Search Count: 2,949
All
Selected
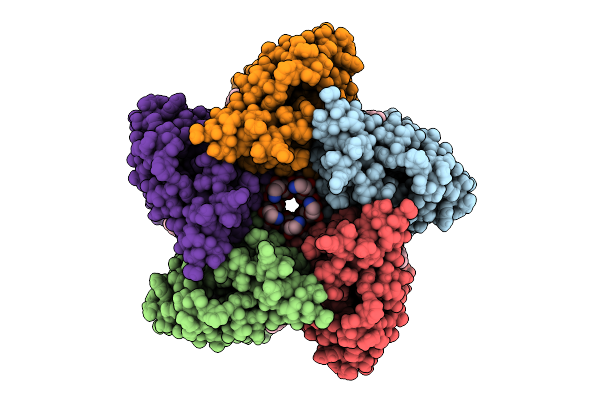 |
Cryo-Em Structure Of The Freshwater Actinorhodopsin, Rhodoluna Lacicola (Rlactr)
Organism: Rhodoluna lacicola
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: PROTON TRANSPORT Ligands: A1INB, A1IVO, A1INC, RET |
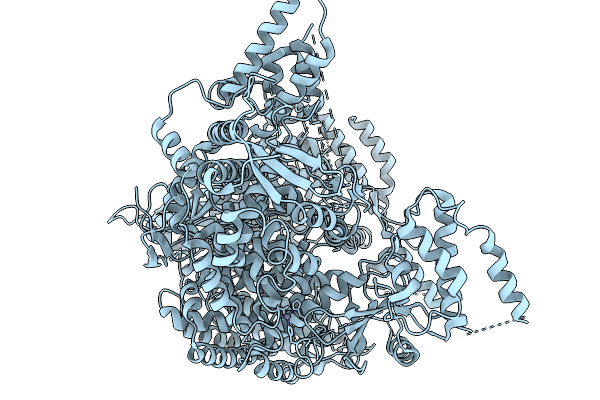 |
2.62A Cryo-Em Structure Of Rna-Directed Rna Polymerase L Of Crimean-Congo Hemorrhagic Fever Virus (Apo State)
Organism: Crimean-congo hemorrhagic fever virus
Method: ELECTRON MICROSCOPY Resolution:2.62 Å Release Date: 2026-03-04 Classification: TRANSFERASE Ligands: ZN |
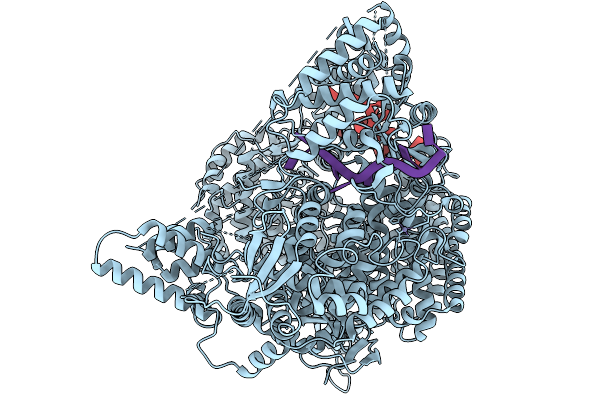 |
2.53A Cryo-Em Structure Of Rna-Directed Rna Polymerase L Of Crimean-Congo Hemorrhagic Fever Virus (Rna Bound)
Organism: Crimean-congo hemorrhagic fever virus, Crimean-congo hemorrhagic fever virus strain ibar10200
Method: ELECTRON MICROSCOPY Resolution:2.53 Å Release Date: 2026-03-04 Classification: TRANSFERASE/RNA Ligands: ZN |
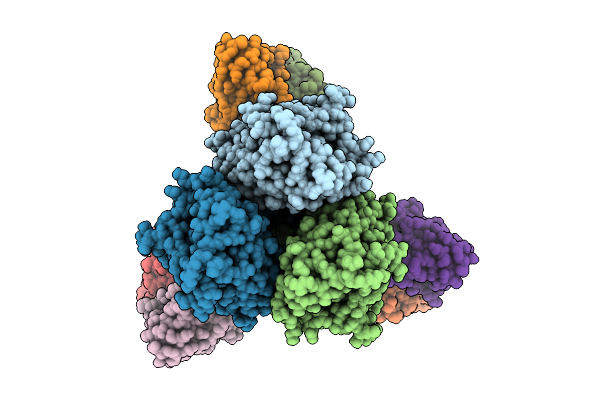 |
Organism: Alphainfluenzavirus influenzae, Macaca mulatta
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: VIRAL PROTEIN |
 |
Rabbit 80S Ribosome In Complex With Erf1-Aaq, Stalled At The Stop Codon In Mutated F2A Sequence
Organism: Homo sapiens, Foot-and-mouth disease virus sat 2, Oryctolagus cuniculus
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: RIBOSOME Ligands: MG, ZN, K, SPD, GTP, SF4 |
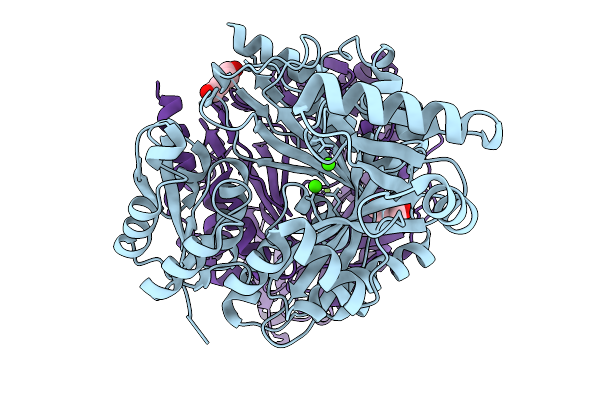 |
Organism: Ankistrodesmus falcatus
