Search Count: 1,643
All
Selected
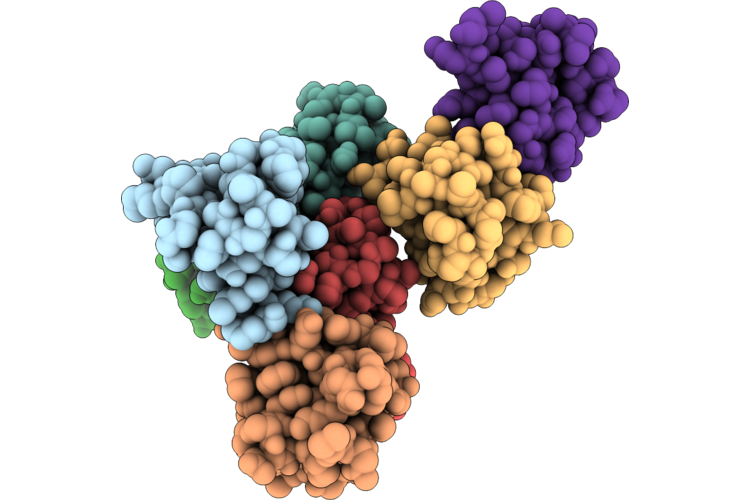 |
Crystal Structure Of Rubredoxin From Psychrophilic Bacterium Polaromonas Glacialis
Organism: Polaromonas glacialis
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2026-05-06 Classification: ELECTRON TRANSPORT Ligands: FE, NA |
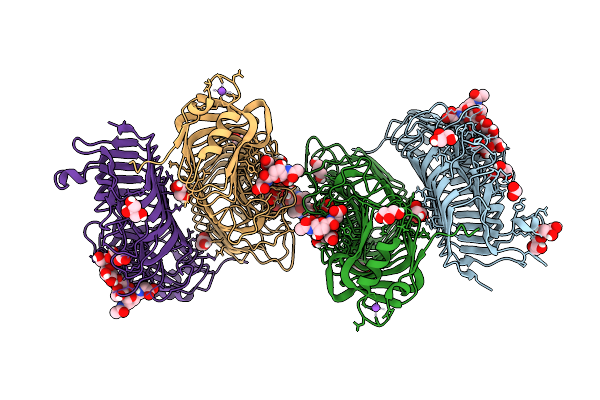 |
Crystal Structure Of The Polysaccharide Lyase Rbmb From Vibrio Cholerae Bound To Vibrio Polysaccharide
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2026-03-11 Classification: LYASE |
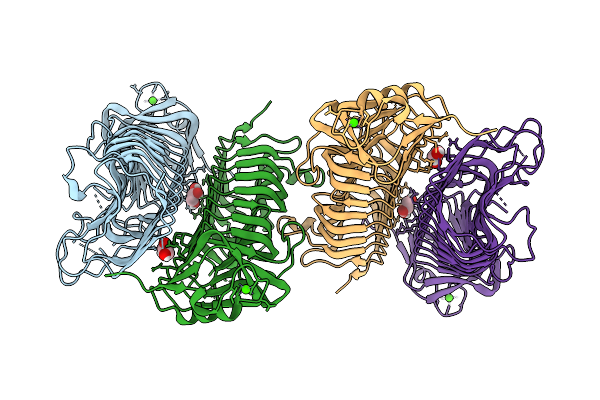 |
Organism: Vibrio cholerae c6706
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-03-11 Classification: LYASE Ligands: GOL, CA |
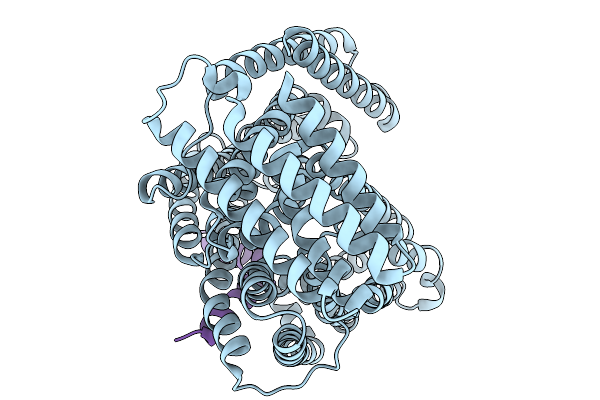 |
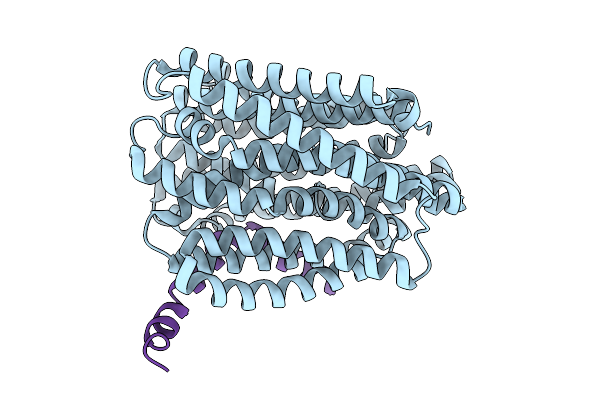
Organism: Escherichia coli k-12, Enterobacteria phage m
Method: ELECTRON MICROSCOPY Resolution:3.60 Å Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN |
 |
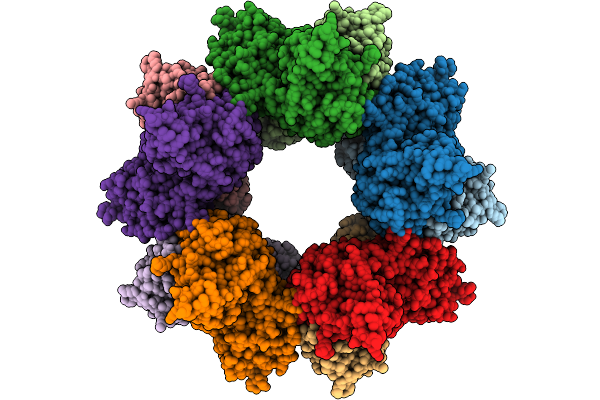
Organism: Escherichia coli, Enterobacteria phage m
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: VIRAL PROTEIN |
 |
Organism: Candidatus methanofastidiosum methylthiophilus
Method: ELECTRON MICROSCOPY Release Date: 2025-08-27 Classification: LYASE Ligands: MG |
 |
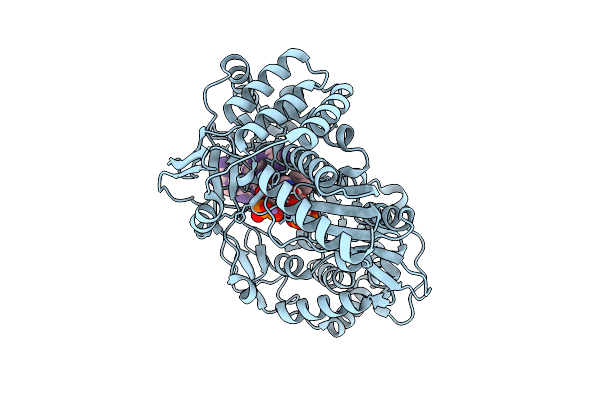
Structural Insights Into The Polymerase Catalyzed Fad-Capping Of Hepatitis C Viral Rna
Organism: Hepatitis c virus jfh-1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.74 Å Release Date: 2025-08-13 Classification: VIRAL PROTEIN Ligands: CDP, FAD, MN |
