Search Count: 1,975
 |
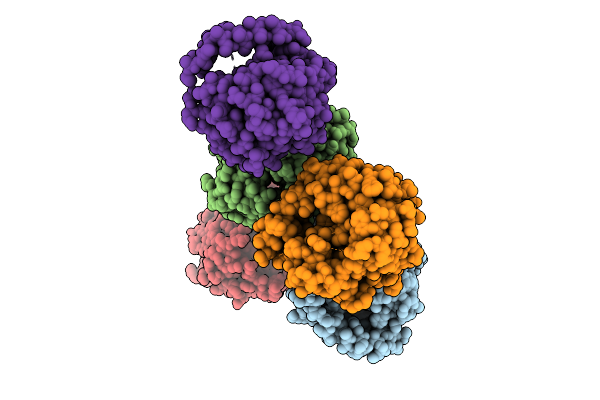 |
Crystal Structure Of Apo Alpha/Beta-Hydrolase Macrolide Esterase Estt From Sphingobacterium Faecium (S126A Mutant)
Organism: Sphingobacterium faecium
Method: X-RAY DIFFRACTION Resolution:3.19 Å Release Date: 2026-04-08 Classification: HYDROLASE |
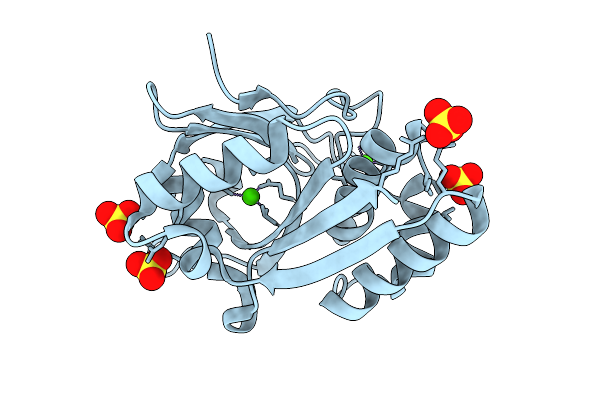 |
Organism: Metagenomes
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-18 Classification: LYASE Ligands: CA, SO4 |
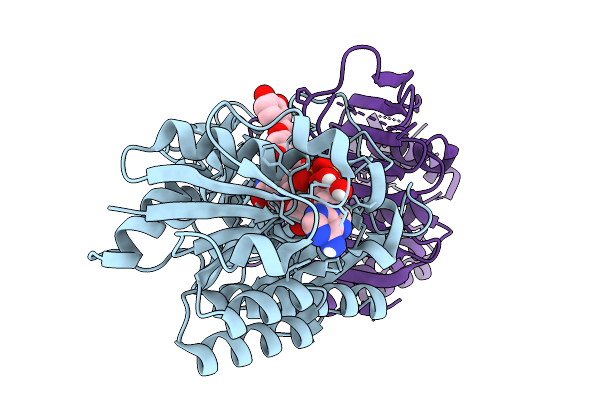 |
Organism: Human gut metagenome
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-03-04 Classification: ISOMERASE Ligands: H9R, NAD |
 |
Organism: Human gut metagenome
Method: X-RAY DIFFRACTION Resolution:1.92 Å Release Date: 2026-03-04 Classification: ISOMERASE Ligands: A1EMQ, NAD |
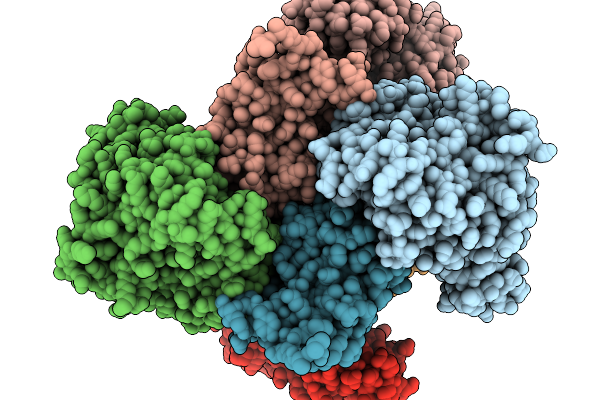 |
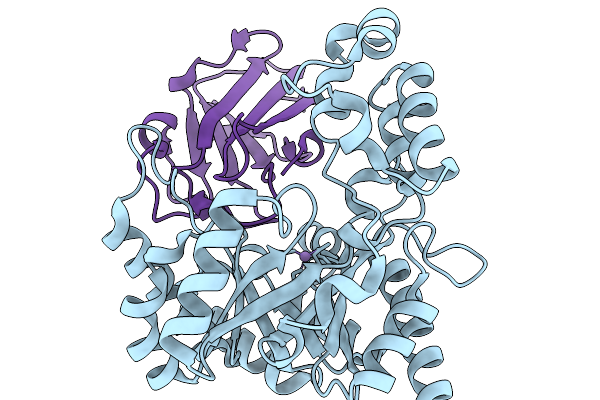
Crystal Structure Of The Dgpb2/C2 Complex From W974-1 In Substrate Free Form
Organism: Human gut metagenome
Method: X-RAY DIFFRACTION Resolution:2.23 Å Release Date: 2026-03-04 Classification: ISOMERASE Ligands: MN |
 |
