Search Count: 7,389
All
Selected
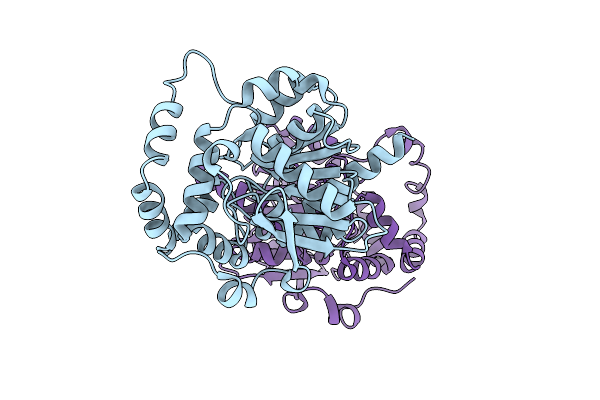 |
Organism: Unidentified
Method: X-RAY DIFFRACTION Resolution:3.30 Å Release Date: 2026-04-01 Classification: HYDROLASE |
 |
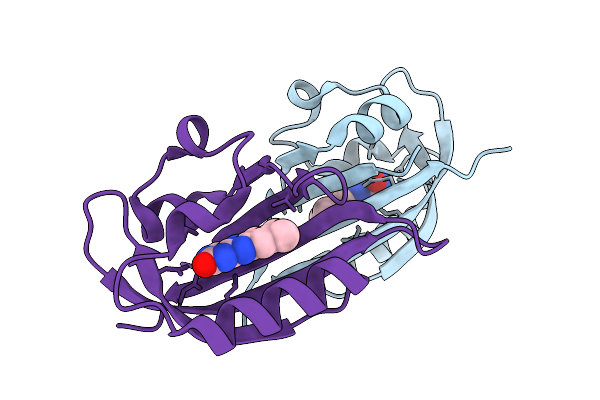
Structure Of Chloroflexus Aggregans Flavin Based Fluorescent Protein (Cagfbfp) Q89E Variant (Complex With Fmn)
Organism: Chloroflexus aggregans (strain md-66 / dsm 9485)
Method: X-RAY DIFFRACTION Resolution:1.27 Å Release Date: 2026-03-25 Classification: FLUORESCENT PROTEIN Ligands: MES, FMN |
 |
Structure Of Chloroflexus Aggregans Flavin Based Fluorescent Protein (Cagfbfp) Q89E Variant (Complex With Lc)
Organism: Chloroflexus aggregans (strain md-66 / dsm 9485)
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2026-03-25 Classification: FLUORESCENT PROTEIN Ligands: LUM |
 |
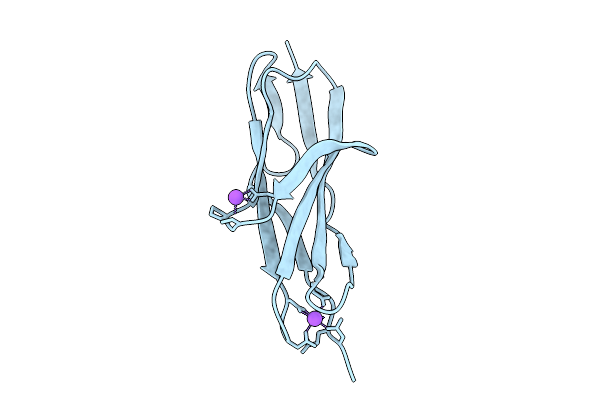
Organism: Aquimarina sp. 2-a2
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-03-18 Classification: SUGAR BINDING PROTEIN |
 |
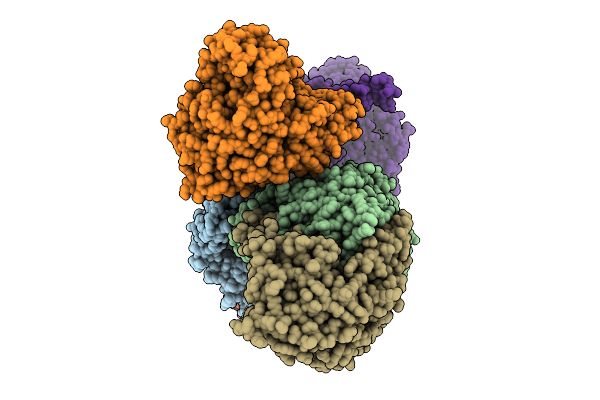
Organism: Aquimarina sp. 2-a2
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-03-18 Classification: SUGAR BINDING PROTEIN |
 |
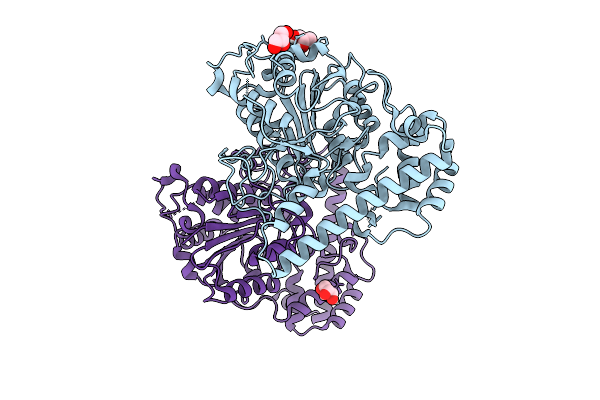
Organism: Paecilomyces variotii
Method: X-RAY DIFFRACTION Resolution:1.92 Å Release Date: 2026-03-11 Classification: BIOSYNTHETIC PROTEIN |
