Search Count: 565
All
Selected
 |
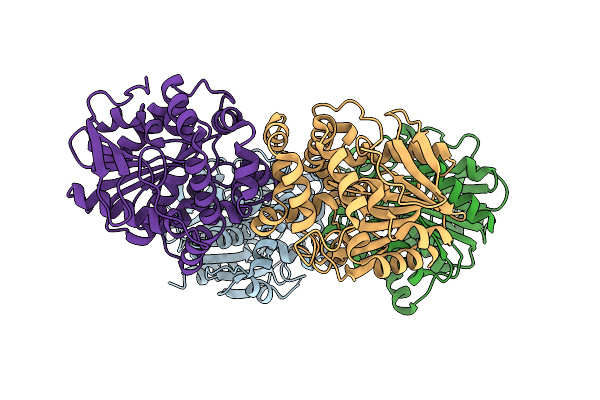
Crystal Structure Of The Fluoroacetate Dehalogenase Rpa1163 - Trp185Tyr/His280Ala With 2-Fluoro-3-Phenylpropanoic Acid
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2026-04-01 Classification: TRANSFERASE |
 |
Crystal Structure Of The Fluoroacetate Dehalogenase Rpa1163 - Lys181Met/Trp185Tyr/His280Ala With 2-Fluoro-3-Phenylpropanoic Acid
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2026-03-25 Classification: TRANSFERASE |
 |
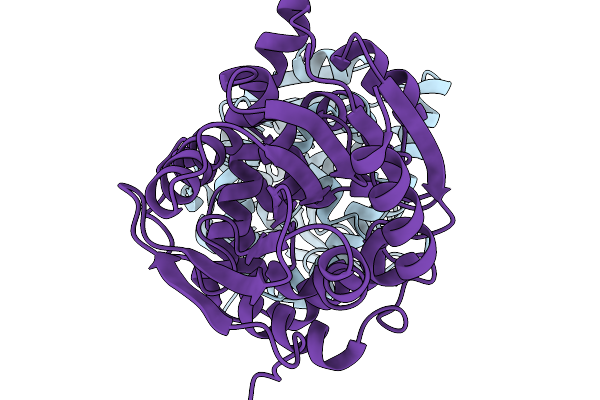
Crystal Structure Of The Fluoroacetate Dehalogenase Rpa1163 - Lys181Met/His280Ala With 2-Fluoro-3-Phenylpropanoic Acid
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2026-01-21 Classification: TRANSFERASE |
 |
Crystal Structure Of The Fluoroacetate Dehalogenase Rpa1163-His280Ala With (S)-2-Fluoro-3-Phenylpropanoic Acid
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2026-01-07 Classification: HYDROLASE |
 |
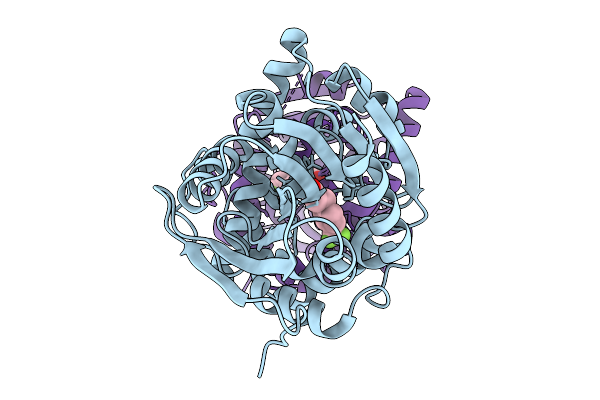
Crystal Structure Of The Fluoroacetate Dehalogenase Rpa1163 - Lys181Lys/Trp185Tyr/His280Ala With (S)-2-Fluoro-3-(4-(Trifluoromethyl)Phenyl)Propanoic Acid
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2025-12-31 Classification: TRANSFERASE Ligands: A1ES3 |
 |
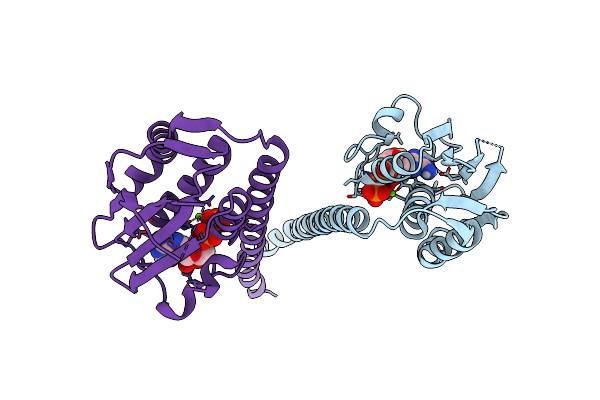
