Search Count: 2,755
All
Selected
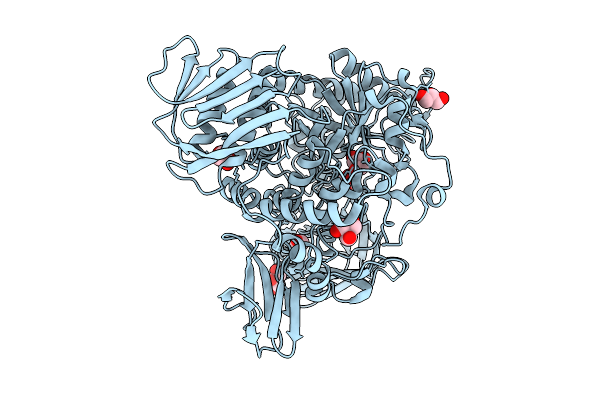 |
Sdacs: A Synergistic Strategy Using Both Crystallographic And Alphafold Structures To Improve Thermostability And Activity Of N-Acetylhexosaminidase
Organism: Cellvibrio mixtus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: TRS, GOL |
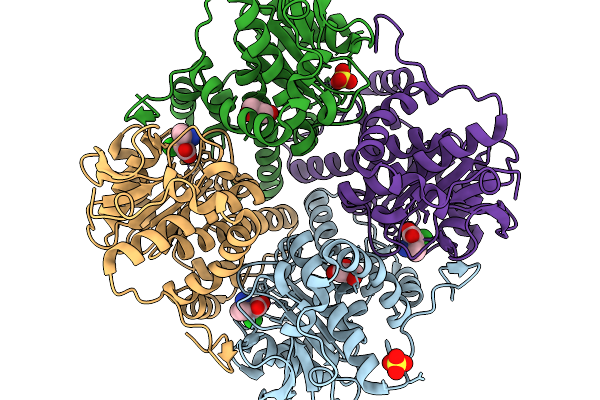 |
Organism: Penicillium aethiopicum
Method: X-RAY DIFFRACTION Resolution:2.17 Å Release Date: 2026-03-04 Classification: METAL BINDING PROTEIN Ligands: A1L7T, GOL, MN, CL, SO4 |
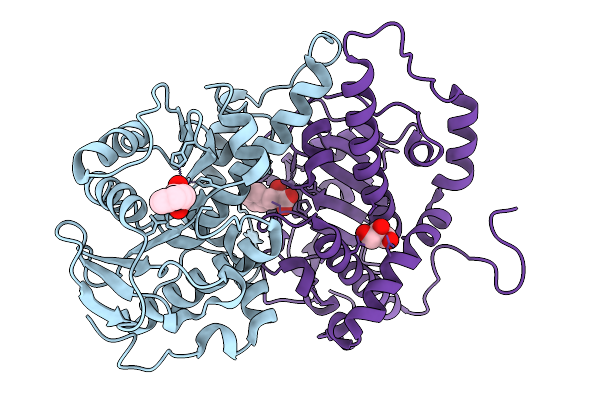 |
Structure Of Tqam From Penicillium Aethiopicum In Complex With D-(+)-3-Phenyllactic Acid
Organism: Penicillium aethiopicum
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2026-03-04 Classification: METAL BINDING PROTEIN Ligands: HF2, GOL, MN, CL |
 |
The Cryo-Em Structure Of C. Crescentus Drid-Ssdna-Rnap-Sigma73-Bape Promoter Transcription Activation Complex
Method: ELECTRON MICROSCOPY
Resolution:4.24 Å Release Date: 2026-02-25 Classification: TRANSCRIPTION/DNA Ligands: MG, ZN |
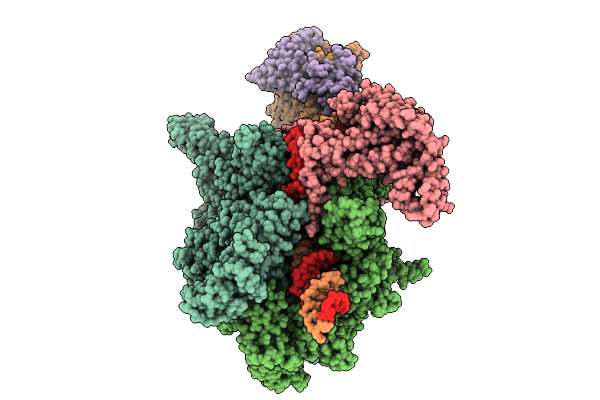 |
The Cryo-Em Structure Of C. Crescentus Drid-Ssdna-Rnap-Sigma73-Ccna_03891/Ccna_01149 Promoter Transcription Activation Complex
