Search Count: 1,227
All
Selected
 |
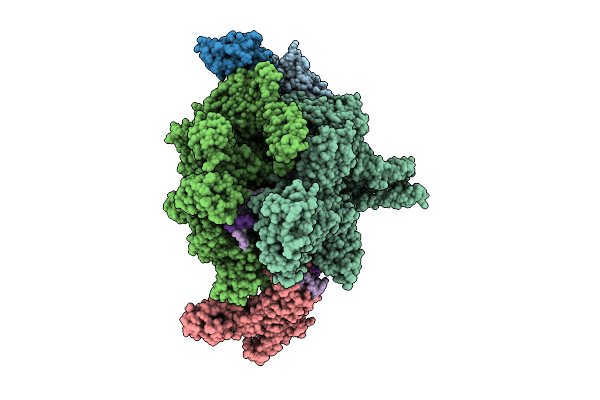
Cryo-Em Structure Of Vibrio Cholerae Rna Polymerase Transcription Activation Complex With Toxr Transcription Factor And Ompu Promoter Dna
Organism: Vibrio cholerae o395, Vibrio cholerae o1 biovar el tor str. n16961
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: TRANSCRIPTION Ligands: ZN, MG |
 |
Cryo-Em Structure Of Vibrio Cholerae Rna Polymerase Transcription Activation Complex With Toxr And Tcpp Transcription Factors And A Toxt Promoter Dna Fragment
Organism: Vibrio cholerae o395, Vibrio cholerae o1 biovar el tor str. n16961
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: TRANSCRIPTION Ligands: MG, ZN |
 |
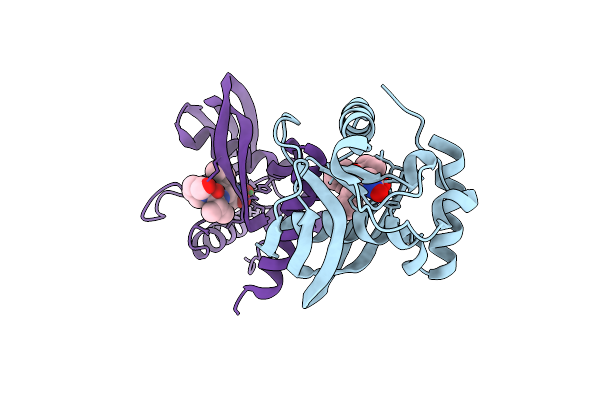
Organism: Vibrio cholerae o1 biovar el tor str. n16961
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2025-12-10 Classification: HYDROLASE Ligands: NI, BB2 |
 |
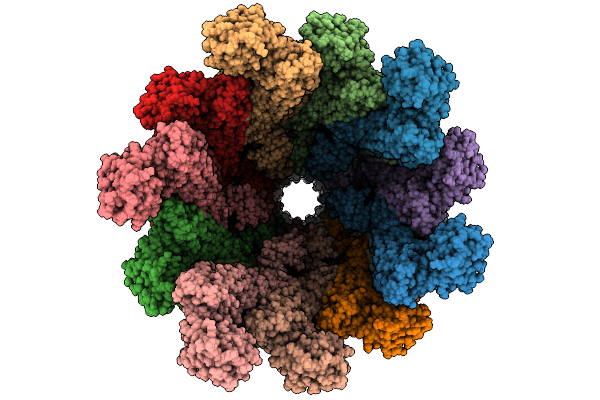
Organism: Vibrio cholerae o1 biovar el tor str. n16961
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2025-12-10 Classification: HYDROLASE Ligands: BB2, NI |
 |
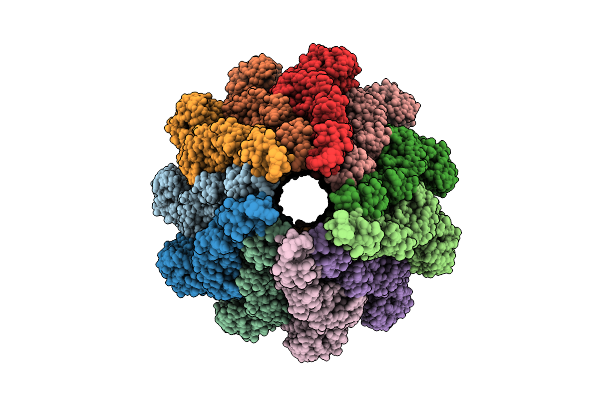