Crystal Structure Of The Fluoroacetate Dehalogenase Rpa1163 - Lys181Ser/Trp185Gly/His280Ala With (S)-2-Fluoro-3-(4-(Trifluoromethyl)Phenyl)Propanoic Acid
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:1.38 Å Release Date: 2025-12-31 Classification: TRANSFERASE Ligands: A1ES3 |
 |
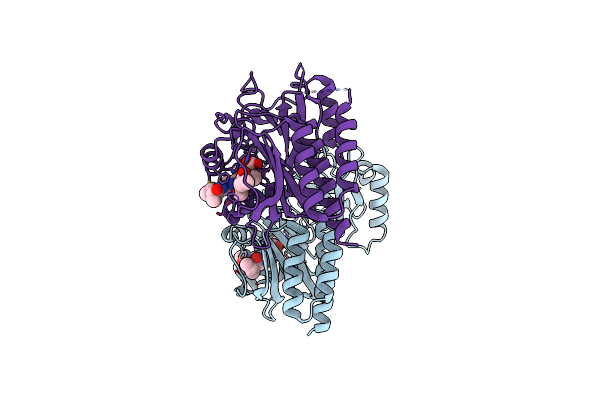
Crystal Structure Of The Histidine Kinase Domain Of Bacteriophytochrome Rpbphp2
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:3.19 Å Release Date: 2023-11-22 Classification: SIGNALING PROTEIN, TRANSFERASE Ligands: ATP, MG |
 |
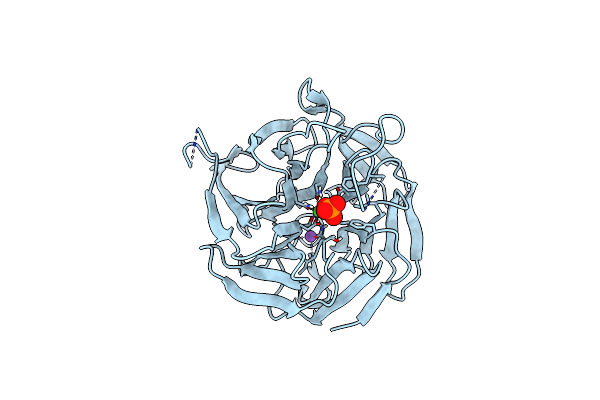
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:2.02 Å Release Date: 2023-08-16 Classification: FLUORESCENT PROTEIN Ligands: CYC |
 |
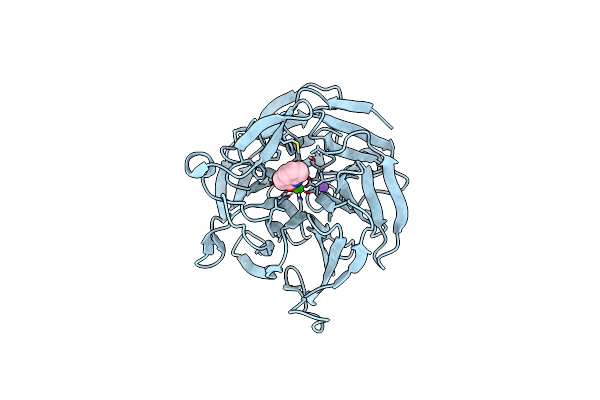
Crystal Structure Of Rpa3624, A Beta-Propeller Lactonase From Rhodopseudomonas Palustris, With Active-Site Bound Phosphate
Organism: Rhodopseudomonas palustris (strain atcc baa-98 / cga009)
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2023-01-11 Classification: HYDROLASE Ligands: PO4, CA, NA |
 |
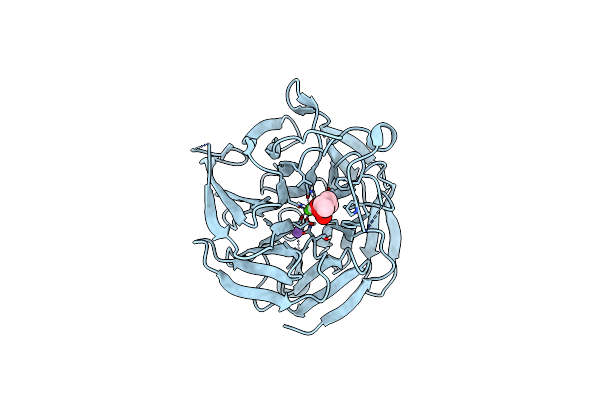
Crystal Structure Of Rpa3624, A Beta-Propeller Lactonase From Rhodopseudomonas Palustris, With Active-Site Bound 2-Hydroxyquinoline
Organism: Rhodopseudomonas palustris (strain atcc baa-98 / cga009)
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2023-01-11 Classification: HYDROLASE Ligands: OCH, CA, NA |
 |