 |
Organism: Viola dissecta
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2026-03-11 Classification: LIGASE Ligands: GOL |
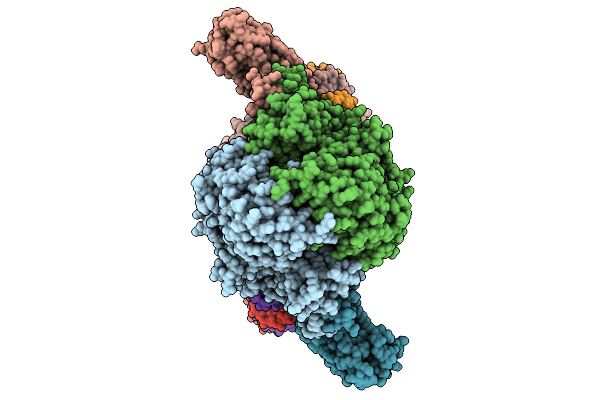 |
Cryoem Structure Of M. Mazei Topoisomerase Vi(A-E342Q)-Minicircle Dna Complex In Cleavage State
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: CA, ANP, K |
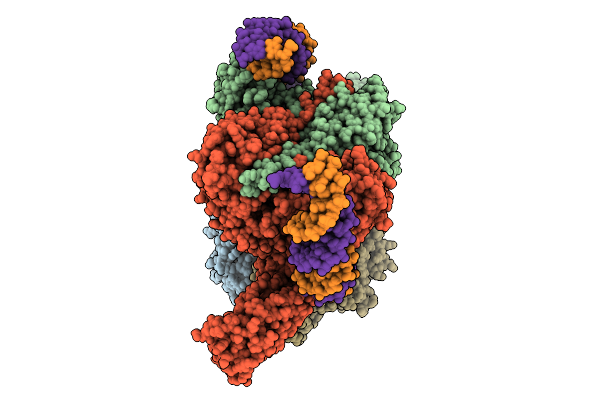 |
Cryoem Structure Of M. Mazei Topoisomerase Vi(A-E342Q)-Minicircle Dna Complex In Asymmetric State
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: ANP, CA, K |
 |
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: ANP, MG, K |
 |
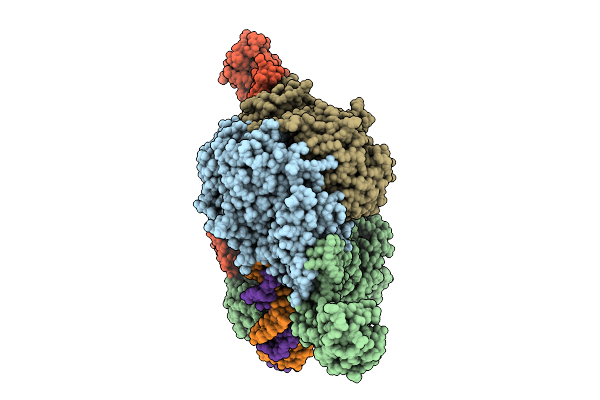
Cryoem Structure Of M. Mazei Topoisomerase Vi-Minicircle Dna Complex In Partially Unfolded Transducer State
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: MG, ANP, K |
 |
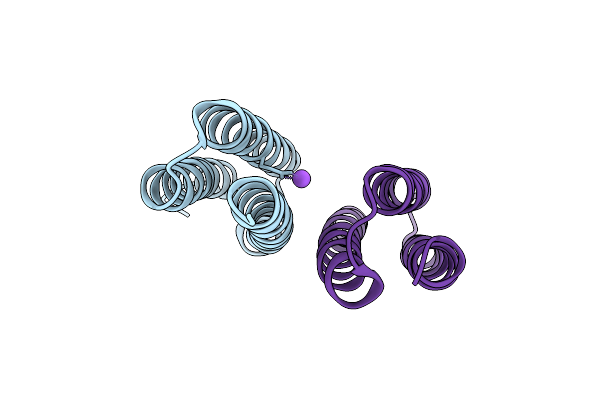
Cryoem Structure Of M. Mazei Topoisomerase Vi-Minicircle Dna Complex In Asymmetric State
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: ANP, MG, K |
 |
Mini-Protein Binder Of N-Terminal Domain Of Nucleocapsid Protein Of Sars-Cov2
Organism: Unidentified
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-02-25 Classification: DE NOVO PROTEIN |
 |
Organism: Unidentified
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-02-18 Classification: RECOMBINATION |
