Search Count: 5,480
All
Selected
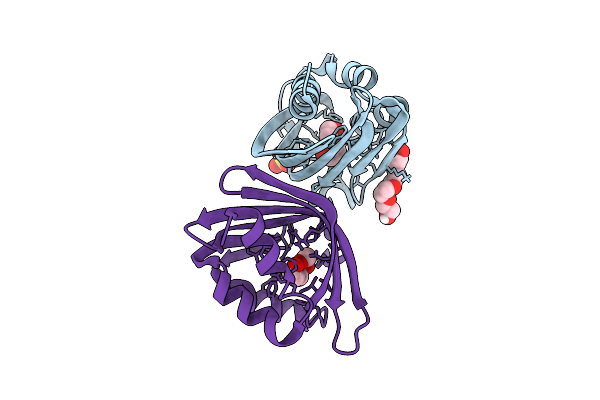 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2026-03-18 Classification: HYDROLASE Ligands: PG4, PGE, SO4 |
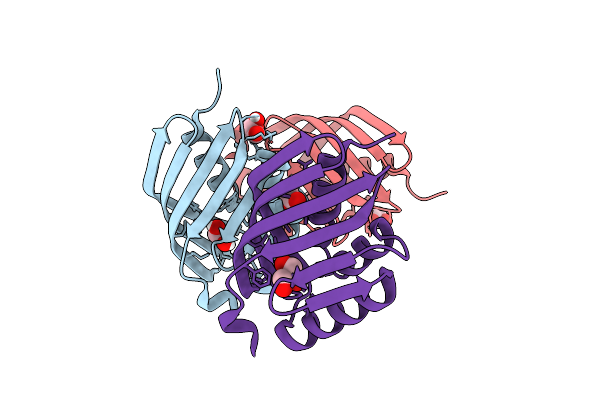 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-03-18 Classification: HYDROLASE Ligands: EDO, GOL, ACY |
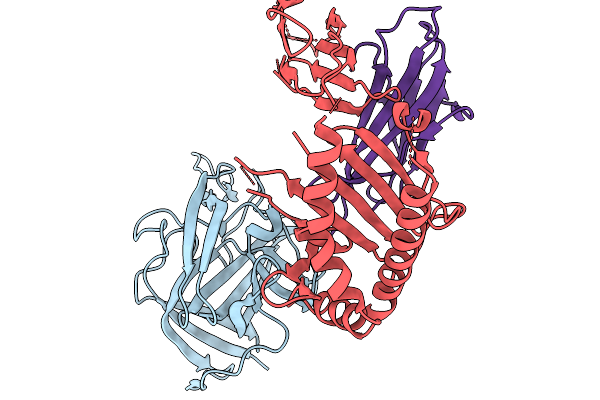 |
Organism: Homo sapiens, Human astrovirus 6
Method: X-RAY DIFFRACTION Resolution:2.97 Å Release Date: 2025-11-19 Classification: IMMUNE SYSTEM |
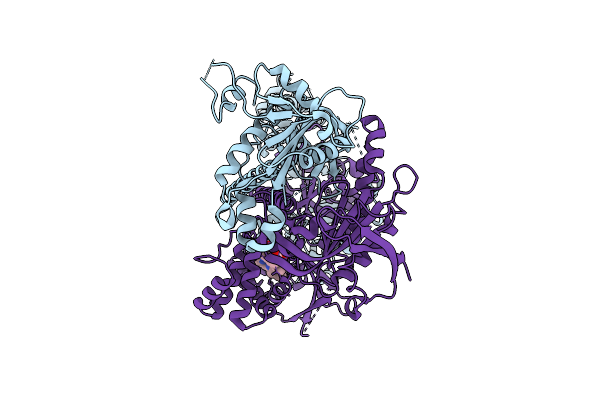 |
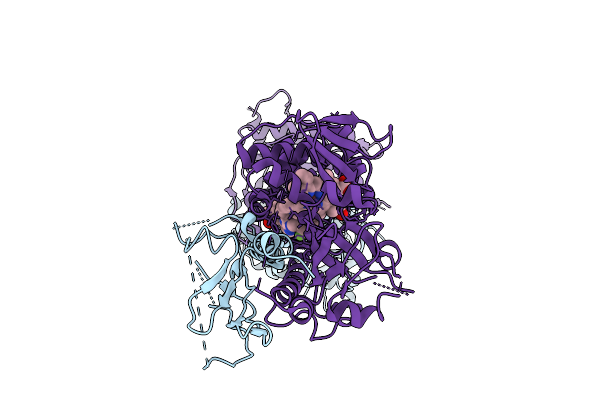
Compact, Ligand-Free State Of Manduca Sexta Soluble Guanylate Cyclase Mutant Beta C122S
Organism: Manduca sexta
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2025-11-12 Classification: SIGNALING PROTEIN Ligands: HEM |
 |
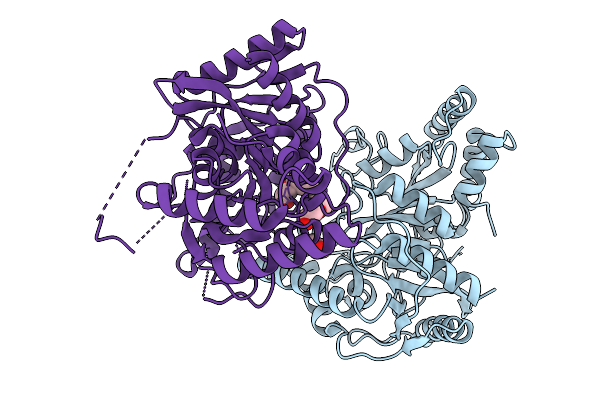
Extended, Cyr715-Bound State Of Manduca Sexta Soluble Guanylate Cyclase Mutant Beta C122S
Organism: Manduca sexta
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: SIGNALING PROTEIN Ligands: G2P, HEM, A1CGK |
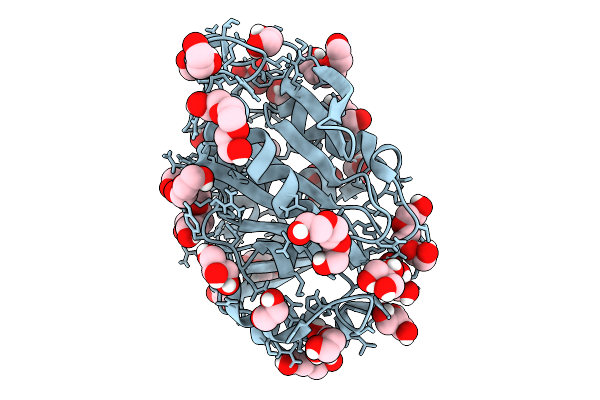 |
