Search Count: 1,381
All
Selected
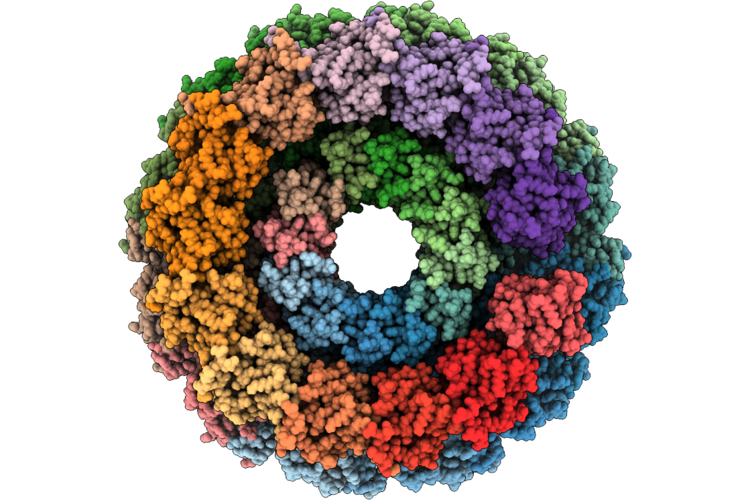 |
Organism: Escherichia phage theodorherzl, Klebsiella pneumoniae
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: HYDROLASE Ligands: MG |
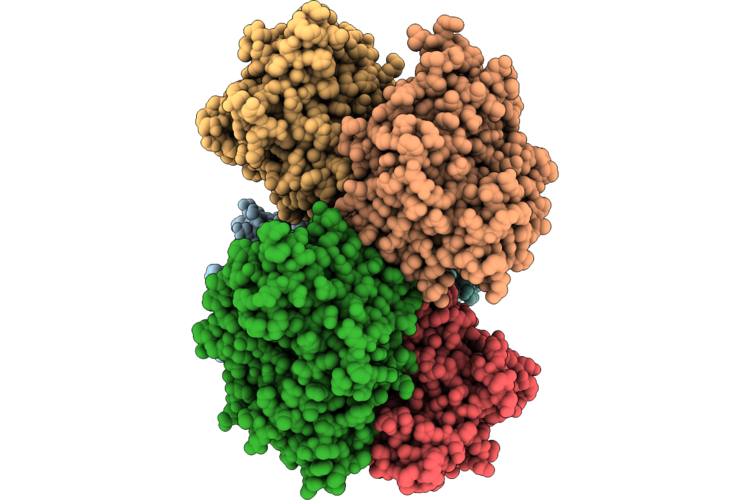 |
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Resolution:2.74 Å Release Date: 2026-05-13 Classification: HYDROLASE/RNA Ligands: ATP, ZN, MG |
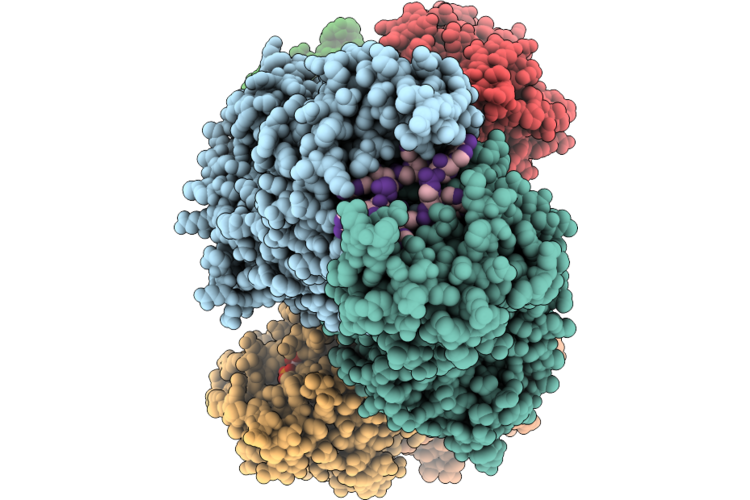 |
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2026-05-13 Classification: HYDROLASE/RNA Ligands: ATP, ZN, MG |
 |
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: HYDROLASE/RNA Ligands: ATP, MG, ZN |
 |
Structure Of The Type Iii Crispr-Associated Deaminase In Complex Ca6 And Atp, State 4
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Resolution:2.94 Å Release Date: 2026-05-13 Classification: HYDROLASE/RNA Ligands: ATP, ZN, MG |
 |
Structure Of The Type Iii Crispr-Associated Deaminase In Complex Ca6 And Atp, State 5