Method: ELECTRON MICROSCOPY
Resolution:3.70 Å Release Date: 2026-02-25 Classification: TRANSCRIPTION/DNA Ligands: MG, ZN |
 |
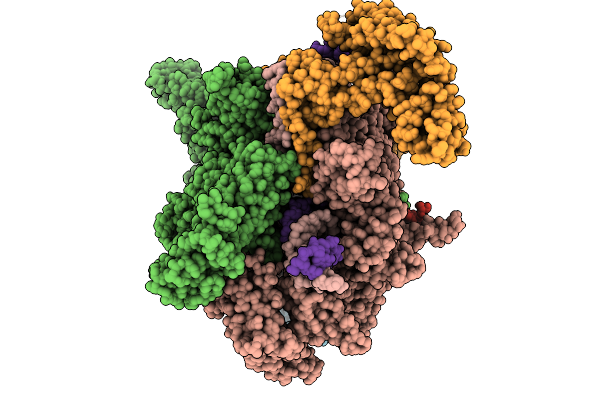
The Cryo-Em Structure Of C. Crescentus Drid-Ssdna-Rnap-Sigma73-Dida Promoter Transcription Activation Complex
Method: ELECTRON MICROSCOPY
Resolution:3.98 Å Release Date: 2026-02-25 Classification: TRANSCRIPTION/DNA Ligands: MG, ZN |
 |
The Cryo-Em Structure Of C. Crescentus Rnap-Sigma73-Ccna_03891/Ccna_01149 Promoter Complex
Method: ELECTRON MICROSCOPY
Resolution:3.54 Å Release Date: 2026-02-25 Classification: TRANSCRIPTION/DNA Ligands: MG, ZN |
 |
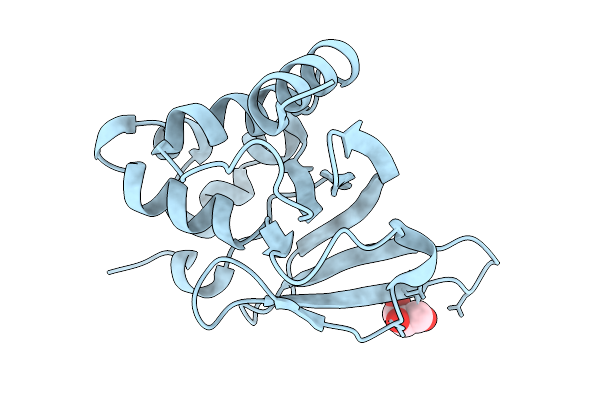
Organism: Geobacillus phage gbsv1
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: VIRAL PROTEIN |
 |
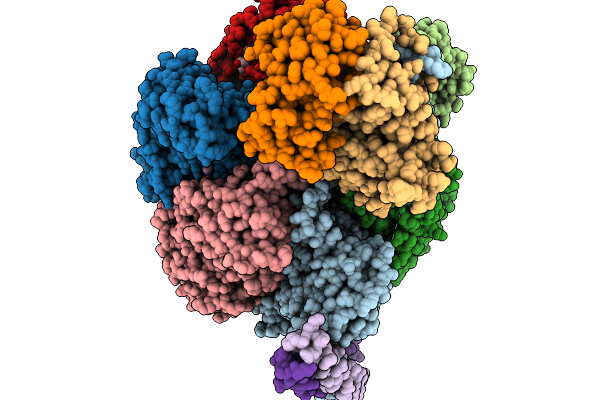
Organism: Scenedesmus sp. oki-4n
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-02-04 Classification: TRANSPORT PROTEIN Ligands: GOL |
 |
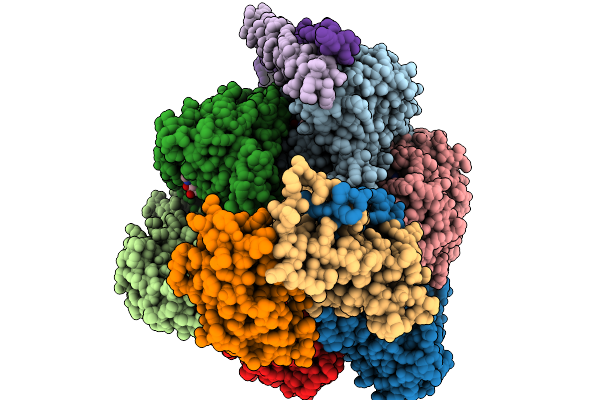
Cryo-Em Structure Of F-Atp Synthase From Mycobacteroides Abscessus (Rotational State 1)
Organism: Mycobacteroides abscessus subsp. abscessus