Crystal Structure Of Rpa3624, A Beta-Propeller Lactonase From Rhodopseudomonas Palustris, With Active-Site Bound Tetrahedral Intermediate
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2023-01-11 Classification: HYDROLASE Ligands: CA, NA, SGU |
 |
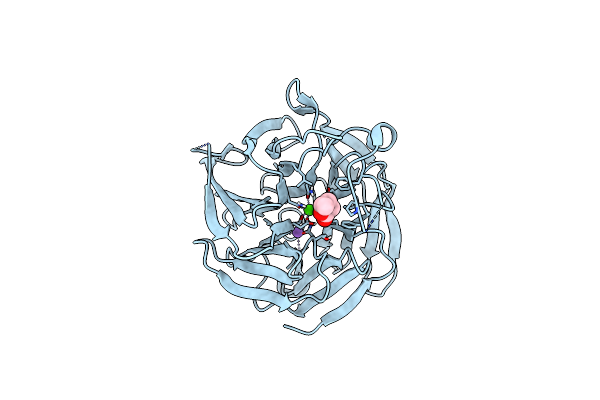
Crystal Structure Of Rpa3624, A Beta-Propeller Lactonase From Rhodopseudomonas Palustris, With Active-Site Bound Product
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2023-01-11 Classification: HYDROLASE Ligands: CA, NA, SJ3 |
 |
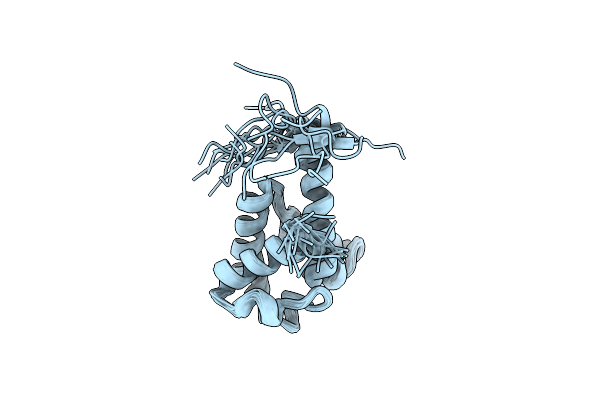
Crystal Structure Of Rpa3624, A Beta-Propeller Lactonase From Rhodopseudomonas Palustris, With Active-Site Bound (S)Gamma-Valerolactone
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2023-01-11 Classification: HYDROLASE Ligands: CA, NA, YVR |
 |
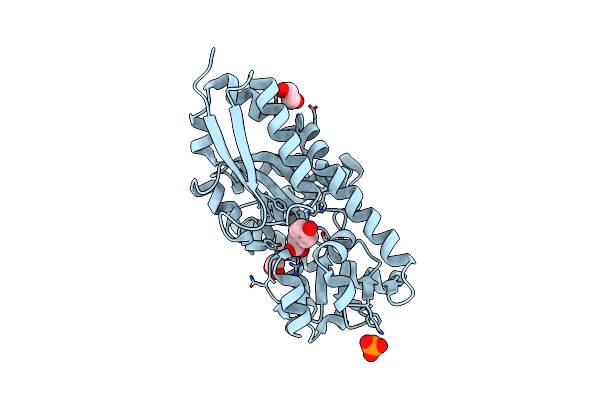
Solution Nmr Of The Specialized Apo-Acyl Carrier Protein (Rpa2022) From Rhodopseudomonas Palustris, Refined Without Rdcs. Northeast Structural Genomics Consortium Target Rpr324
Organism: Rhodopseudomonas palustris (strain atcc baa-98 / cga009)
Method: SOLUTION NMR Release Date: 2022-08-10 Classification: TRANSFERASE |
 |
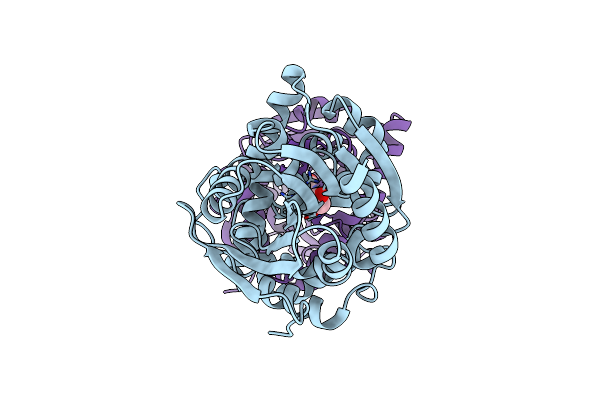
Organism: Rhodopseudomonas palustris (strain atcc baa-98 / cga009)
Method: X-RAY DIFFRACTION Resolution:1.10 Å Release Date: 2021-10-06 Classification: TRANSPORT PROTEIN Ligands: 4HP, EDO, PO4 |
