Search Count: 4,381
All
Selected
 |
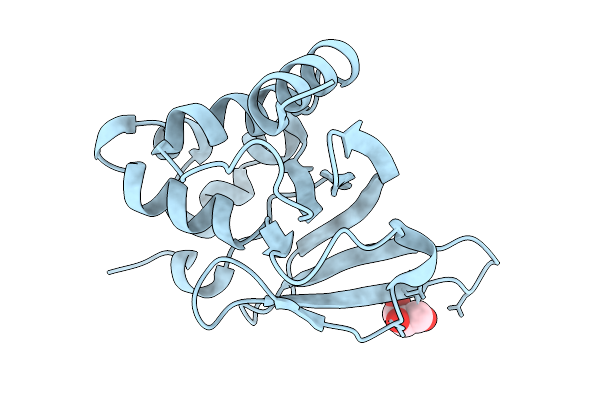
Organism: Dolichospermum circinale
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: GOL |
 |
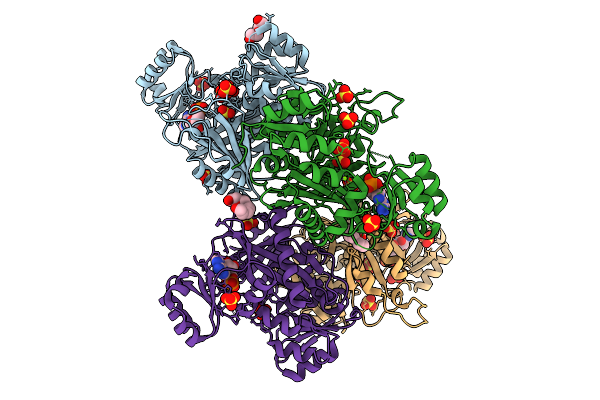
Organism: Scenedesmus sp. oki-4n
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-02-04 Classification: TRANSPORT PROTEIN Ligands: GOL |
 |
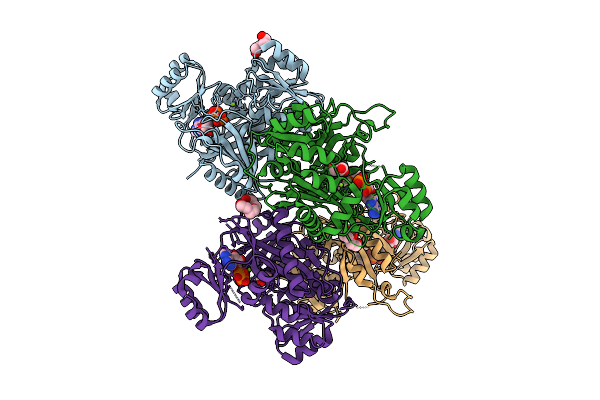
Crystal Structure Of Phosphoribosylaminoimidazole Carboxylase From Burkholderia Xenovorans (Amp, Adp And Sulfate Complex)
Organism: Paraburkholderia xenovorans lb400
Method: X-RAY DIFFRACTION Resolution:2.49 Å Release Date: 2026-01-21 Classification: LYASE Ligands: AMP, PEG, MG, SO4, PG4, ADP |
 |
Crystal Structure Of Phosphoribosylaminoimidazole Carboxylase From Burkholderia Xenovorans (Atp Complex)
Organism: Paraburkholderia xenovorans lb400
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2025-12-24 Classification: LYASE Ligands: ATP, PEG, MG, PGE, GOL, ADP, PG4 |
 |
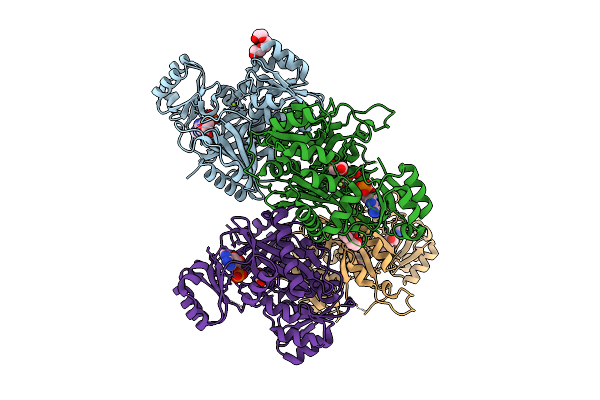
Crystal Structure Of Phosphoribosylaminoimidazole Carboxylase From Burkholderia Xenovorans (Adp Complex)
Organism: Paraburkholderia xenovorans lb400
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2025-12-24 Classification: LYASE Ligands: ADP, PEG, MG, PGE, GOL, PG4 |
 |
Crystal Structure Of Phosphoribosylaminoimidazole Carboxylase From Burkholderia Xenovorans (Amp Complex)
Organism: Paraburkholderia xenovorans lb400
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2025-12-24 Classification: LYASE Ligands: AMP, PGE, MG, PEG, PG4, GOL |
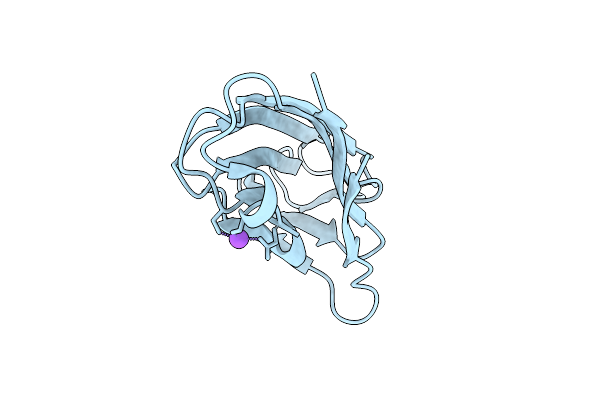 |
Apo Form Of Tumor Necrosis Factor-Like Lectin Pltl From Photorhabdus Laumondii