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2026-05-13 Classification: HYDROLASE/RNA Ligands: ATP, ZN, MG |
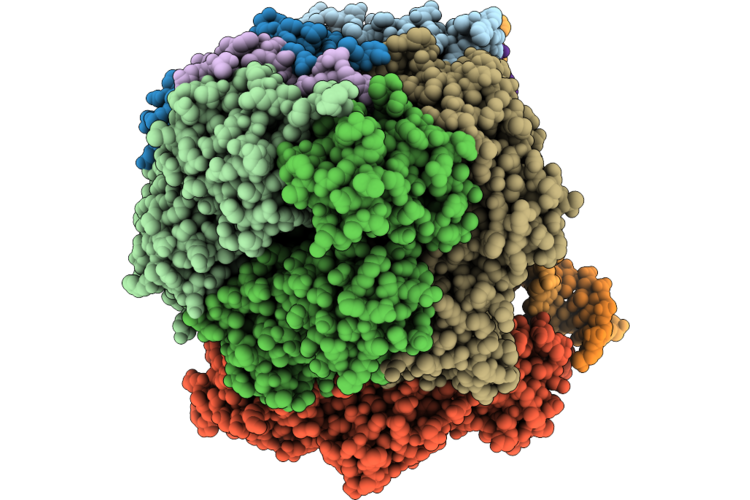 |
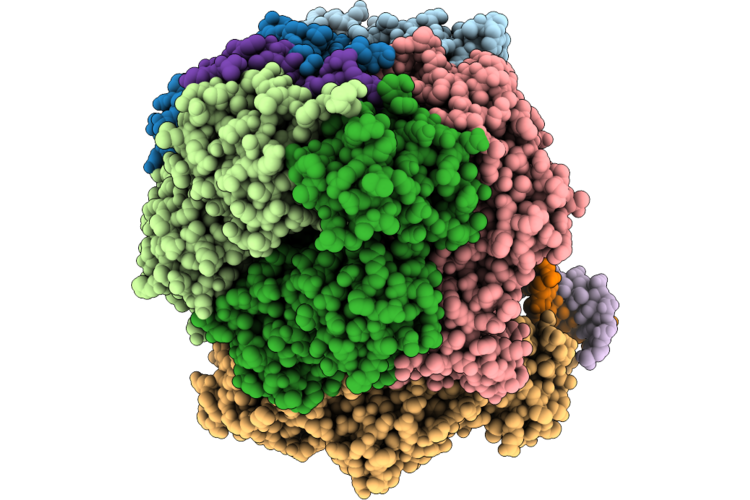
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 1-Nt Gapped Dna In State 1 Conformer 2 With Flexibly Bound Dna At The Shoulder
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:2.71 Å Release Date: 2026-04-29 Classification: REPLICATION/DNA Ligands: ZN, AGS, MG |
 |
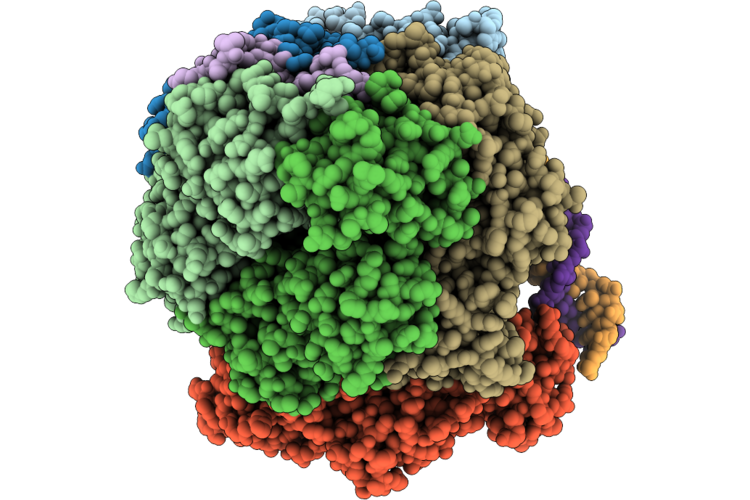
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 1-Nt Gapped Dna In State 1 Conformer 3 With Disordered Dna At The Shoulder
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:2.73 Å Release Date: 2026-04-29 Classification: REPLICATION/DNA Ligands: ZN, AGS, MG |
 |
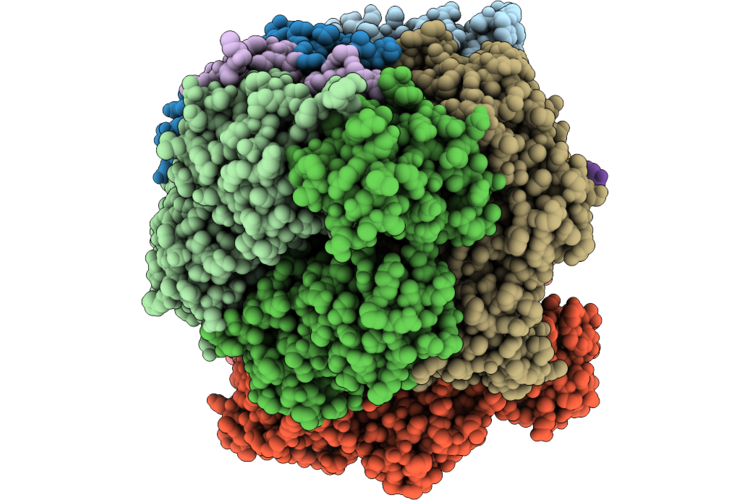
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 1-Nt Gapped Dna In State 2 Conformer 1 With Sharply Bent Dna
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:2.72 Å Release Date: 2026-04-29 Classification: REPLICATION/DNA Ligands: ZN, AGS, MG |
