Search Count: 2,433
All
Selected
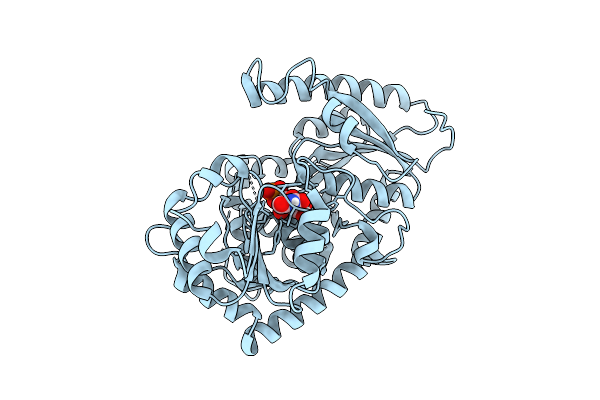 |
Crystal Structure Of The Glycosyltransferase Qsfuct From Quillaja Saponaria
Organism: Quillaja saponaria
Method: X-RAY DIFFRACTION Resolution:2.11 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: UDP |
 |
Organism: Azotobacter vinelandii dj
Method: ELECTRON MICROSCOPY Resolution:2.16 Å Release Date: 2026-03-04 Classification: METAL BINDING PROTEIN Ligands: S5Q, SF4, MG |
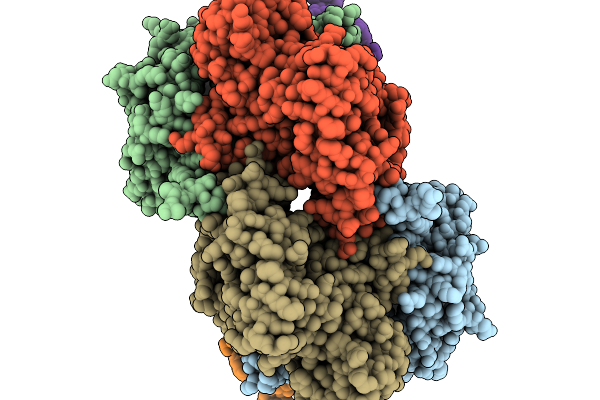 |
Organism: Azotobacter vinelandii dj
Method: ELECTRON MICROSCOPY Resolution:2.14 Å Release Date: 2026-03-04 Classification: METAL BINDING PROTEIN Ligands: S5Q, SF4, MG |
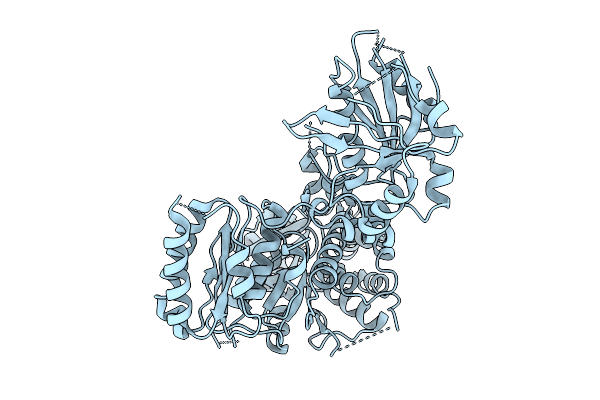 |
Organism: Shewanella baltica
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2026-02-25 Classification: LIPID BINDING PROTEIN |
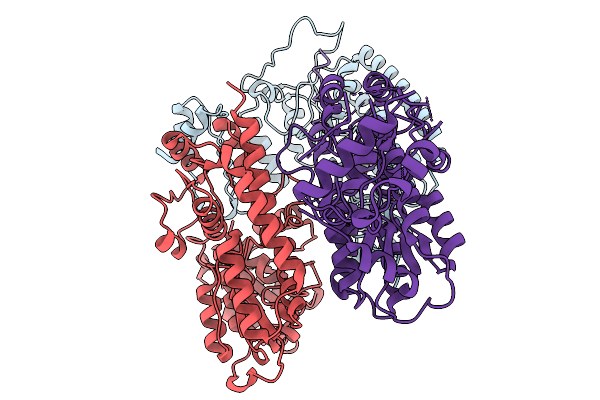 |
Organism: Azotobacter vinelandii dj
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: OXIDOREDUCTASE |
