Search Count: 107
All
Selected
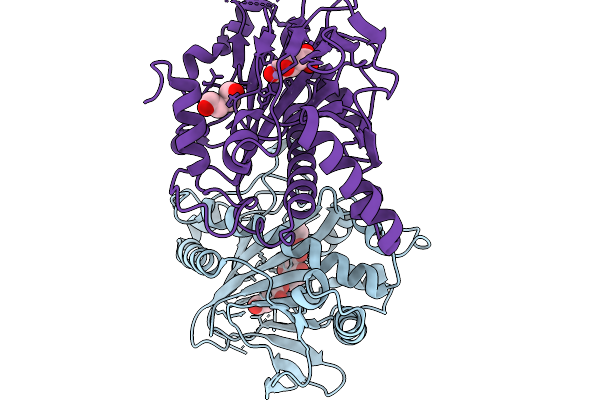 |
Fe/Alpha-Kg-Dependent Decarboxylase Trah In Complexes With Mn2+, Alpha-Kg, And Substrate
Organism: Penicillium crustosum
Method: X-RAY DIFFRACTION Resolution:2.89 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: AKG, MN, A1I30, PEG |
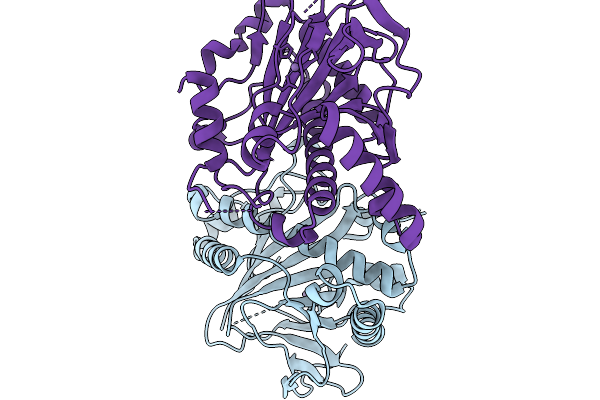 |
Crystal Structure Of Fe/Alphakg-Dependent Decarboxylase Trah In Complexes With Mn2+
Organism: Penicillium crustosum
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: MN |
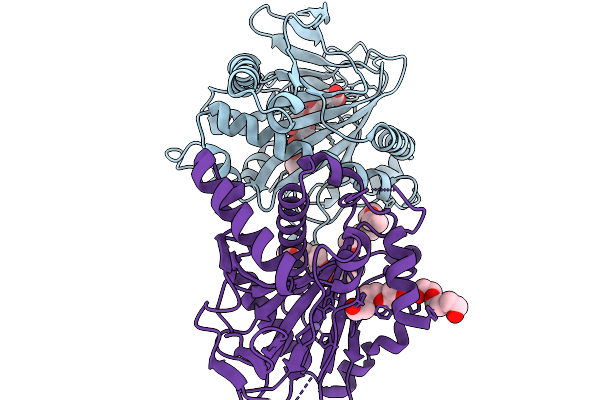 |
Fe/Alpha-Kg-Dependent Decarboxylase Trah In Complexes With Mn2+, Alpha-Kg, And Substrate
Organism: Penicillium crustosum
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: MN, AKG, A1I31, 1PG, PGE |
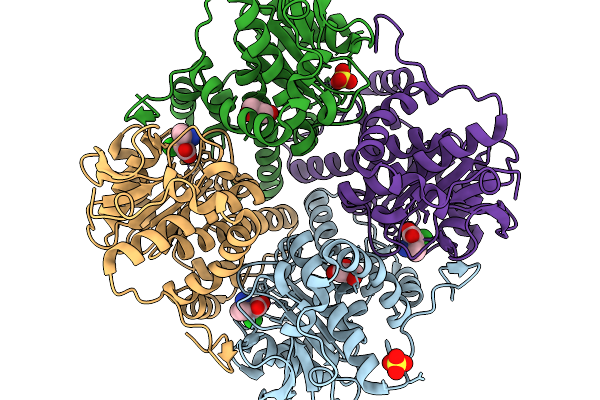 |
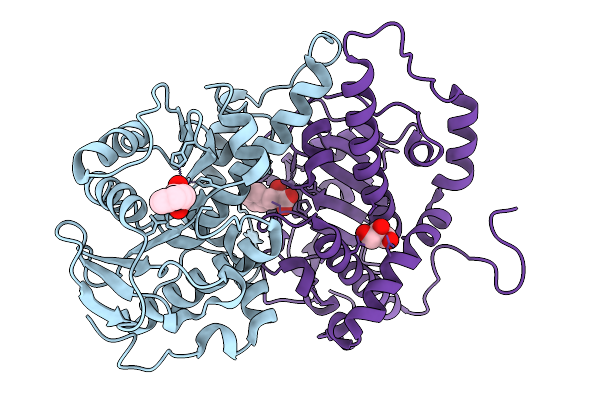
Organism: Penicillium aethiopicum
Method: X-RAY DIFFRACTION Resolution:2.17 Å Release Date: 2026-03-04 Classification: METAL BINDING PROTEIN Ligands: A1L7T, GOL, MN, CL, SO4 |
 |
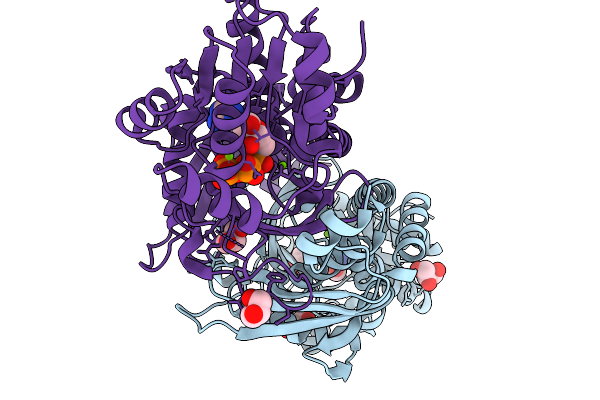