 |
Structural Insights Into The Polymerase Catalyzed Fad-Capping Of Hepatitis C Viral Rna
Organism: Hepatitis c virus jfh-1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.96 Å Release Date: 2025-08-13 Classification: VIRAL PROTEIN Ligands: FAD, CDP, MN |
 |
Structural Insights Into The Polymerase Catalyzed Fad-Capping Of Hepatitis C Viral Rna
Organism: Hepatitis c virus jfh-1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.46 Å Release Date: 2025-08-13 Classification: VIRAL PROTEIN Ligands: CDP, MN, FAD |
 |
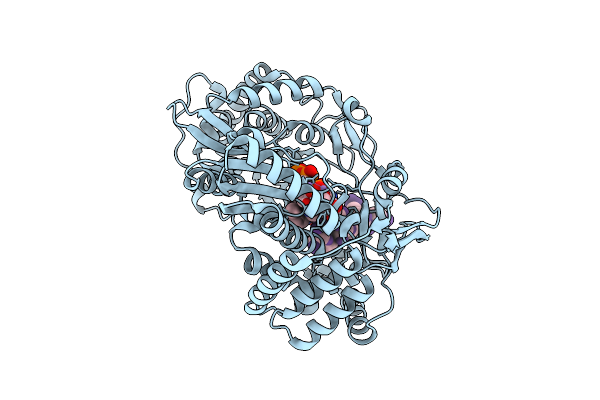
Structural Insights Into The Polymerase Catalyzed Fad-Capping Of Hepatitis C Viral Rna
Organism: Hepatitis c virus jfh-1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.92 Å Release Date: 2025-08-13 Classification: VIRAL PROTEIN Ligands: FAD, GDP, MN |
 |
Structural Insights Into The Polymerase Catalyzed Fad-Capping Of Hepatitis C Viral Rna
Organism: Hepatitis c virus jfh-1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2025-08-13 Classification: VIRAL PROTEIN Ligands: C5P, MN, FAD |
 |
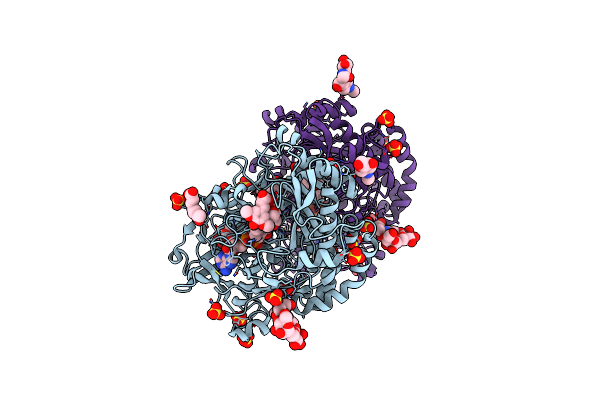
Structural Insights Into The Polymerase Catalyzed Fad-Capping Of Hepatitis C Viral Rna
Organism: Hepatitis c virus genotype 2a (isolate jfh-1), Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.13 Å Release Date: 2025-08-13 Classification: VIRAL PROTEIN Ligands: GDP, MN, FAD |
 |
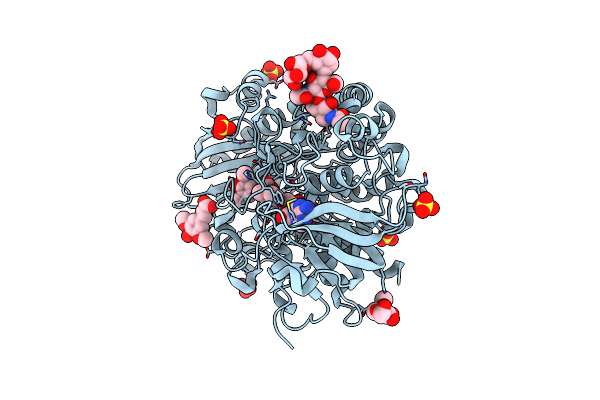