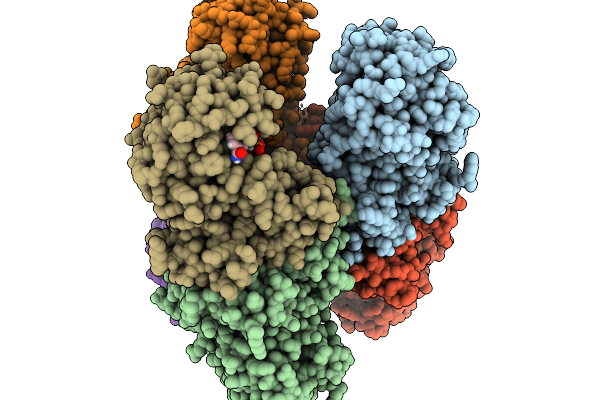
Organism: Human gut metagenome
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2026-03-04 Classification: ISOMERASE Ligands: MN |
 |
Organism: Human gut metagenome
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-04 Classification: ISOMERASE Ligands: NAD |
 |
Organism: Cervus canadensis, Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: VIRAL PROTEIN Ligands: NAG |
 |
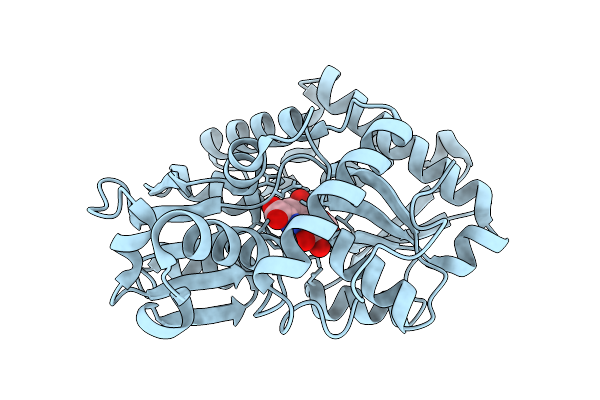
Organism: Severe acute respiratory syndrome coronavirus 2, Cervus canadensis
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: VIRAL PROTEIN Ligands: NAG |
 |
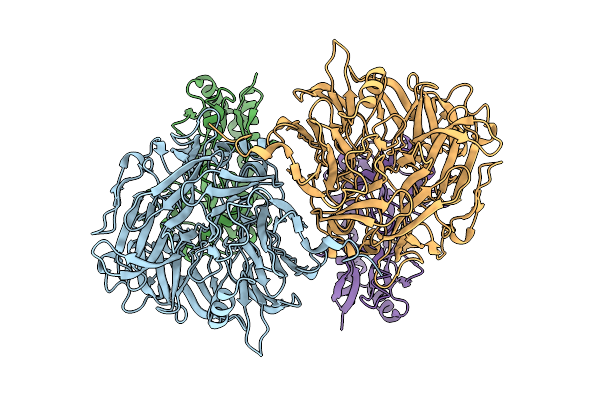
Organism: Methylorubrum extorquens
Method: X-RAY DIFFRACTION Resolution:1.98 Å Release Date: 2025-10-15 Classification: TRANSPORT PROTEIN Ligands: PQQ, CL |
 |
Organism: Methylorubrum extorquens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-23 Classification: PROTEIN BINDING |
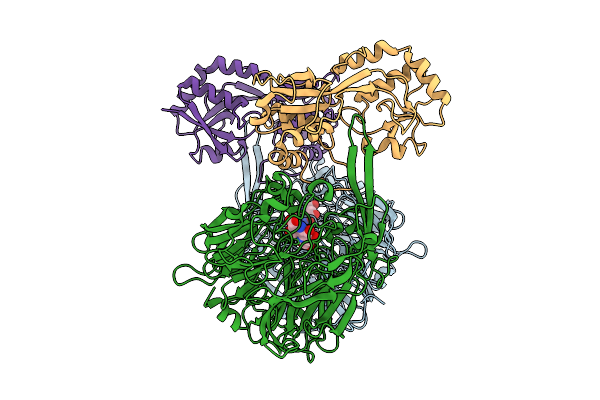 |
Organism: Methylorubrum extorquens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-23 Classification: PROTEIN BINDING Ligands: PQQ |