Structure Of Tqam From Penicillium Aethiopicum In Complex With D-(+)-3-Phenyllactic Acid
Organism: Penicillium aethiopicum
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2026-03-04 Classification: METAL BINDING PROTEIN Ligands: HF2, GOL, MN, CL |
 |
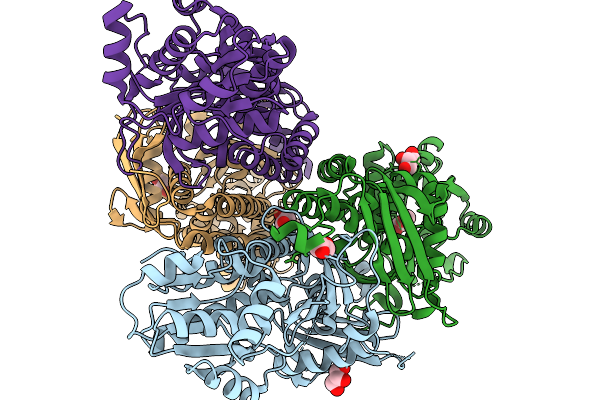
Organism: Penicillium aurantiogriseum
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2026-01-14 Classification: BIOSYNTHETIC PROTEIN Ligands: GOL, TRS, MG, ATP |
 |
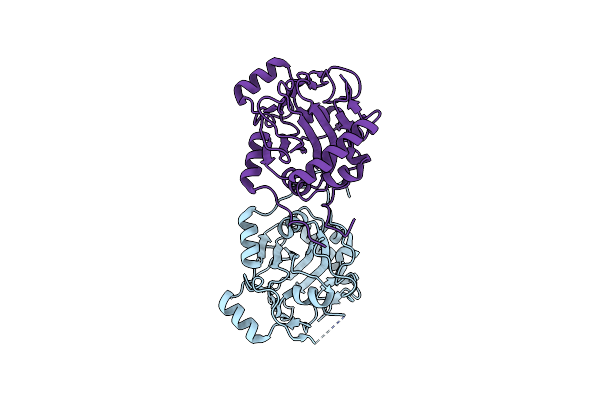
Organism: Penicillium aurantiogriseum
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-01-14 Classification: BIOSYNTHETIC PROTEIN Ligands: GOL |
 |
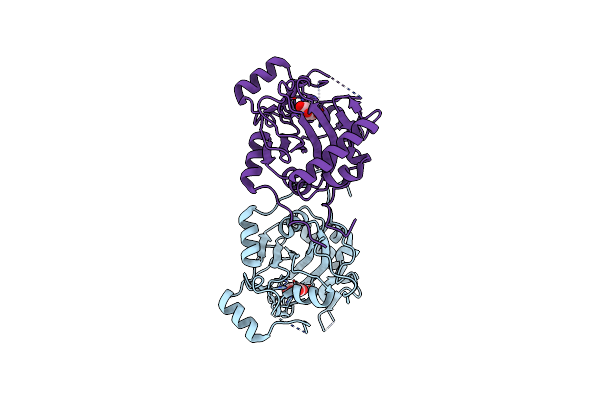
Organism: Uncultured bacterium esnapd13
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2023-04-19 Classification: STRUCTURAL PROTEIN Ligands: CL |
 |
Organism: Uncultured bacterium esnapd13
Method: X-RAY DIFFRACTION Resolution:1.97 Å Release Date: 2023-04-19 Classification: STRUCTURAL PROTEIN Ligands: AKG, FE |
 |
Organism: Uncultured bacterium esnapd13
