Search Count: 2,577
All
Selected
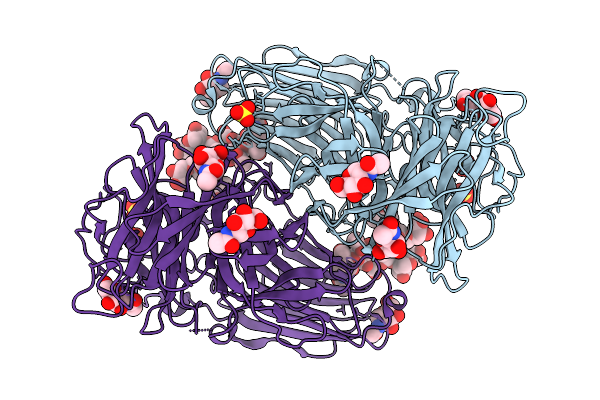 |
Organism: Komagataella phaffii cbs 7435
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: SO4, NAG |
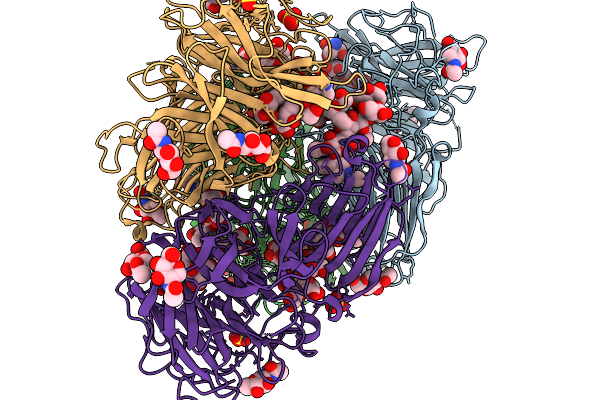 |
Structure Of D81A-Fructofuranosidase From Purpureocilum Lilacinum In Complex With Raffinose
Organism: Komagataella phaffii cbs 7435
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: FRU, TRS, NAG, GLC, GLA, SO4 |
 |
Structure Of D81A-Fructofuranosidase From Purpureocilum Lilacinum In Complex With Fructose
Organism: Komagataella phaffii cbs 7435
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: FRU, NAG, SO4 |
 |
Structure Of D81A-Fructofuranosidase From Purpureocilum Lilacinum In Complex With Sucrose
Organism: Komagataella phaffii cbs 7435
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: GLC, FRU, SO4, TRS, NAG, GOL |
 |
Organism: Adeno-associated virus 2, Adeno-associated virus 8
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: VIRUS Ligands: ADP, MG |
 |
Organism: Adeno-associated virus 8
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: VIRUS Ligands: ADP, MG |
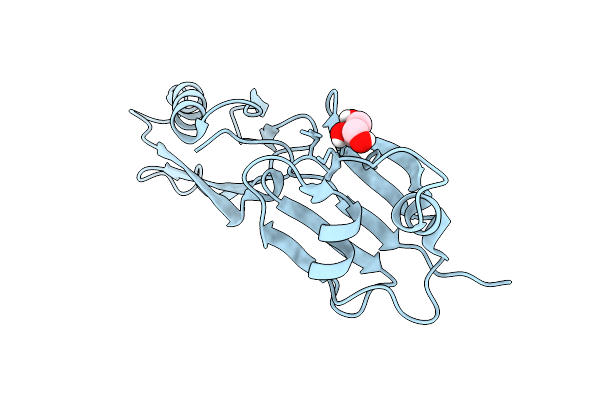 |
Organism: Myxococcus xanthus dz2
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-01-28 Classification: TRANSLOCASE Ligands: GOL |
 |
Organism: Adeno-associated virus - 8, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-23 Classification: VIRAL PROTEIN/HYDROLASE |
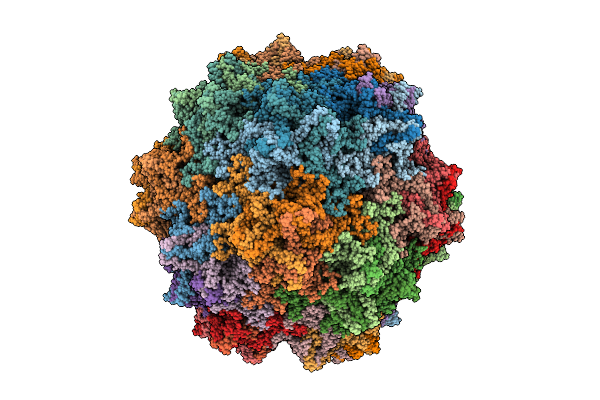 |
Organism: Adeno-associated virus - 8
Method: ELECTRON MICROSCOPY Release Date: 2025-07-23 Classification: VIRUS |
 |
Organism: Adeno-associated virus - 8, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-23 Classification: VIRAL PROTEIN/HYDROLASE |
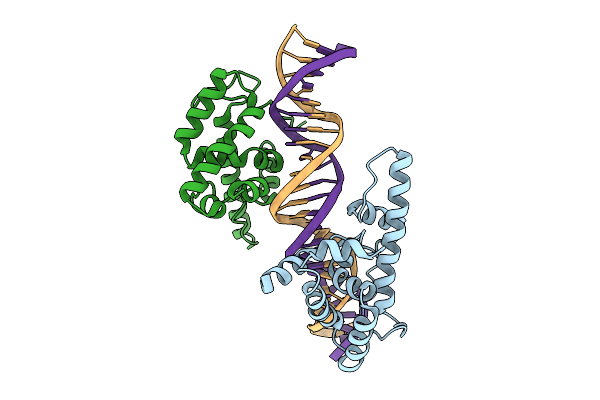 |