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2025-11-19 Classification: LIGASE Ligands: PEG, CA, MG |
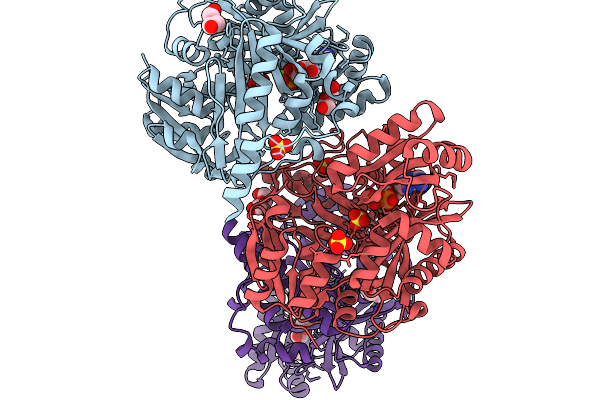 |
Crystal Structure Of Biotin Carboxylase From Ankistrodesmus In Complex With Adp
Organism: Ankistrodesmus falcatus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-11-19 Classification: LIGASE Ligands: MG, ADP, PEG, SO4 |
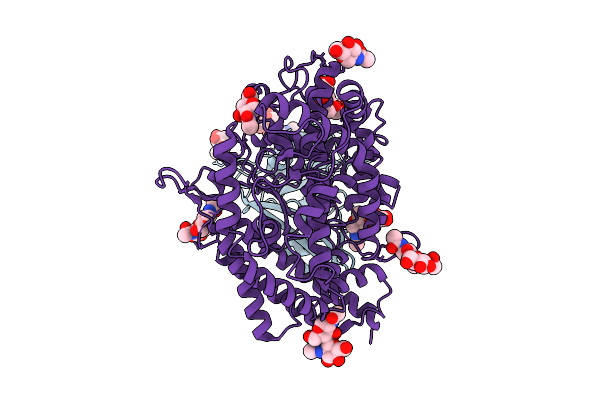 |
Organism: Bat coronavirus hku5, Pipistrellus abramus
Method: ELECTRON MICROSCOPY Release Date: 2025-11-05 Classification: VIRAL PROTEIN Ligands: NAG |
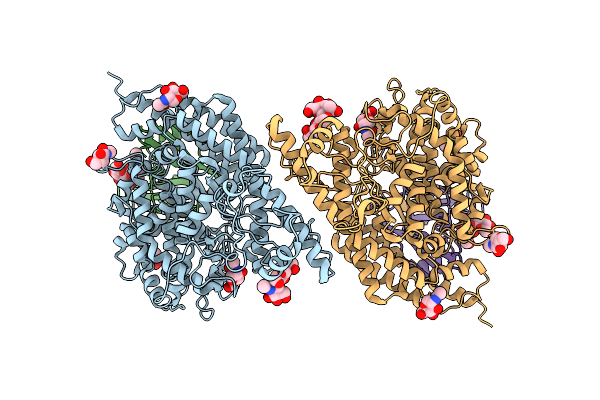 |
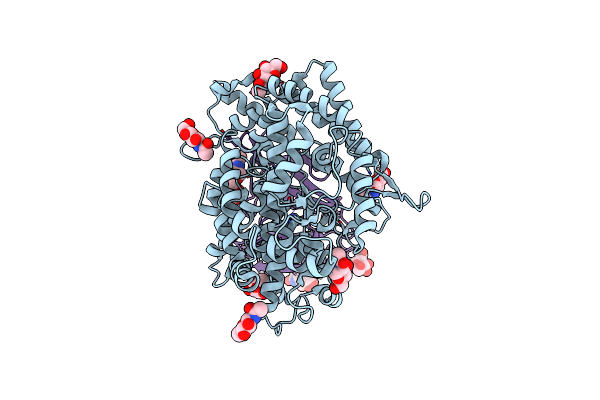
Structure Of Hku5 Spike C-Terminal Domain In Complex With Ace2 From Pipistrellus Abramus
Organism: Pipistrellus abramus, Pipistrellus bat coronavirus hku5
Method: ELECTRON MICROSCOPY Resolution:4.20 Å Release Date: 2025-07-30 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Organism: Pipistrellus abramus, Pipistrellus bat coronavirus hku5
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2025-02-19 Classification: VIRAL PROTEIN/HYDROLASE Ligands: NAG, ZN |
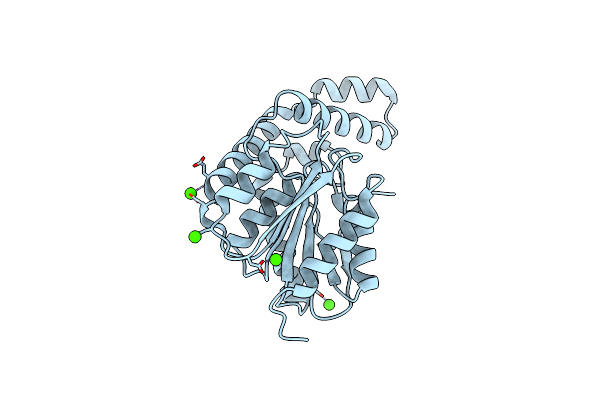 |
Fphh, Staphylococcus Aureus Fluorophosphonate-Binding Serine Hydrolases H, Apo Crystal Form 2
Organism: Staphylococcus aureus usa300-ca-263
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2025-01-08 Classification: HYDROLASE Ligands: CA |
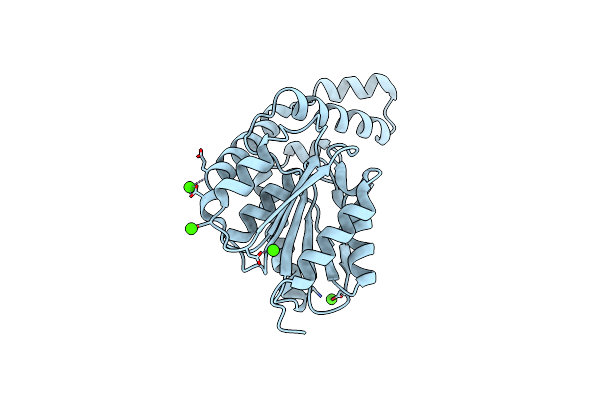 |
Fphh, Staphylococcus Aureus Fluorophosphonate-Binding Serine Hydrolases H, Apo Form 2 At Room Temperature
Organism: Staphylococcus aureus usa300-ca-263
