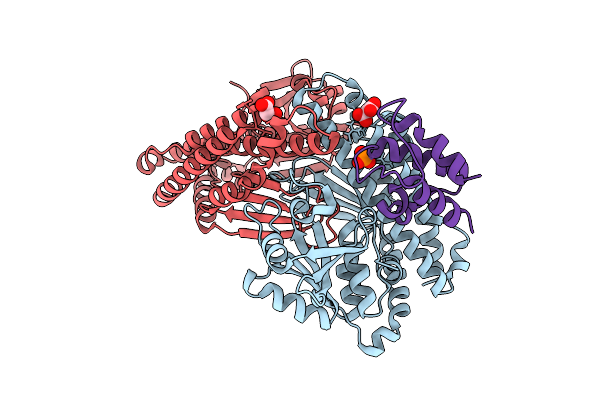 |
Crystal Structure Of Ccbd C19S Mutant Complexed With Single Pcp Domain Of Ccbz
Organism: Streptomyces caelestis
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2025-09-17 Classification: BIOSYNTHETIC PROTEIN Ligands: MLI, PNS |
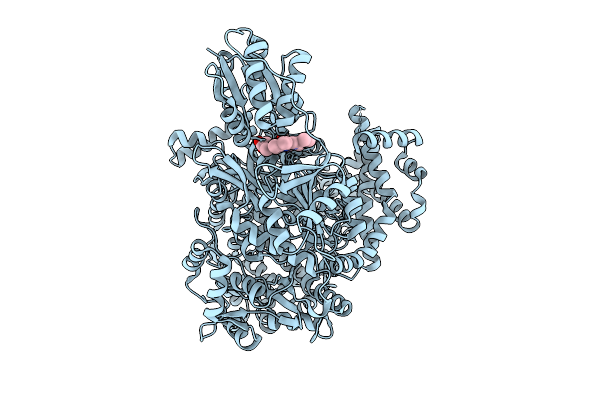 |
Organism: Bacteroides
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: METAL TRANSPORT Ligands: HEM, NA |
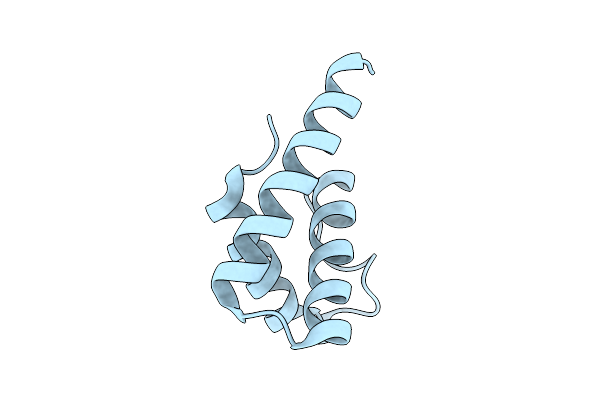 |
Distinct Quaternary States, Intermediates, And Autoinhibition During Loading Of The Dnab-Replicative Helicase By The Phage Lambda P Helicase Loader
Organism: Escherichia phage lambda
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2025-07-16 Classification: REPLICATION |
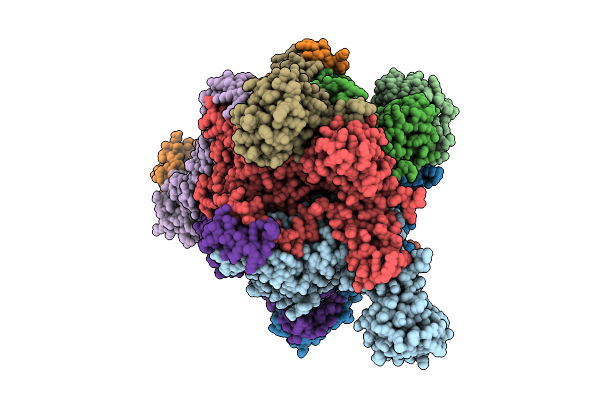 |
Ecoli Dnab Helicase And Phage Lambda Loader P With Adp-Mg In A 6:5 Stoichiometry Ratio.