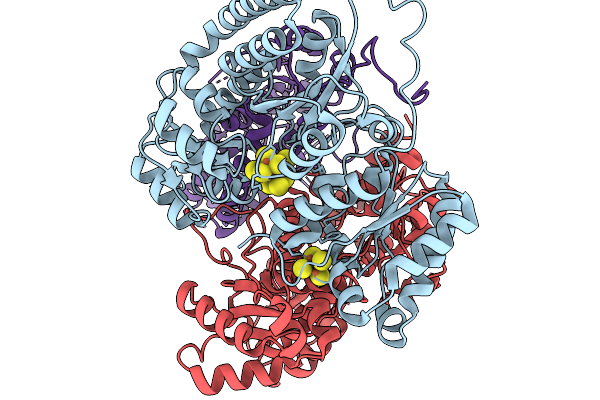 |
Organism: Azotobacter vinelandii dj
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: OXIDOREDUCTASE Ligands: S5Q, SF4 |
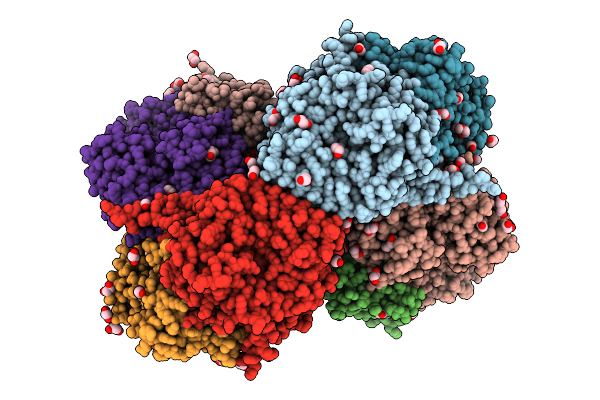 |
Organism: Azotobacter vinelandii dj
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2025-12-03 Classification: METAL BINDING PROTEIN Ligands: SF4, S5Q, PEG, EDO, 1PE, PGE, MG, PG4 |
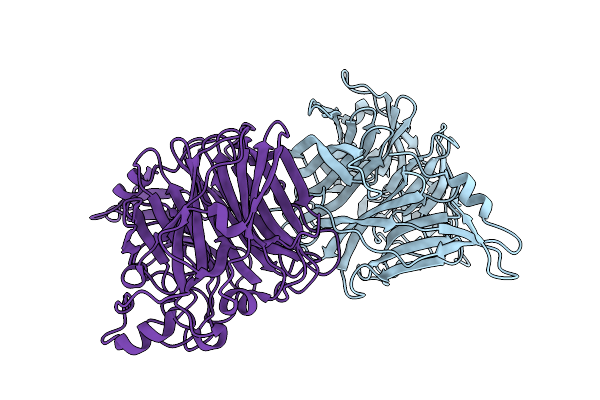 |
Organism: J virus
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-11-19 Classification: VIRAL PROTEIN |
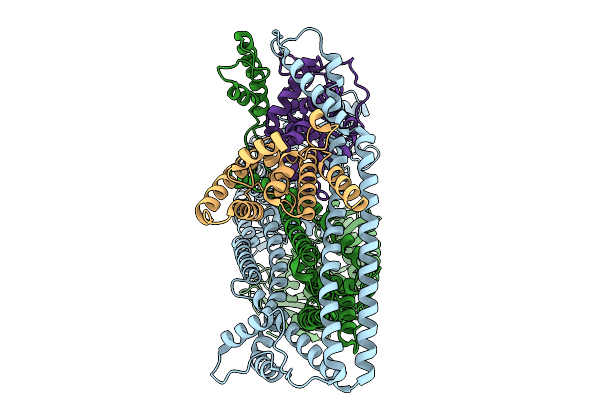 |
Organism: Salmonella enterica subsp. enterica serovar tennessee
Method: ELECTRON MICROSCOPY Release Date: 2025-10-01 Classification: IMMUNE SYSTEM |
 |
Organism: Salmonella enterica subsp. enterica serovar tennessee, Bacillus subtilis subsp. subtilis str. 168
Method: ELECTRON MICROSCOPY Release Date: 2025-10-01 Classification: IMMUNE SYSTEM |