In Situ Structure Of The Sheathed Flad Flagellar Filament In Vibrio Cholerae
Organism: Vibrio cholerae o1 biovar el tor str. n16961
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: MOTOR PROTEIN |
 |
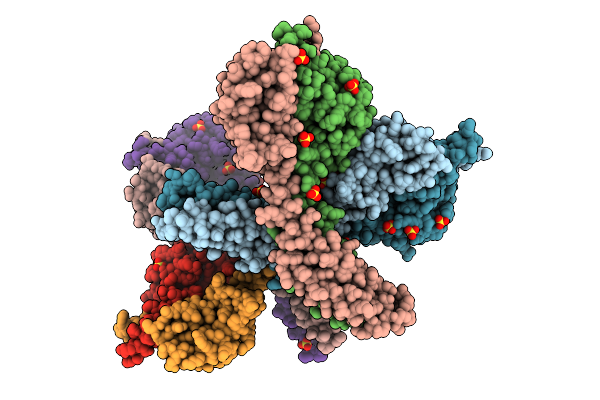
In Situ Unsheathed Flagellar Filament Of Vibrio Cholerae Resolved With Helical Reconstruction.
Organism: Vibrio cholerae o1 biovar el tor str. n16961
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: MOTOR PROTEIN |
 |
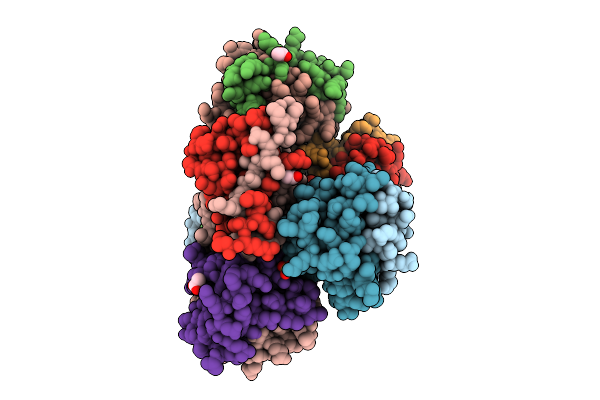
Crystal Structure Of The Histidine Kinase Vc2136 From Vibrio Cholerae Serotype O1
Organism: Vibrio cholerae o1 biovar el tor str. n16961
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2025-08-20 Classification: TRANSFERASE Ligands: SO4, CL |
 |
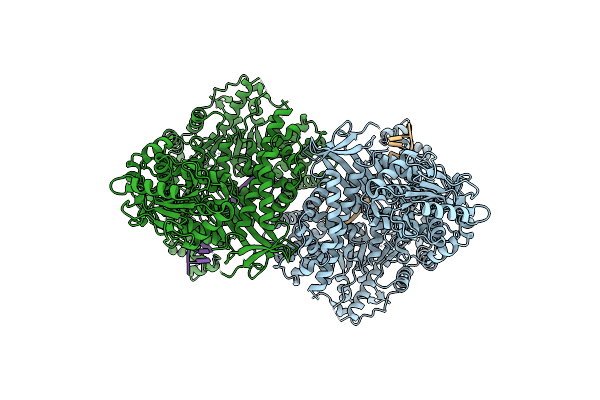
Organism: Vibrio cholerae serotype o1 (strain atcc 39315 / el tor inaba n16961)
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-07-30 Classification: TOXIN Ligands: SO4, ACT |
 |
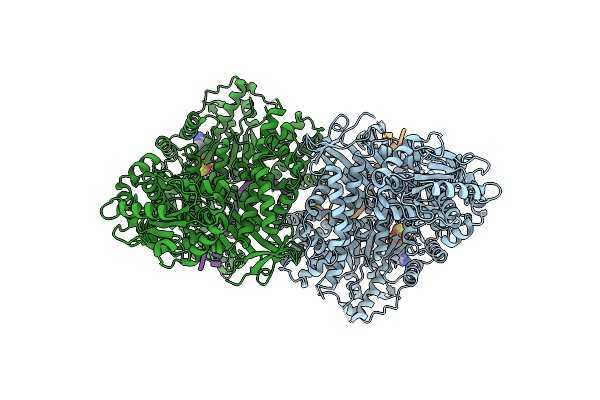
Organism: Vibrio cholerae o1 biovar el tor str. n16961, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-02-19 Classification: DNA BINDING PROTEIN/DNA |