Organism: Escherichia coli, Escherichia phage lambda
Method: ELECTRON MICROSCOPY Release Date: 2025-07-16 Classification: DNA BINDING PROTEIN Ligands: ADP, MG |
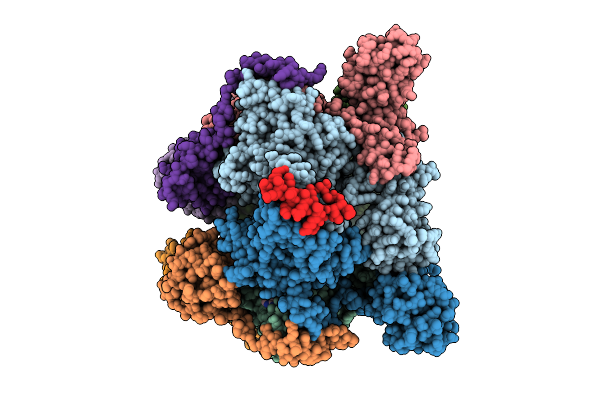 |
Ecoli Dnab Helicase And Phage Lambda Loader P With Adp-Mg In A 6:6 Stoichiometry Ratio.
Organism: Escherichia coli, Escherichia phage lambda
Method: ELECTRON MICROSCOPY Release Date: 2025-07-16 Classification: DNA BINDING PROTEIN Ligands: ADP, MG |
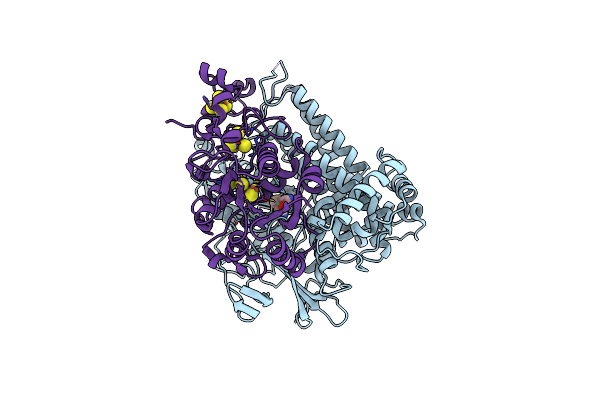 |
Crystal Structure Of The C120G Variant Of The Membrane-Bound [Nife]-Hydrogenase From Cupriavidus Necator In The Air-Oxidized State At 1.93 A Resolution.
Organism: Cupriavidus necator h16
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2025-02-26 Classification: OXIDOREDUCTASE Ligands: NFV, MG, SF4, F3S, 35L, SF3, CL |
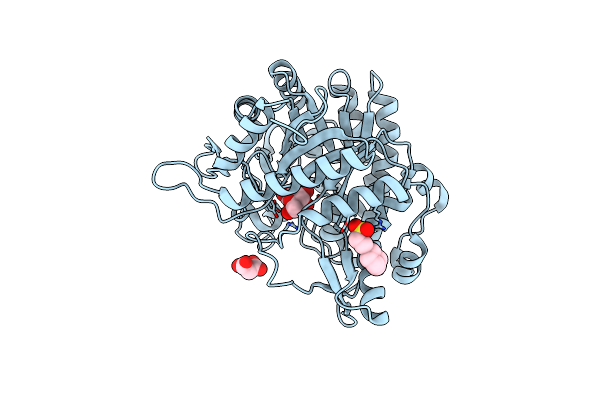 |
Organism: Metagenomes
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-01-15 Classification: HYDROLASE Ligands: MLA, BGC, NHE |
 |
Organism: Metagenomes
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2025-01-15 Classification: HYDROLASE Ligands: NHE, NA, MLA |

