Search Count: 2,783
All
Selected
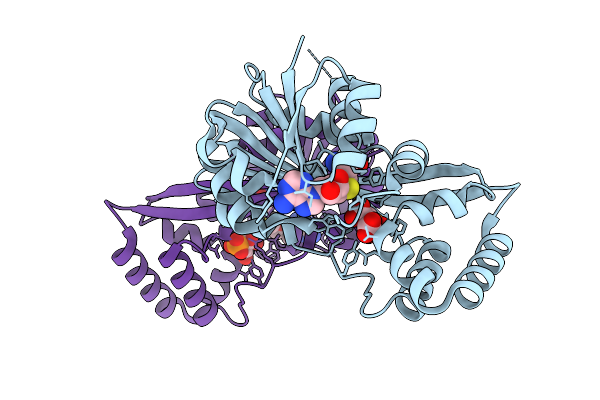 |
Organism: Pantoea ananatis
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2026-03-18 Classification: BIOSYNTHETIC PROTEIN Ligands: SAH, A1BHZ, NI |
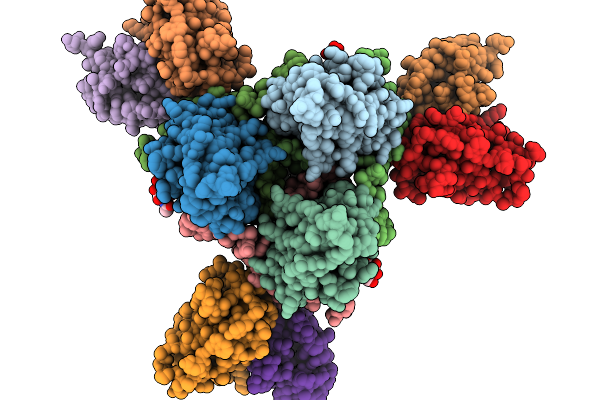 |
Cryo-Em Structure Of Sudv Glycoprotein With Modified Hr1C (L579P) And Hr2 Stalk Bound To Ca45 Fab
Organism: Sudan ebolavirus (strain human/uganda/gulu/2000), Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.13 Å Release Date: 2026-02-18 Classification: VIRAL PROTEIN Ligands: NAG |
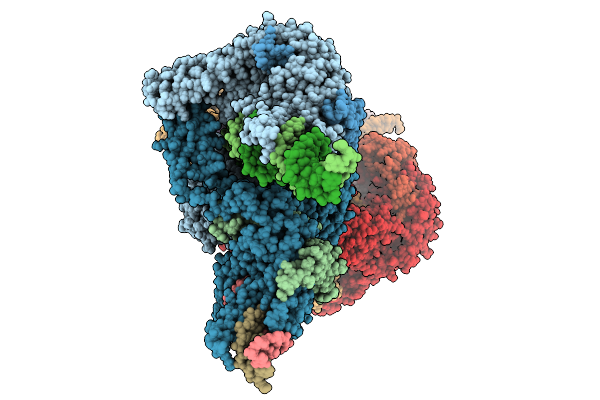 |
Organism: Bacillus
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: DNA BINDING PROTEIN Ligands: ZN |
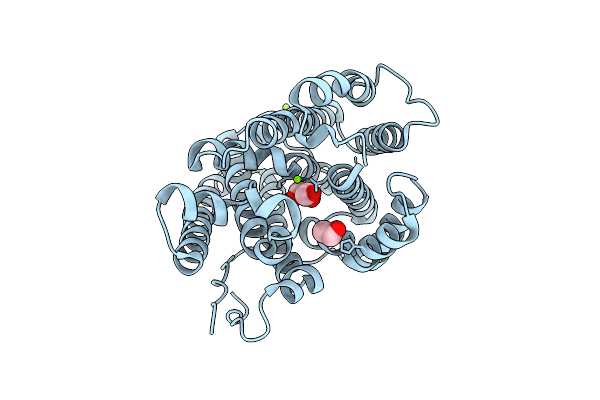 |
Organism: Nonomuraea coxensis
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-12-24 Classification: LYASE Ligands: MG, GOL, EDO |
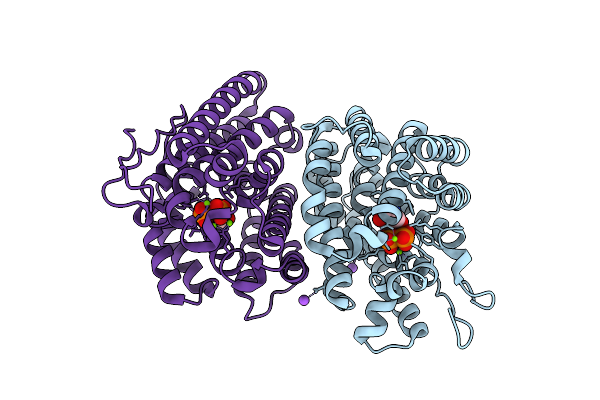 |
1-Epi-Cubenol Synthase From Nonomuraea Coxensis (Ncecs) In Complex With Pyrophosphate
Organism: Nonomuraea coxensis
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-12-24 Classification: LYASE Ligands: POP, MG, NA, TRS, GOL |
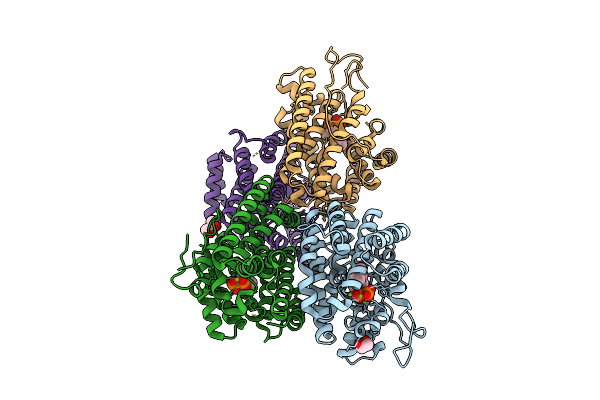 |
1-Epi-Cubenol Synthase From Nonomuraea Coxensis (Ncecs) In Complex With 2,3-Dhfpp
Organism: Nonomuraea coxensis
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2025-12-24 Classification: LYASE Ligands: 3E9, MG, BU3, NA |
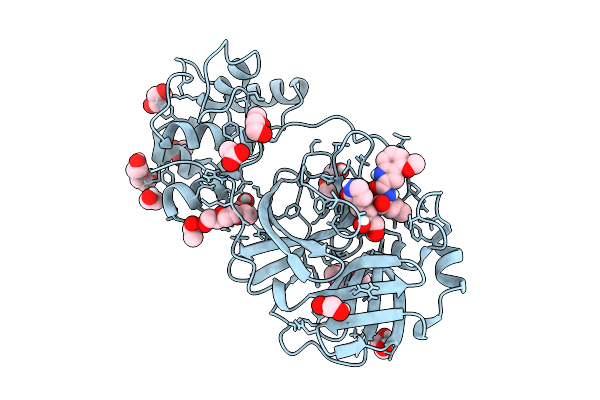 |
