Search Count: 1,557
All
Selected
 |
Organism: Bacteroidales, Bacteroidales bacterium
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: IMMUNE SYSTEM/RNA Ligands: MG, ZN |
 |
Organism: Lacticaseibacillus rhamnosus
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2025-12-24 Classification: LYASE |
 |
Organism: Lacticaseibacillus rhamnosus
Method: X-RAY DIFFRACTION Resolution:2.64 Å Release Date: 2025-12-24 Classification: LYASE |
 |
Organism: Lacticaseibacillus rhamnosus
Method: X-RAY DIFFRACTION Resolution:2.69 Å Release Date: 2025-12-24 Classification: LYASE Ligands: PLP |
 |
Organism: Lacticaseibacillus rhamnosus
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2025-12-24 Classification: LYASE |
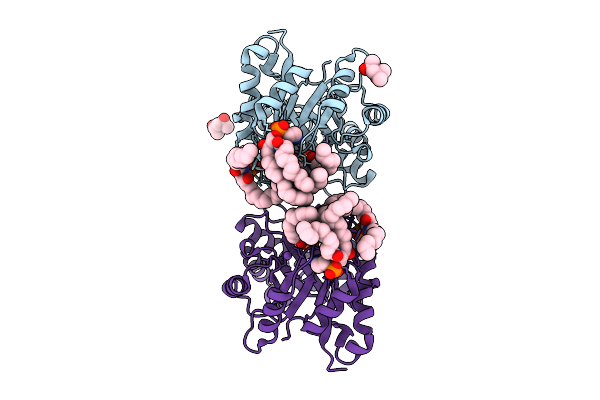 |
Structure Of Phospholipase D Betaib1I From Sicarius Terrosus Venom, H47N Mutant Bound To Product And Substrate Sphingolipids At 2.2 A Resolution From A 2-Day Old Crystal
Organism: Sicarius terrosus
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-12-03 Classification: LYASE Ligands: A1A43, A1A44, NA, MG, MPD |
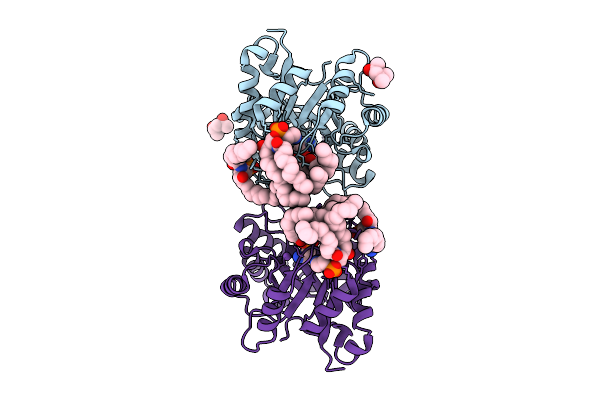 |
Structure Of Phospholipase D Betaib1I From Sicarius Terrosus Venom, H47N Mutant Bound To Substrate Sphingolipids At 2.60 A Resolution
Organism: Sicarius terrosus
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2025-12-03 Classification: LYASE Ligands: A1A43, NA, MG, MPD |
 |
Crystal Structure Of The Cyp154C5 F92A/R114A/T248D/E282A Variant From Nocardia Farcinica
Organism: Nocardia farcinica ifm 10152
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2025-11-05 Classification: OXIDOREDUCTASE Ligands: TES, HEM, CD, CL, K, GOL |
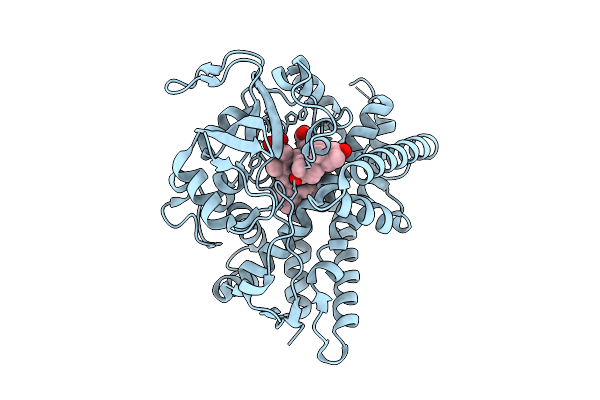 |
Crystal Structure Of The Cyp154C5 F92A/R114A/T248G/E282A Variant From Nocardia Farcinica
Organism: Nocardia farcinica ifm 10152
Method: X-RAY DIFFRACTION Resolution:1.89 Å Release Date: 2025-10-22 Classification: OXIDOREDUCTASE Ligands: HEM, TES |
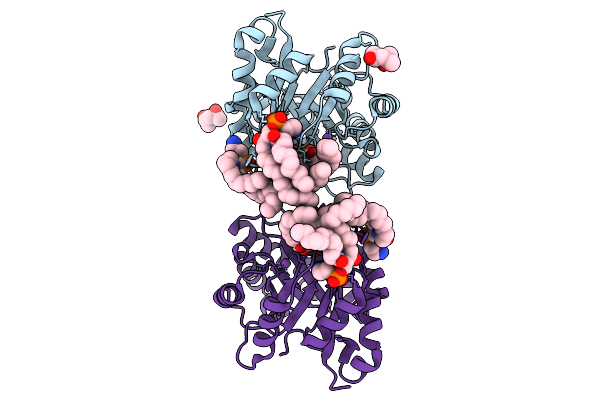 |
Structure Of Phospholipase D Betaib1I From Sicarius Terrosus Venom, H47N Mutant Bound To Product And Substrate Sphingolipids At 1.85 A Resolution
Organism: Sicarius terrosus
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2025-09-10 Classification: LYASE Ligands: A1A43, A1A44, NA, MG, MPD |
 |
Organism: Dissulfurispira thermophila
Method: ELECTRON MICROSCOPY Release Date: 2025-08-27 Classification: IMMUNE SYSTEM Ligands: ZN |
 |
Organism: Aedes aegypti
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-07-02 Classification: OXIDOREDUCTASE Ligands: CO, A1L7D |
 |
Organism: Aedes aegypti
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2025-04-16 Classification: UNKNOWN FUNCTION Ligands: MLT |
 |
Organism: Aedes aegypti
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2025-01-15 Classification: OXIDOREDUCTASE Ligands: CO, A1LW3 |
 |
Organism: Acinetobacter pittii phea-2
Method: X-RAY DIFFRACTION Resolution:2.74 Å Release Date: 2024-10-02 Classification: HYDROLASE Ligands: CA |
 |
Organism: Candidatus nitrospira inopinata
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2024-08-14 Classification: METAL BINDING PROTEIN Ligands: SO4, MN, NI |
 |
Capsular Polysaccharide Synthesis Multienzyme Of Actinobacillus Pleuropneumoniae Serotype 3
Organism: Actinobacillus pleuropneumoniae
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2024-07-03 Classification: BIOSYNTHETIC PROTEIN Ligands: SO4, ZN |
 |
Organism: Aedes aegypti, Apocrypta bakeri
Method: ELECTRON MICROSCOPY Release Date: 2024-06-19 Classification: MEMBRANE PROTEIN |
 |
Organism: Aedes aegypti, Apocrypta bakeri
Method: ELECTRON MICROSCOPY Release Date: 2024-06-19 Classification: MEMBRANE PROTEIN Ligands: JZ0 |
 |
Organism: Ensifer aridi
Method: X-RAY DIFFRACTION Resolution:1.36 Å Release Date: 2024-06-12 Classification: ANTIVIRAL PROTEIN Ligands: ZN |