Crystal Structure Of Halo-Tolerant Petase From Marine Metagenome (Halopetase1)
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.16 Å Release Date: 2025-10-01 Classification: HYDROLASE Ligands: GOL, PGE, PG4, PEG, EDO |
 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.89 Å Release Date: 2025-09-03 Classification: HYDROLASE Ligands: GOL, P6G |
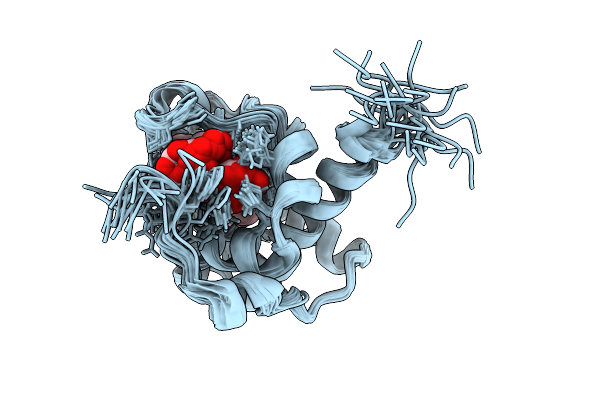 |
Nmr Solution Structure Of Termini-Truncated Oxidized Cytochrome C552 From Thioalkalivibrio Paradoxus
Organism: Thioalkalivibrio paradoxus arh 1
Method: SOLUTION NMR Release Date: 2025-07-30 Classification: ELECTRON TRANSPORT Ligands: HEC |
 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2025-07-02 Classification: HYDROLASE |
 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-07-02 Classification: HYDROLASE Ligands: CL |
 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2025-07-02 Classification: HYDROLASE Ligands: CL |
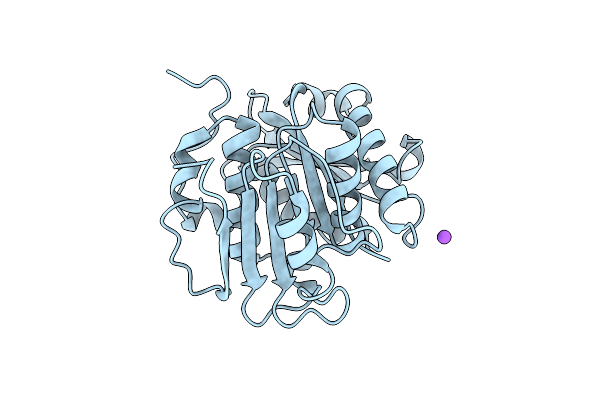 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-07-02 Classification: HYDROLASE Ligands: CL, NA |
 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2025-07-02 Classification: HYDROLASE Ligands: EDO, NA |