Organism: Photorhabdus laumondii subsp. laumondii tto1
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2025-10-29 Classification: SUGAR BINDING PROTEIN |
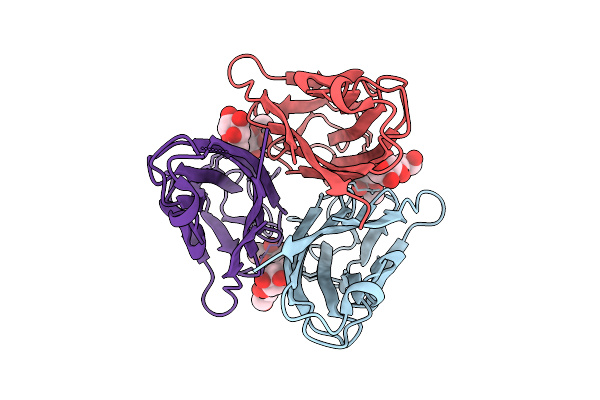 |
Tumor Necrosis Factor-Like Lectin Pltl From Photorhabdus Laumondii In Complex With Blood Group B Trisaccharide
Organism: Photorhabdus laumondii subsp. laumondii tto1
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-10-29 Classification: SUGAR BINDING PROTEIN Ligands: FUC, GLA |
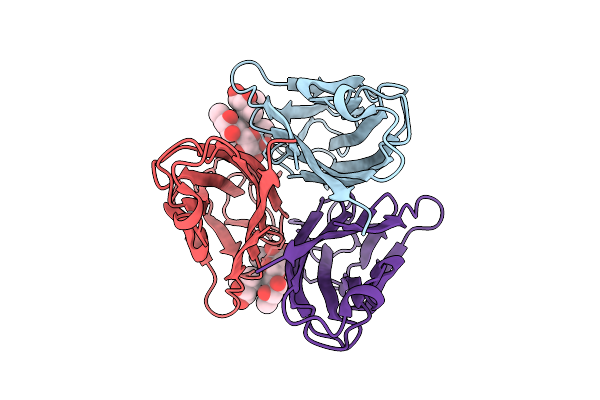 |
Tumor Necrosis Factor-Like Lectin Pltl From Photorhabdus Laumondii In Complex With Lewis Y Tetrasaccharide
Organism: Photorhabdus laumondii subsp. laumondii tto1
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-10-29 Classification: SUGAR BINDING PROTEIN |
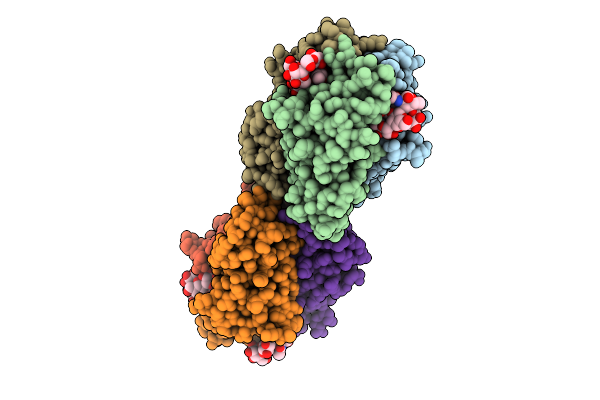 |
Tumor Necrosis Factor-Like Lectin Pltl From Photorhabdus Laumondii In Complex With B Lewis B Pentasaccharide
Organism: Photorhabdus laumondii subsp. laumondii tto1
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2025-10-29 Classification: SUGAR BINDING PROTEIN |
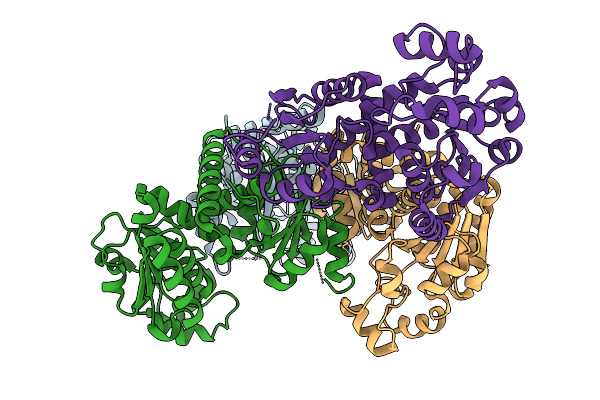 |
Crystal Structure Of Ornithine Carbamoyltransferase From Burkholderia Xenovorans
Organism: Paraburkholderia xenovorans lb400
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2025-09-24 Classification: TRANSFERASE Ligands: CL |
 |
Crystal Structure Of Ornithine Carbamoyltransferase From Burkholderia Xenovorans In Complex With Phosphono Carbamate
Organism: Paraburkholderia xenovorans lb400
Method: X-RAY DIFFRACTION Resolution:2.67 Å Release Date: 2025-09-24 Classification: TRANSFERASE Ligands: CP, PO4, CL |
 |
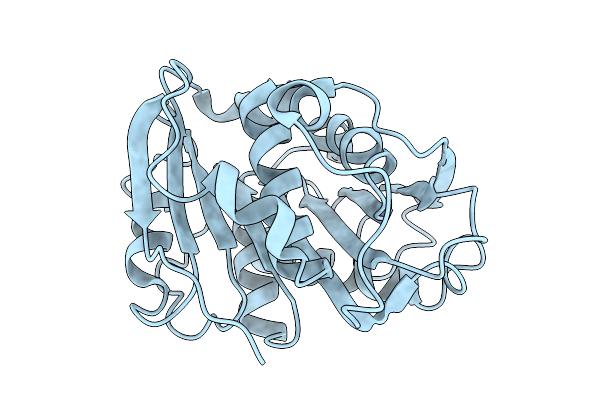