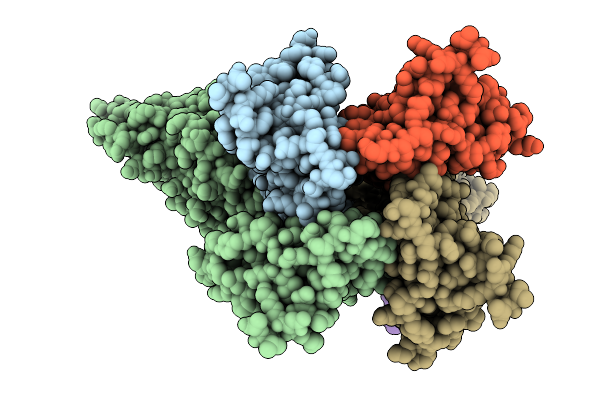 |
Organism: Salmonella enterica subsp. enterica serovar tennessee, Bacillus subtilis subsp. subtilis str. 168
Method: ELECTRON MICROSCOPY Release Date: 2025-10-01 Classification: IMMUNE SYSTEM |
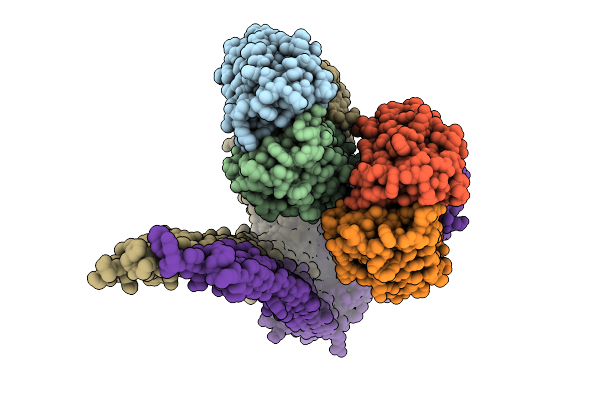 |
Organism: Pleurotus ostreatus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: MEMBRANE PROTEIN |
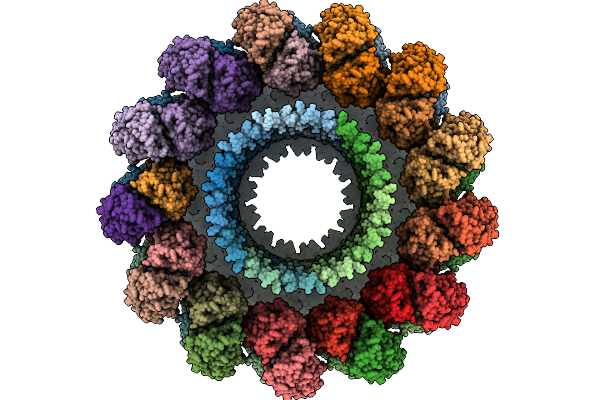 |
Organism: Pleurotus ostreatus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: MEMBRANE PROTEIN |
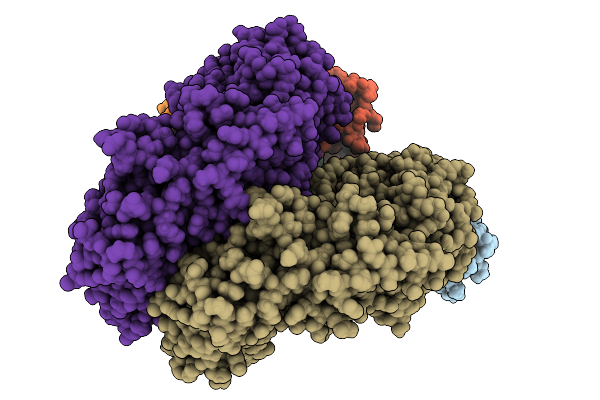 |
Organism: Pleurotus ostreatus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: MEMBRANE PROTEIN |
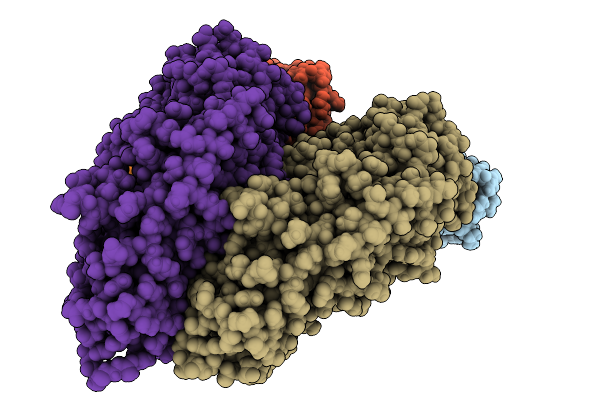 |
Organism: Pleurotus ostreatus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: MEMBRANE PROTEIN Ligands: A1A8R, CLR |
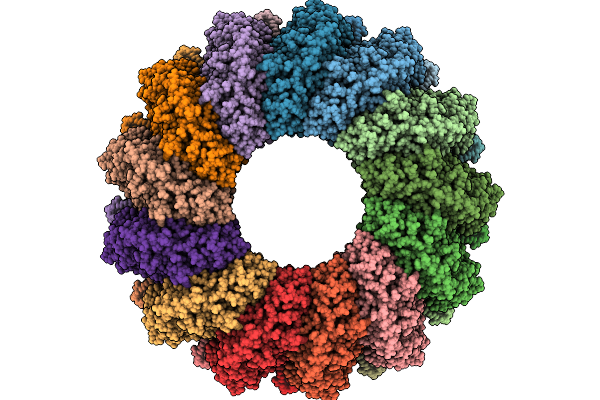 |
Organism: Pleurotus ostreatus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: MEMBRANE PROTEIN |