 |
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 1-Nt Gapped Dna In State 2 Conformer 2 With Less Bent Dna
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:3.12 Å Release Date: 2026-04-29 Classification: REPLICATION/DNA Ligands: ZN, AGS, MG |
 |
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 10-Nt Gapped Dna In State 1 The Dna Recognition State
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:2.54 Å Release Date: 2026-04-29 Classification: REPLICATION/DNA Ligands: ZN, AGS, MG |
 |
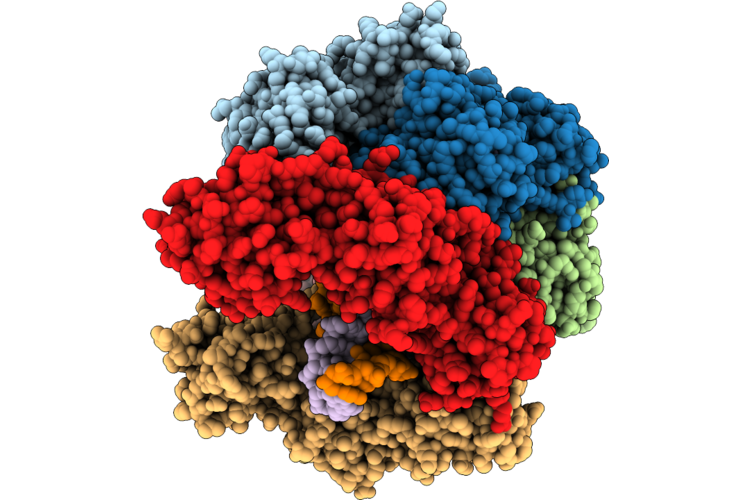
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 10-Nt Gapped Dna In State 2 Conformer 1 With Fully Open Clamp And Unsettled Dna
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:2.54 Å Release Date: 2026-04-29 Classification: REPLICATION/DNA Ligands: ZN, AGS, MG |
 |
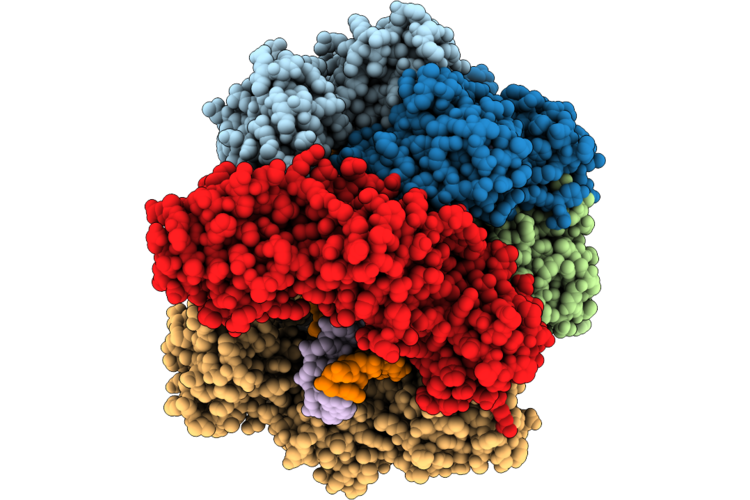
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 10-Nt Gapped Dna In State 2 Conformer 2 With Fully Open Clamp And Settled Dna