Crystal Structure Of The Polyester Hydrolase Leipzig 7 (Phl7) Mut3 Variant With Glycosylation By Expression In Pichia Pastoris (P_Phl7Mut3)
Organism: Compost metagenome
Method: X-RAY DIFFRACTION Resolution:1.37 Å Release Date: 2025-09-10 Classification: HYDROLASE Ligands: SO4 |
 |
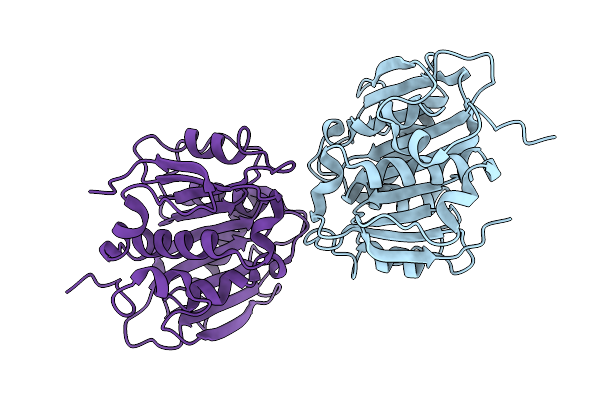
Crystal Structure Of The Non-Glycosylated Polyester Hydrolase Leipzig 7 (Phl7) Mut3 Variant Expressed In Pichia Pastoris (P_Phl7Mut3_Ng)
Organism: Compost metagenome
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2025-09-10 Classification: HYDROLASE Ligands: CL |
 |
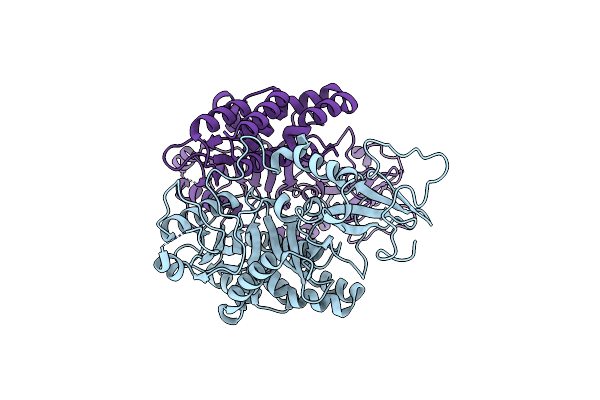
Crystal Structure Of The Polyester Hydrolase Leipzig 7 (Phl7) Mut3 Variant Expressed In E. Coli (E_Phl7Mut3)
Organism: Compost metagenome
Method: X-RAY DIFFRACTION Resolution:0.89 Å Release Date: 2025-09-10 Classification: HYDROLASE |
 |
Organism: Pseudomonas phage vb_pa32_gums
Method: ELECTRON MICROSCOPY Release Date: 2025-07-02 Classification: UNKNOWN FUNCTION |
 |
Organism: Photorhabdus laumondii subsp. laumondii tto1
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-03-05 Classification: CELL ADHESION Ligands: PEG, MFU |
 |
Organism: Photorhabdus laumondii subsp. laumondii tto1
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2025-03-05 Classification: CELL ADHESION Ligands: MFU |
 |
Organism: Photorhabdus laumondii subsp. laumondii tto1
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-03-05 Classification: CELL ADHESION Ligands: MFU, EDO, PEG |
 |
Organism: Photorhabdus laumondii subsp. laumondii tto1
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2025-03-05 Classification: CELL ADHESION Ligands: MFU, CL, MG |