Crystallographic Structure Of Oligosaccharide Dehydrogenase From Pycnoporus Cinnabarinus Bound To Sinapic Acid, Tetragonal Crystal
Organism: Trametes cinnabarina
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2025-07-02 Classification: OXIDOREDUCTASE Ligands: FAD, SXX, SO4, NAG |
 |
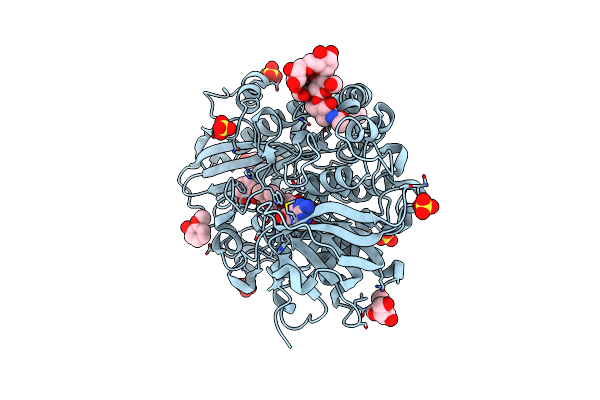
Crystallographic Structure Of Oligosaccharide Dehydrogenase From Pycnoporus Cinnabarinus Bound To Sinapic Acid, Orthorhombic Crystal
Organism: Trametes cinnabarina
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2025-07-02 Classification: OXIDOREDUCTASE Ligands: FAD, SXX, NAG, SO4 |
 |
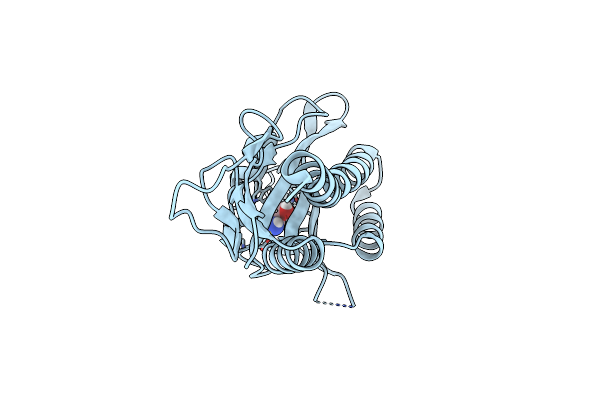
Crystallographic Structure Of Oligosaccharide Dehydrogenase From Pycnoporus Cinnabarinus Bound To Guaiacol, Orthorhombic Crystal
Organism: Trametes cinnabarina
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2025-07-02 Classification: OXIDOREDUCTASE Ligands: FAD, NAG, SO4, JZ3 |
 |
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2025-07-02 Classification: SIGNALING PROTEIN Ligands: CL, ETA |
 |
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2025-07-02 Classification: SIGNALING PROTEIN Ligands: 2A1, CL |
 |
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2025-07-02 Classification: SIGNALING PROTEIN Ligands: SEL, CL |
 |
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-07-02 Classification: SIGNALING PROTEIN |
 |
Organism: Xanthomonas sacchari
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-05-21 Classification: SUGAR BINDING PROTEIN Ligands: EDO, A1BNY |