 |
Crystal Structure Of Leaf Branch Compost Cutinase Quintuple Variant Iccg L50Y
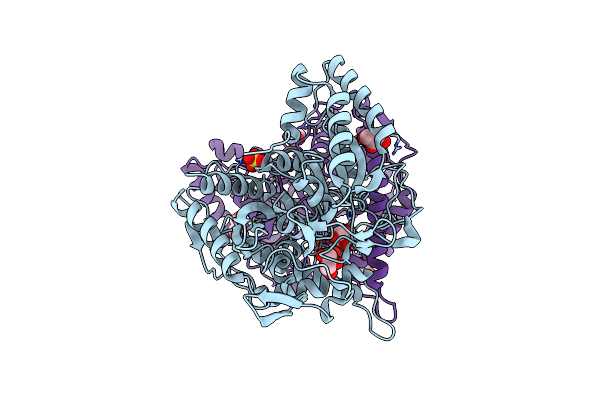
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.51 Å Release Date: 2025-07-02 Classification: HYDROLASE Ligands: EDO, CL |
 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:2.59 Å Release Date: 2025-05-21 Classification: HYDROLASE Ligands: GOL, SO4 |
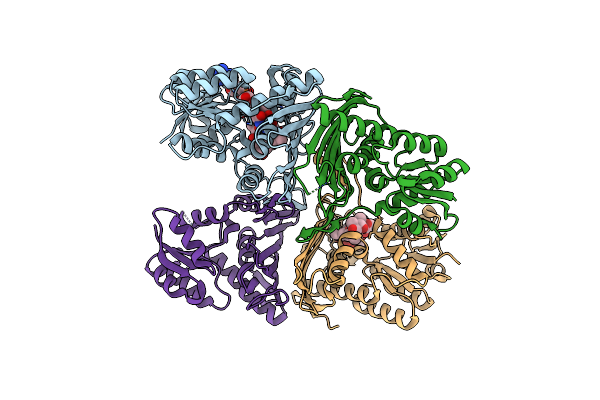 |
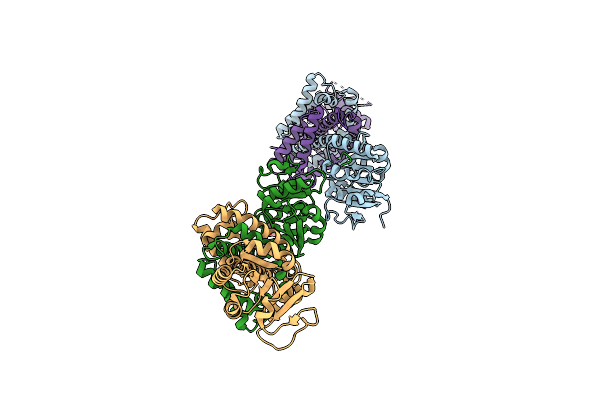
Organism: Amycolatopsis mediterranei s699
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2025-03-12 Classification: OXIDOREDUCTASE Ligands: NAD, US6 |
 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2025-03-05 Classification: BIOSYNTHETIC PROTEIN Ligands: IOD, CL, GOL, EDO |
 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:2.32 Å Release Date: 2025-02-05 Classification: HYDROLASE Ligands: PG6, SO4 |
 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:2.18 Å Release Date: 2025-02-05 Classification: HYDROLASE Ligands: SO4, PMS |
 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-02-05 Classification: HYDROLASE Ligands: ACT, A1IHK |