Method: X-RAY DIFFRACTION Resolution:2.14 Å Release Date: 2023-04-19 Classification: STRUCTURAL PROTEIN Ligands: FE, AKG, PRO |
 |
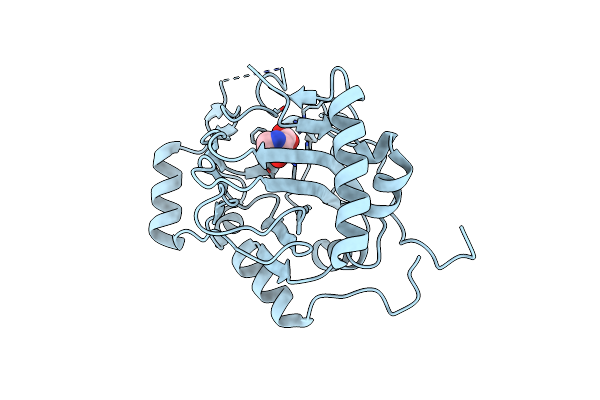
Organism: Uncultured bacterium esnapd13
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2023-04-19 Classification: STRUCTURAL PROTEIN Ligands: HYP |
 |
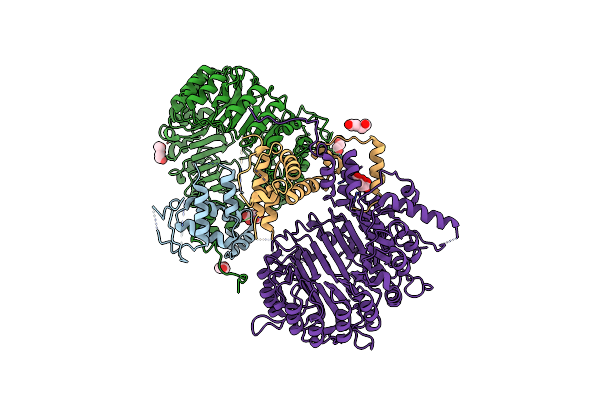
Organism: Uncultured bacterium esnapd13
Method: X-RAY DIFFRACTION Resolution:2.38 Å Release Date: 2023-04-19 Classification: STRUCTURAL PROTEIN Ligands: HY3 |
 |
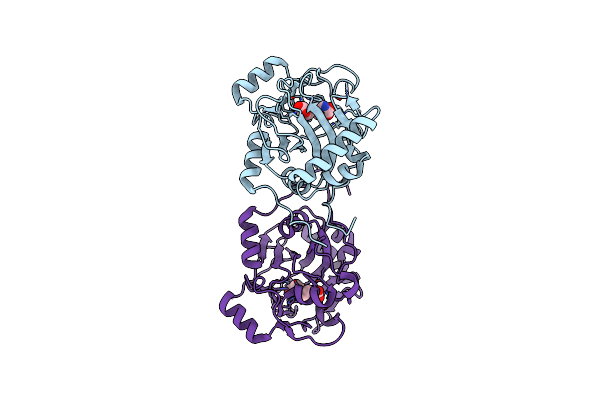
Organism: Arabidopsis thaliana, Oryza sativa subsp. japonica, Oryza sativa tropical japonica subgroup
Method: X-RAY DIFFRACTION Resolution:3.21 Å Release Date: 2022-04-20 Classification: SIGNALING PROTEIN Ligands: PEG, CIT |
 |
Organism: Uncultured bacterium esnapd13
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2022-02-02 Classification: HYDROLASE Ligands: PRO, SIN, O |
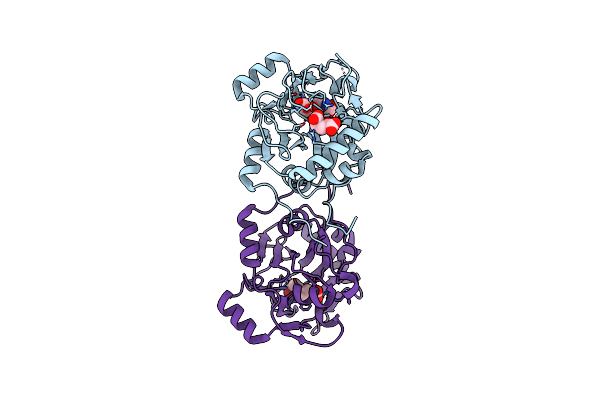 |
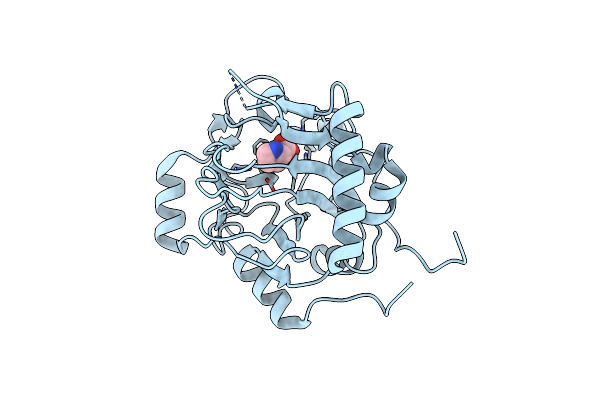