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:2.53 Å Release Date: 2026-04-29 Classification: REPLICATION Ligands: ZN, AGS, MG |
 |
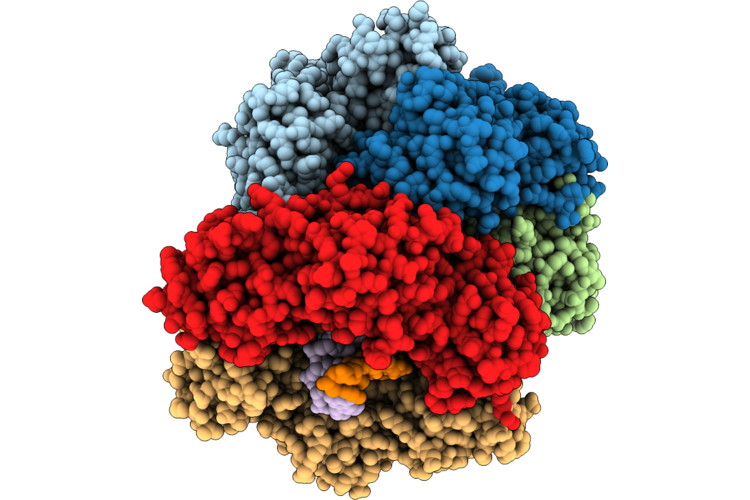
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 10-Nt Gapped Dna In State 2 Conformer 3 With Partially Closed Clamp
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-04-29 Classification: REPLICATION/DNA Ligands: ZN, AGS, MG |
 |
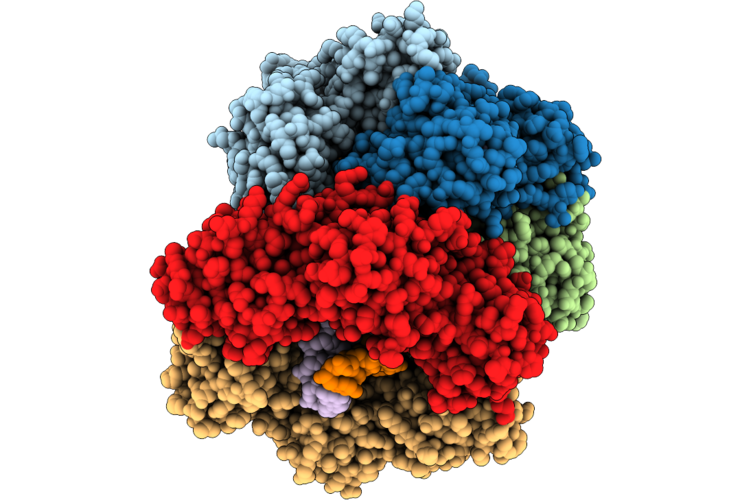
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 10-Nt Gapped Dna In State 2 Conformer 4 With Fully Closed Clamp
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:2.88 Å Release Date: 2026-04-29 Classification: REPLICATION Ligands: ZN, AGS, MG |
 |
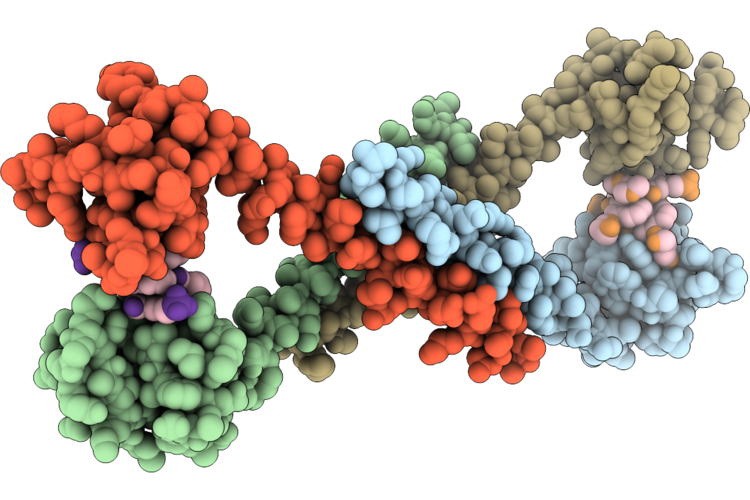
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 10-Nt Gapped Dna In State 2 Conformer 5 With Fully Closed Clamp
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-04-29 Classification: REPLICATION/DNA Ligands: ZN, AGS, MG |