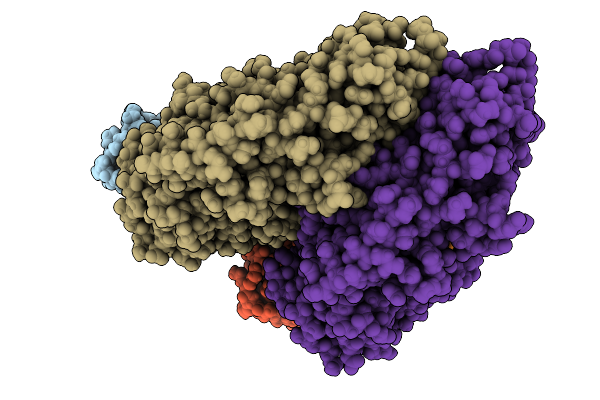 |
Organism: Pleurotus ostreatus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: MEMBRANE PROTEIN Ligands: A1A8R |
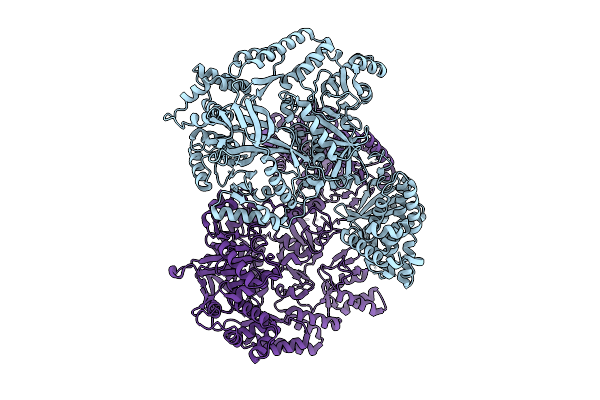 |
Structure Of Madb, A Class I Dehydrates From Clostridium Maddingley In The Apo State
Organism: Clostridium sp. maddingley mbc34-26
Method: ELECTRON MICROSCOPY Release Date: 2025-07-16 Classification: BIOSYNTHETIC PROTEIN |
 |
Structure Of Madb, A Class I Dehydrates From Clostridium Maddingley, In Complex With Its Substrate
Organism: Clostridium sp. maddingley mbc34-26
Method: ELECTRON MICROSCOPY Release Date: 2025-07-16 Classification: BIOSYNTHETIC PROTEIN |
 |
The Crystal Structure Of The Ha1 Domain Of Hemagglutinin From A/Shanghai/02/2013 (H7N9) Bound To H7-235 Fab
Organism: Influenza a virus (a/shanghai/02/2013(h7n9)), Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-06-25 Classification: IMMUNE SYSTEM Ligands: NAG, SIA |