Co-Crystal Structure Of Feline Coronavirus Uu23 Main Protease With Pfizer Intravenous Compound Pf-00835231
Organism: Feline alphacoronavirus 1
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-11-26 Classification: VIRAL PROTEIN Ligands: V2M, ACY, PG4, EDO, PEG |
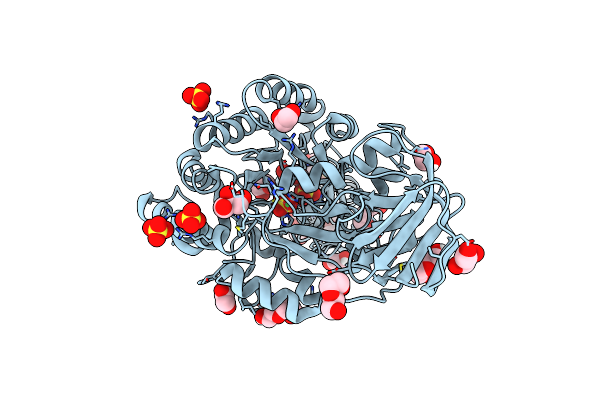 |
N-Acyl-D-Amino-Acid Deacylase (D-Acylase) From Klebsiella Pneumoniae In An Open Conformation
Organism: Klebsiella pneumoniae subsp. pneumoniae kp13
Method: X-RAY DIFFRACTION Resolution:2.27 Å Release Date: 2025-06-18 Classification: HYDROLASE Ligands: SO4, PEG, EDO, GOL, PGE, NI |
 |
N-Acyl-D-Amino-Acid Deacylase (D-Acylase) From Klebsiella Pneumoniae In The Absence Of Glycerol
Organism: Klebsiella pneumoniae subsp. pneumoniae kp13
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2025-06-18 Classification: HYDROLASE Ligands: EDO, SO4, NI |
 |
Organism: Thermus antranikianii dsm 12462
Method: ELECTRON MICROSCOPY Release Date: 2025-01-29 Classification: TRANSLOCASE/DNA Ligands: ANP |
 |
Organism: Thermus antranikianii dsm 12462
Method: ELECTRON MICROSCOPY Release Date: 2025-01-22 Classification: TRANSLOCASE |
 |
Organism: Thermus antranikianii dsm 12462
Method: ELECTRON MICROSCOPY Release Date: 2025-01-22 Classification: TRANSLOCASE/DNA Ligands: ANP |
 |
Organism: Bacillus, Bacillus phage spo1
Method: ELECTRON MICROSCOPY Release Date: 2024-12-04 Classification: TRANSCRIPTION |
 |
Organism: Bacillus, Bacillus phage spo1