 |
Crystal Structure Of Affitin C10 Fused To A Coiled-Coil Domain In Complex With A Quinoline Oligoamide Foldamer
Organism: Sulfolobus acidocaldarius, Mus musculus, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-22 Classification: DE NOVO PROTEIN |
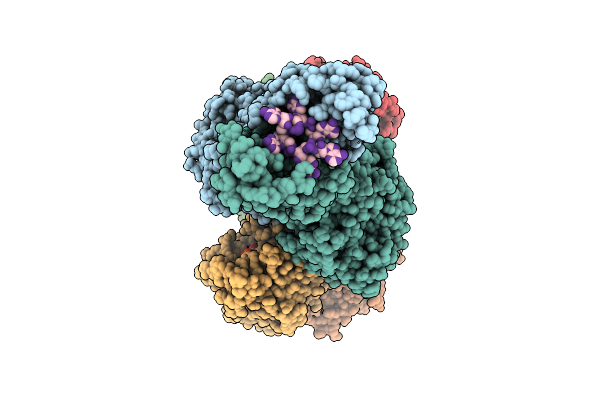 |
The Structure Of Type Iii Crispr-Associated Deaminase In Complex Ca6 And Atp, State 3
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: HYDROLASE/RNA Ligands: ATP, MG, ZN |
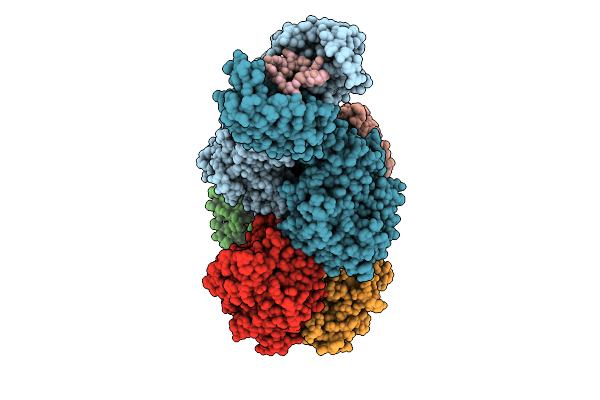 |
The Structure Of Type Iii Crispr-Associated Deaminase In Complex Ca6 And Atp, State 4
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Resolution:2.94 Å Release Date: 2026-04-15 Classification: HYDROLASE/RNA Ligands: ATP, ZN, MG |
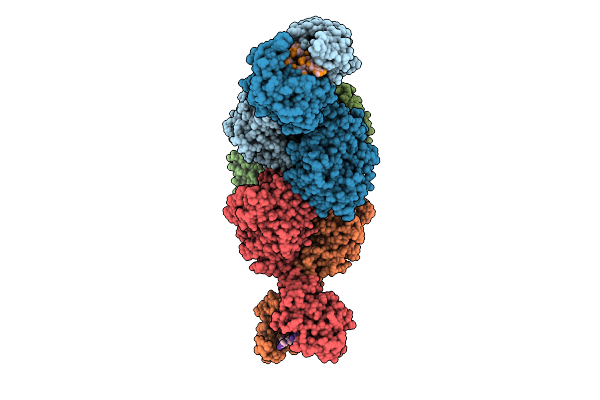 |
The Structure Of Type Iii Crispr-Associated Deaminase In Complex Ca6 And Atp, State 5
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2026-04-15 Classification: HYDROLASE/RNA Ligands: ATP, ZN, MG |