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-01-08 Classification: HYDROLASE Ligands: CA |
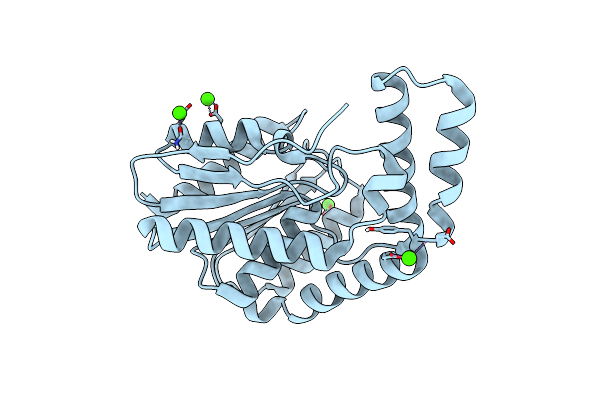 |
Fphi, Staphylococcus Aureus Fluorophosphonate-Binding Serine Hydrolases I, Apo Form At Room Temperature
Organism: Staphylococcus aureus usa300-ca-263
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-01-08 Classification: HYDROLASE Ligands: CA |
 |
Organism: Crimean-congo hemorrhagic fever virus strain ibar10200
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2024-12-11 Classification: VIRAL PROTEIN Ligands: NAG, A2G |
 |
Organism: Crimean-congo hemorrhagic fever virus strain ibar10200, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-12-11 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Organism: Foot-and-mouth disease virus sat 2, Bos taurus
Method: ELECTRON MICROSCOPY Release Date: 2024-10-30 Classification: VIRUS LIKE PARTICLE |
 |
Organism: Escherichia coli k-12, Guillardia theta
Method: ELECTRON MICROSCOPY Release Date: 2024-09-11 Classification: RNA BINDING PROTEIN/RNA/DNA Ligands: ZN |
 |
Organism: Escherichia coli k-12, Guillardia theta, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2024-09-11 Classification: RNA BINDING PROTEIN/RNA/DNA Ligands: ZN |
 |
Organism: Escherichia coli k-12, Guillardia theta, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2024-09-11 Classification: RNA BINDING PROTEIN/RNA/DNA Ligands: ZN |
 |
Organism: Guillardia theta
Method: ELECTRON MICROSCOPY Resolution:2.71 Å Release Date: 2024-09-04 Classification: MEMBRANE PROTEIN Ligands: RET, PC1 |