 |
Organism: Vibrio cholerae o1 biovar el tor str. n16961, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-02-19 Classification: DNA BINDING PROTEIN/DNA Ligands: AGS, MG |
 |
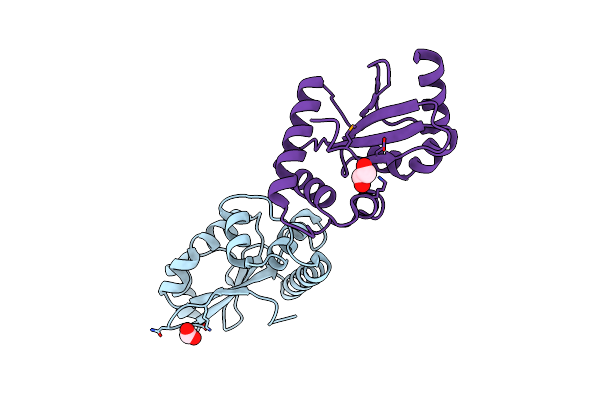
Makc And Makd Are Two Proteins Associated With A Tripartite Toxin Of Vibrio Cholerae
Organism: Vibrio cholerae o1 biovar el tor str. n16961
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2024-09-25 Classification: UNKNOWN FUNCTION |
 |
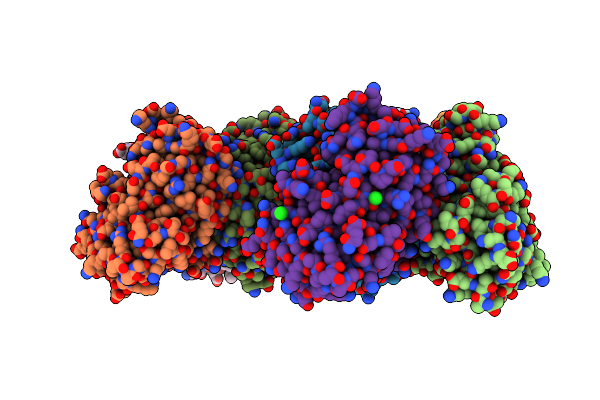
Crystal Structure Of The Thiol:Disulfide Interchange Protein Dsbc From Vibrio Cholerae
Organism: Vibrio cholerae o1 biovar el tor str. n16961
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2024-09-04 Classification: OXIDOREDUCTASE Ligands: FMT, CL, EDO |
 |
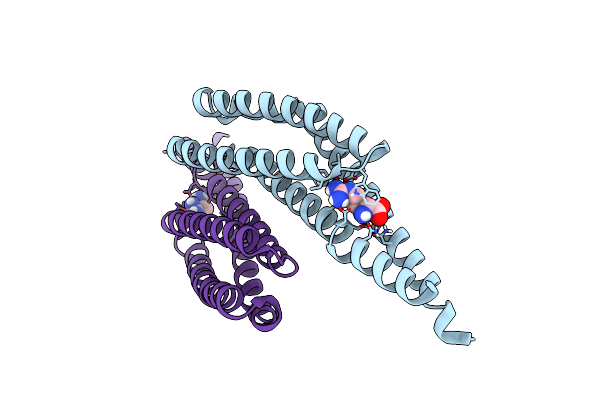
Crystal Structure Of The Substrate Binding Domain Protein Of The Abc Transporter Pbp2_Yxem From Vibrio Cholerae
Organism: Vibrio cholerae o1 biovar el tor str. n16961
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2024-08-28 Classification: TRANSPORT PROTEIN Ligands: PGO, CL, K, PGR, SO4 |
 |
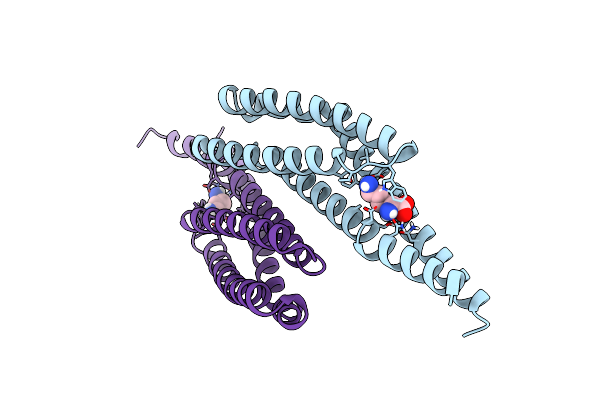