Structure Of The Nothoceros Aenigmaticus Lfy Dna-Binding Domain Bound To Dna
Organism: Nothoceros aenigmaticus
Method: X-RAY DIFFRACTION Resolution:3.09 Å Release Date: 2025-07-16 Classification: GENE REGULATION Ligands: HOH |
 |
Structure Of The Nothoceros Aenigmaticus Lfy Dna-Binding Domain Bound To Dna
Organism: Nothoceros aenigmaticus
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2025-07-09 Classification: DNA BINDING PROTEIN |
 |
Organism: Adeno-associated virus 8
Method: ELECTRON MICROSCOPY Release Date: 2024-11-27 Classification: VIRUS |
 |
Organism: Adeno-associated virus 8, Camelidae
Method: ELECTRON MICROSCOPY Release Date: 2024-11-27 Classification: VIRUS |
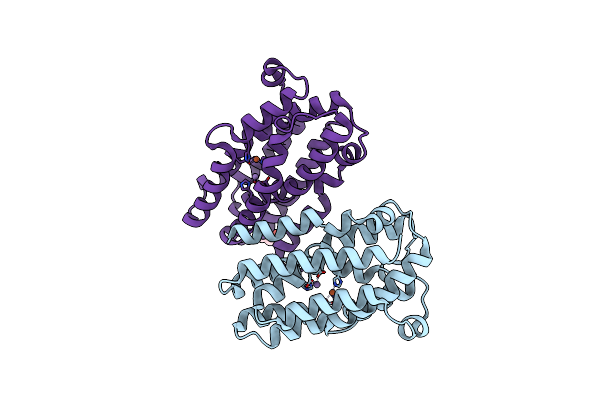 |
Organism: Chlamydia trachomatis
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2024-11-06 Classification: OXIDOREDUCTASE Ligands: FE2, MN, GOL |
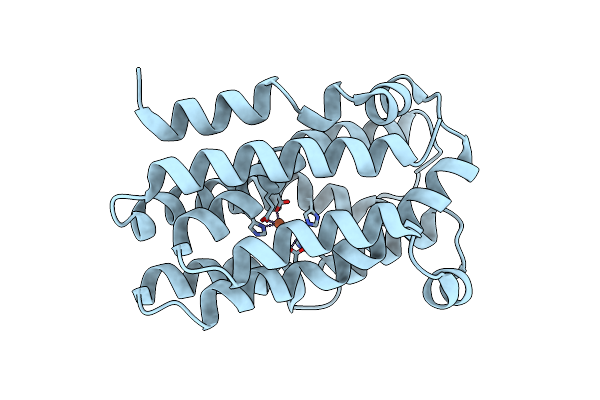 |
Organism: Chlamydia trachomatis
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2024-11-06 Classification: OXIDOREDUCTASE Ligands: FE2 |
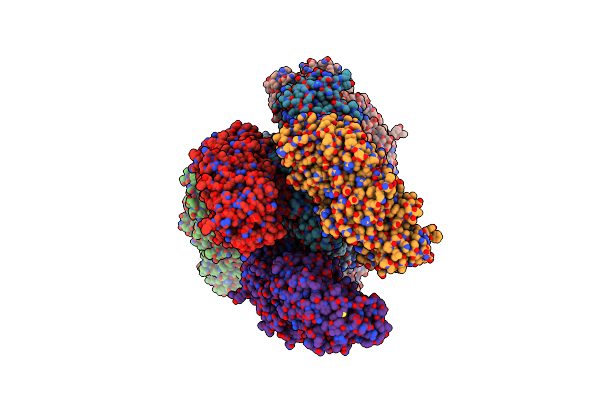 |
Crystal Structure Of 1,4-Alpha-Glucan Branching Protein From Rhodothermus Profundi
Organism: Rhodothermus profundi
Method: X-RAY DIFFRACTION Resolution:2.91 Å Release Date: 2024-10-02 Classification: BIOSYNTHETIC PROTEIN |
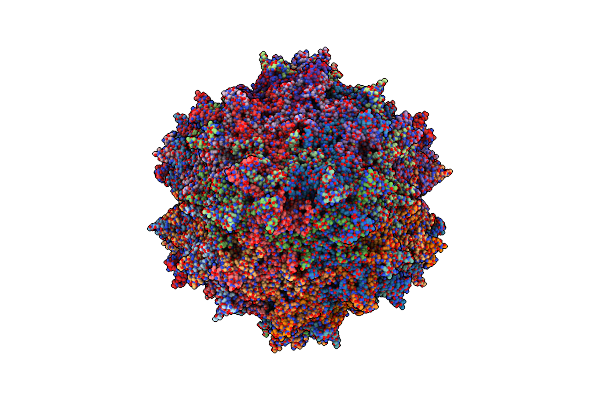 |
Organism: Adeno-associated virus - 8
Method: ELECTRON MICROSCOPY Release Date: 2024-04-24 Classification: VIRUS |
 |
Organism: Haemophilus influenzae 86-028np
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2024-01-17 Classification: PEPTIDE BINDING PROTEIN |
 |
Organism: Chlamydia trachomatis
Method: X-RAY DIFFRACTION Resolution:2.66 Å Release Date: 2023-08-09 Classification: HYDROLASE/INHIBITOR Ligands: UER, NA, GOL |