Trans-Proline-Hydroxylase H11 With Succinic And L-Proline In The Fourth Reaction State.
Organism: Uncultured bacterium esnapd13
Method: X-RAY DIFFRACTION Resolution:1.92 Å Release Date: 2022-02-02 Classification: HYDROLASE Ligands: PRO, SIN, AKG, O |
 |
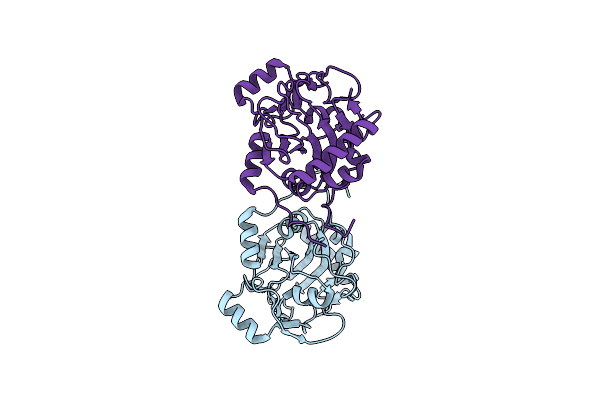
Organism: Uncultured bacterium esnapd13
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2022-02-02 Classification: HYDROLASE Ligands: PRO, O |
 |
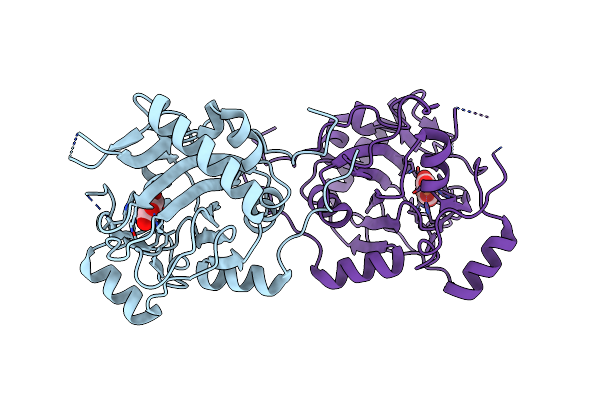
Organism: Uncultured bacterium esnapd13
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2022-02-02 Classification: HYDROLASE |
 |
Organism: Uncultured bacterium esnapd13
Method: X-RAY DIFFRACTION Resolution:1.97 Å Release Date: 2022-02-02 Classification: HYDROLASE Ligands: AKG, FE |
 |
Organism: Uncultured bacterium esnapd13
Method: X-RAY DIFFRACTION Resolution:2.14 Å Release Date: 2022-02-02 Classification: HYDROLASE Ligands: AKG, PRO, FE |
 |
Organism: Uncultured bacterium esnapd13
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2022-02-02 Classification: HYDROLASE Ligands: HYP |