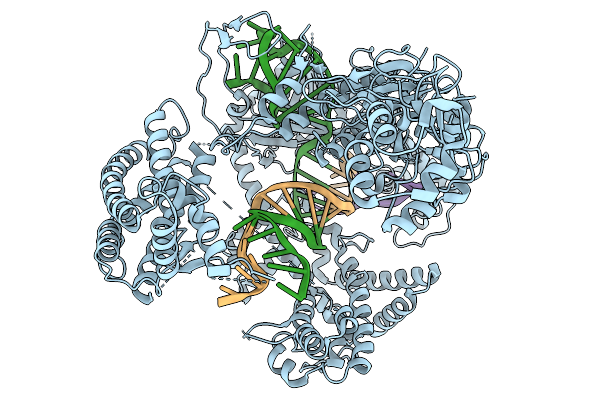 |
Organism: Francisella tularensis subsp. novicida, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.63 Å Release Date: 2026-02-11 Classification: DNA BINDING PROTEIN |
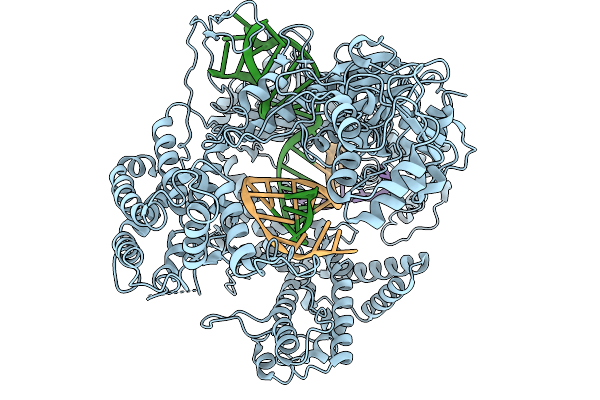 |
Organism: Francisella tularensis subsp. novicida, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.21 Å Release Date: 2026-02-11 Classification: DNA BINDING PROTEIN |
 |
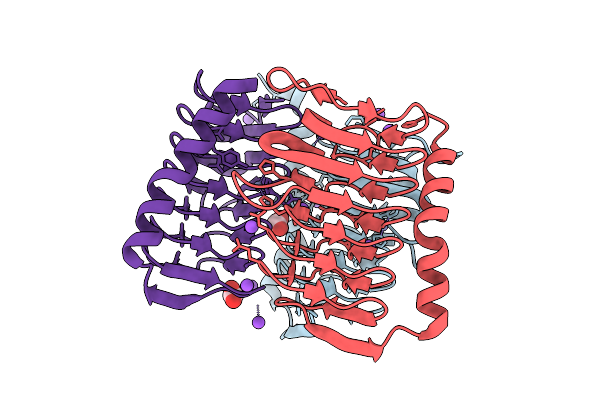
Crystal Structure Of Streptococcus Thermophilus Shp Pheromone Receptor Rgg3 In Complex With Rgg3Bp13
Organism: Streptococcus thermophilus lmg 18311, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2026-02-04 Classification: DNA BINDING PROTEIN Ligands: SO4, GOL, TRS |
 |
Organism: Caloramator australicus
Method: X-RAY DIFFRACTION Resolution:1.11 Å Release Date: 2025-12-10 Classification: METAL BINDING PROTEIN Ligands: ZN, NA, FMT |
 |
Organism: Caloramator australicus rc3, Unidentified
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-12-10 Classification: METAL BINDING PROTEIN Ligands: ZN, NA, EPE, MG |
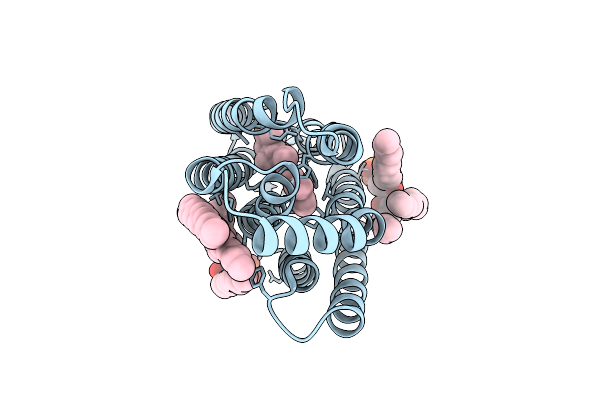 |
Organism: Unidentified
Method: X-RAY DIFFRACTION Resolution:2.07 Å Release Date: 2025-11-26 Classification: MEMBRANE PROTEIN Ligands: RET, 3PE, D12 |