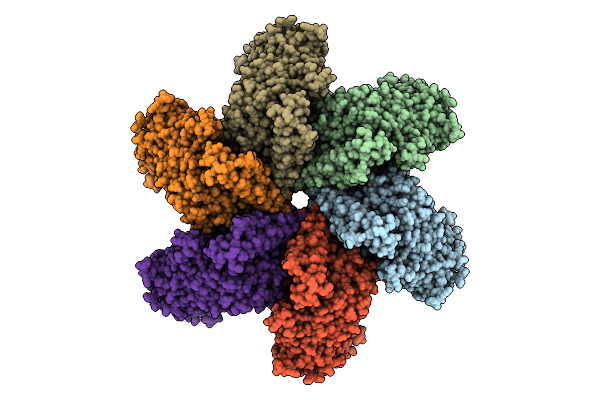 |
Organism: Solanum lycopersicum
Method: ELECTRON MICROSCOPY Release Date: 2025-07-23 Classification: IMMUNE SYSTEM Ligands: ATP |
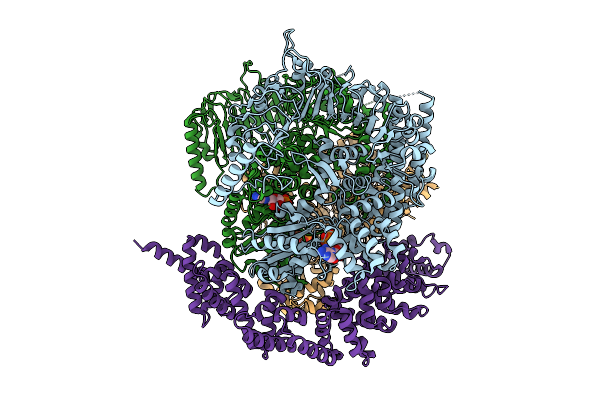 |
Organism: Solanum lycopersicum, Phytophthora infestans
Method: ELECTRON MICROSCOPY Release Date: 2025-07-23 Classification: IMMUNE SYSTEM Ligands: ATP |
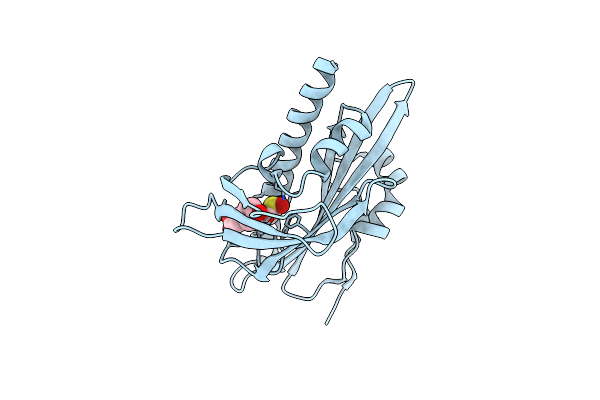 |
Organism: Solanum lycopersicum
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2025-07-02 Classification: PLANT PROTEIN Ligands: HC4, DMS |
 |
Organism: Solanum lycopersicum
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-07-02 Classification: PLANT PROTEIN Ligands: FER, DMS |
 |
Organism: Solanum lycopersicum
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2025-07-02 Classification: PLANT PROTEIN Ligands: DHC, DMS, SO4 |
 |
Organism: Solanum lycopersicum
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2025-07-02 Classification: PLANT PROTEIN Ligands: A1I70, DMS |
 |
Organism: Riemerella anatipestifer
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2025-06-25 Classification: TRANSPORT PROTEIN |