Method: ELECTRON MICROSCOPY Resolution:2.94 Å Release Date: 2026-01-14 Classification: HYDROLASE Ligands: ATP, MG, ADP |
 |
Cryo-Em Structure Of F-Atp Synthase From Mycobacteroides Abscessus (Rotational State 2)
Organism: Mycobacteroides abscessus subsp. abscessus
Method: ELECTRON MICROSCOPY Resolution:3.41 Å Release Date: 2026-01-14 Classification: HYDROLASE Ligands: ATP, MG, ADP |
 |
Cryo-Em Structure Of F-Atp Synthase From Mycobacteroides Abscessus (Rotational State 3)
Organism: Mycobacteroides abscessus subsp. abscessus
Method: ELECTRON MICROSCOPY Resolution:2.79 Å Release Date: 2026-01-14 Classification: HYDROLASE Ligands: ATP, MG, ADP |
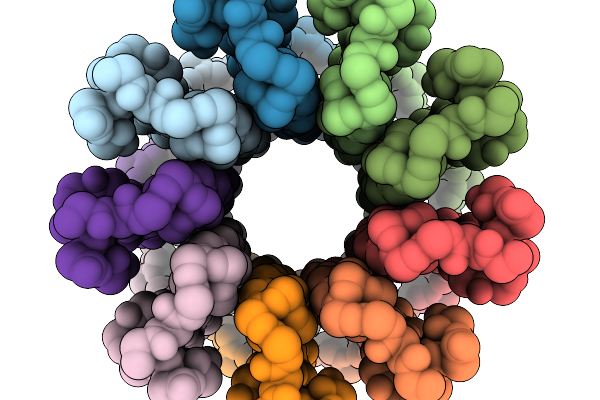 |
Cryo-Em Structure Of F-Atp Synthase C-Ring From Mycobacteroides Abscessus (Backbone)
Organism: Mycobacteroides abscessus subsp. abscessus
Method: ELECTRON MICROSCOPY Resolution:5.61 Å Release Date: 2026-01-14 Classification: HYDROLASE |
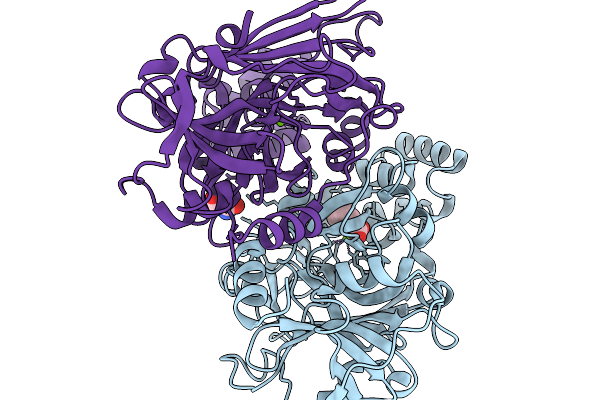 |
Structure-Guided Interface Engineering For Modifying Substrate Binding And Catalytic Activity Of 2-Keto-3-Deoxy-D-Xylonate Dehydratase
Organism: Caulobacter vibrioides na1000
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2025-11-19 Classification: CARBOHYDRATE Ligands: MG, TRS, COI |