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2023-06-07 Classification: MEMBRANE PROTEIN Ligands: DAR |
 |
Organism: Vibrio cholerae
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2023-06-07 Classification: MEMBRANE PROTEIN Ligands: DLY |
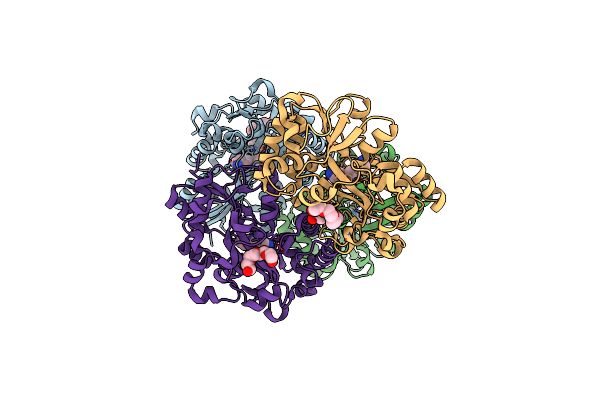 |
Organism: Vibrio cholerae o1 biovar el tor str. n16961
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2023-04-26 Classification: TRANSPORT PROTEIN Ligands: SPD, P6G, PG4 |
 |
Organism: Vibrio cholerae o1 biovar el tor str. n16961
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2023-04-26 Classification: TRANSPORT PROTEIN Ligands: NSD, PG4, P6G |
 |
Organism: Vibrio cholerae o1 biovar el tor str. n16961
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-01-18 Classification: METAL BINDING PROTEIN Ligands: PGE |
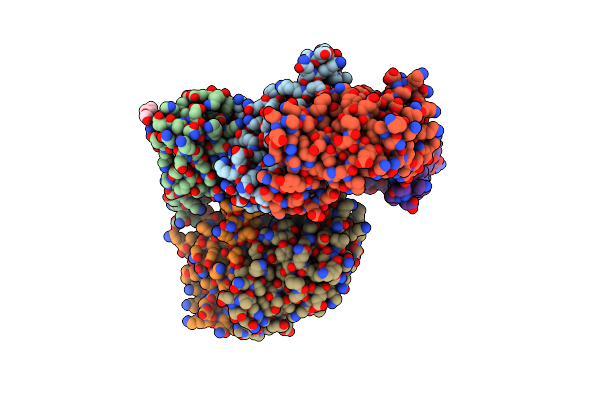 |
Organism: Vibrio cholerae o1 biovar el tor str. n16961
Method: X-RAY DIFFRACTION Resolution:2.41 Å Release Date: 2023-01-18 Classification: METAL BINDING PROTEIN Ligands: ZN, PGE |
 |
Organism: Vibrio cholerae o1 biovar el tor str. n16961
Method: X-RAY DIFFRACTION Resolution:2.95 Å Release Date: 2022-04-27 Classification: TOXIN |