Method: ELECTRON MICROSCOPY Resolution:2.94 Å Release Date: 2024-12-04 Classification: TRANSCRIPTION |
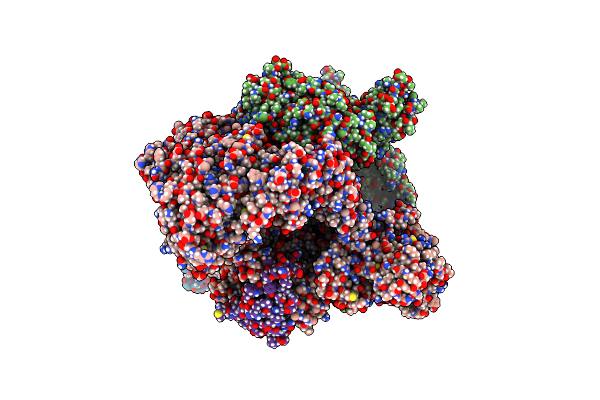 |
Organism: Bacillus
Method: ELECTRON MICROSCOPY Release Date: 2024-12-04 Classification: TRANSCRIPTION Ligands: ZN, MG |
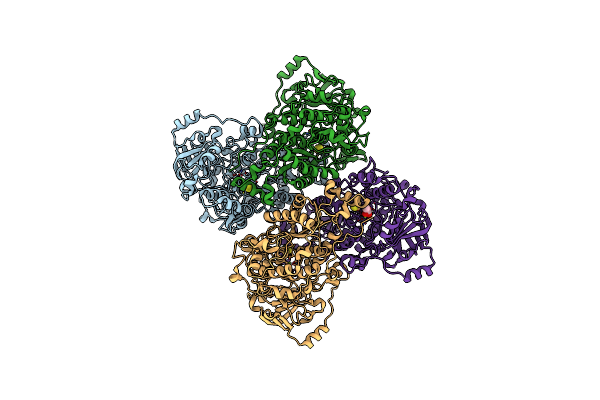 |
His579Leu Variant Of L-Arabinonate Dehydratase Co-Crystallized With 2-Oxobutyrate
Organism: Rhizobium leguminosarum bv. trifolii
Method: X-RAY DIFFRACTION Resolution:2.44 Å Release Date: 2024-09-25 Classification: LYASE Ligands: MG, FES, 2KT |
 |
Organism: Streptomyces yokosukanensis
Method: X-RAY DIFFRACTION Resolution:2.78 Å Release Date: 2024-06-05 Classification: BIOSYNTHETIC PROTEIN Ligands: YG3, DMS, SO4, GOL |
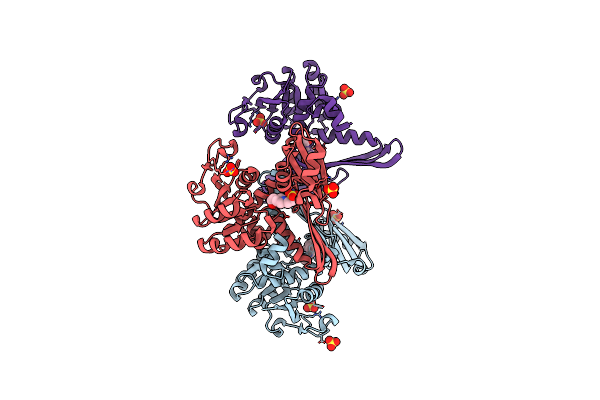 |
Crystal Structure Of Meso-Diaminopimelate Dehydrogenase From Prevotella Timonensis
Organism: Prevotella timonensis
Method: X-RAY DIFFRACTION Resolution:2.47 Å Release Date: 2023-12-13 Classification: OXIDOREDUCTASE Ligands: EPE, SO4 |
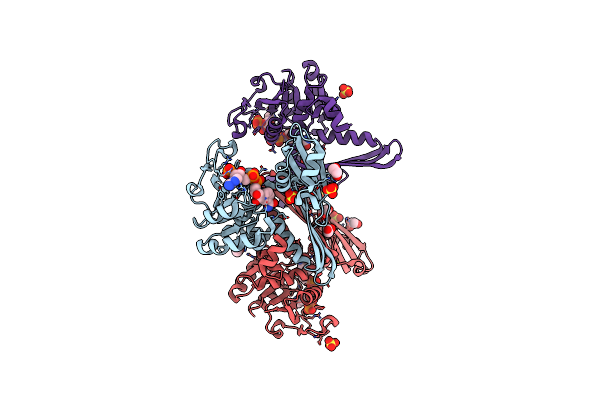 |
Crystal Structure Of Meso-Diaminopimelate Dehydrogenase From Prevotella Timonensis
Organism: Prevotella timonensis
Method: X-RAY DIFFRACTION Resolution:3.07 Å Release Date: 2023-12-13 Classification: OXIDOREDUCTASE Ligands: NDP, SO4, EDO |
 |
Organism: Stutzerimonas chloritidismutans aw-1
Method: X-RAY DIFFRACTION Resolution:1.27 Å Release Date: 2023-08-30 Classification: OXIDOREDUCTASE Ligands: FNR, CL |