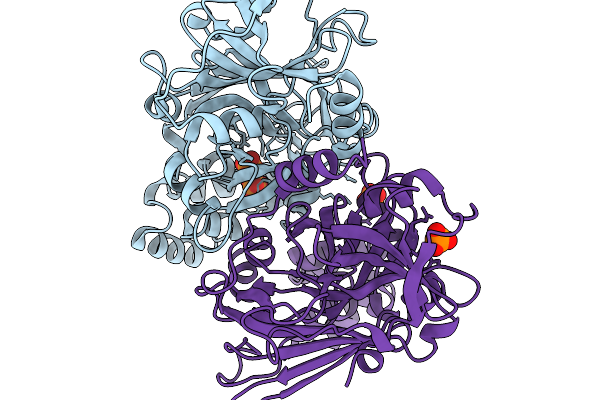 |
Structure-Guided Interface Engineering For Modifying Substrate Binding And Catalytic Activity Of 2-Keto-3-Deoxy-D-Xylonate Dehydratase
Organism: Caulobacter vibrioides na1000
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2025-11-19 Classification: CARBOHYDRATE Ligands: 2HP, NH4 |
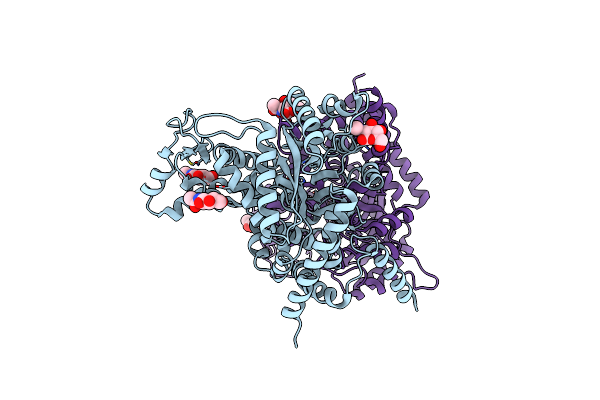 |
Organism: Aplysia californica
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2025-06-25 Classification: HYDROLASE Ligands: NAG, ZN |
 |
Chimeric Adenosine Deaminase Growth Factor (Adgf) In Complex With Pentostatin
Organism: Aplysia californica, Helix pomatia
Method: X-RAY DIFFRACTION Resolution:3.28 Å Release Date: 2025-06-25 Classification: HYDROLASE Ligands: NAG, EDO, ZN, DCF |
 |
Organism: Aplysia californica
Method: ELECTRON MICROSCOPY Release Date: 2025-06-18 Classification: MEMBRANE PROTEIN Ligands: K, MG, CA |
 |
Organism: Aplysia californica
Method: ELECTRON MICROSCOPY Release Date: 2025-06-18 Classification: MEMBRANE PROTEIN Ligands: K |
 |
Organism: Aplysia californica
Method: ELECTRON MICROSCOPY Release Date: 2025-06-18 Classification: MEMBRANE PROTEIN Ligands: K, MG, CA |