 |
Time Resolved Structural Analysis Of The Full Turnover Of An Enzyme - 2256 Ms Covalent Intermediate 1
Organism: Rhodopseudomonas palustris (strain atcc baa-98 / cga009), Rhodopseudomonas palustris
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2019-09-25 Classification: HYDROLASE Ligands: FAH |
 |
Time Resolved Structural Analysis Of The Full Turnover Of An Enzyme - 6788 Ms
Organism: Rhodopseudomonas palustris, Rhodopseudomonas palustris (strain atcc baa-98 / cga009)
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2019-09-25 Classification: HYDROLASE Ligands: FAH |
 |
Fluoroacetate Dehalogenase, Room Temperature Structure Solved By Serial 1 Degree Oscillation Crystallography
Organism: Rhodopseudomonas palustris (strain atcc baa-98 / cga009)
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2019-03-27 Classification: HYDROLASE |
 |
Fluoroacetate Dehalogenase, Room Temperature Structure Solved By Serial 3 Degree Oscillation Crystallography
Organism: Rhodopseudomonas palustris (strain atcc baa-98 / cga009)
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2019-03-27 Classification: HYDROLASE Ligands: CA |
 |
Fluoroacetate Dehalogenase, Room Temperature Structure, Using First 1 Degree Of Total 3 Degree Oscillation
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2019-03-27 Classification: HYDROLASE Ligands: CA |

