Search Count: 2,325
All
Selected
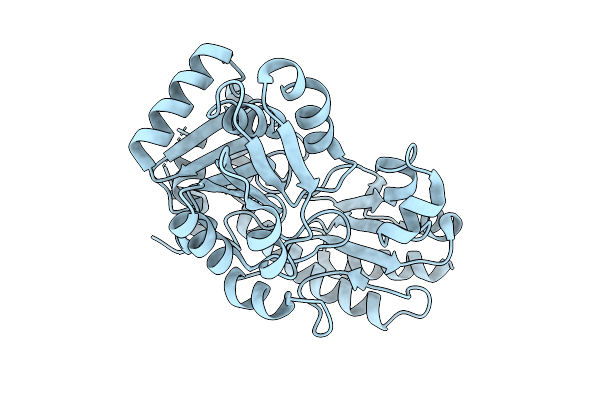 |
Organism: Staphylococcus aureus subsp. aureus mu50
Method: X-RAY DIFFRACTION Resolution:2.33 Å Release Date: 2026-04-15 Classification: LYASE |
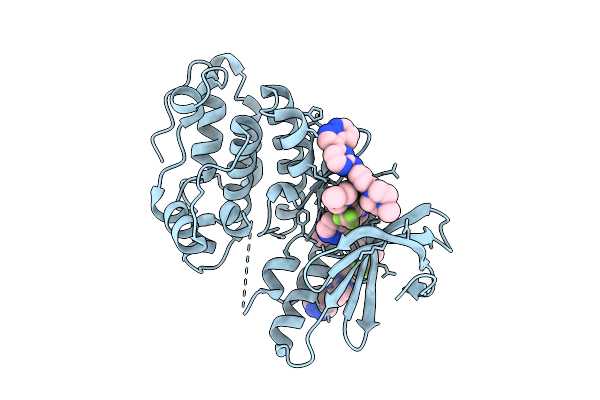 |
Staphylococcus Aureus Serine/Threonine Kinase Stk1 With Inhibitor Gw779439X
Organism: Staphylococcus aureus subsp. aureus n315
Method: X-RAY DIFFRACTION Resolution:2.07 Å Release Date: 2026-03-11 Classification: TRANSFERASE/INHIBITOR Ligands: A1CYJ, BEN, DMS |
 |
Organism: Staphylococcus aureus subsp. aureus mu50
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2026-01-21 Classification: STRUCTURAL PROTEIN |
 |
Organism: Staphylococcus aureus subsp. aureus mu50
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-01-21 Classification: STRUCTURAL PROTEIN Ligands: GOL |
 |
Organism: Rift valley fever virus
Method: ELECTRON MICROSCOPY Resolution:3.40 Å Release Date: 2025-12-24 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Organism: Rift valley fever virus, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2025-12-24 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
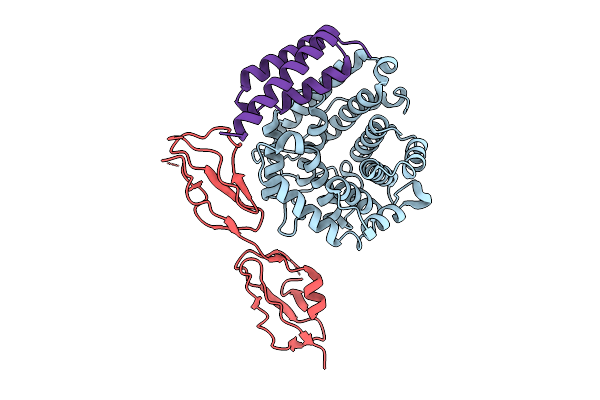 |
Organism: Homo sapiens, Staphylococcus aureus subsp. aureus mu50
Method: X-RAY DIFFRACTION Resolution:3.30 Å Release Date: 2025-12-10 Classification: IMMUNE SYSTEM |
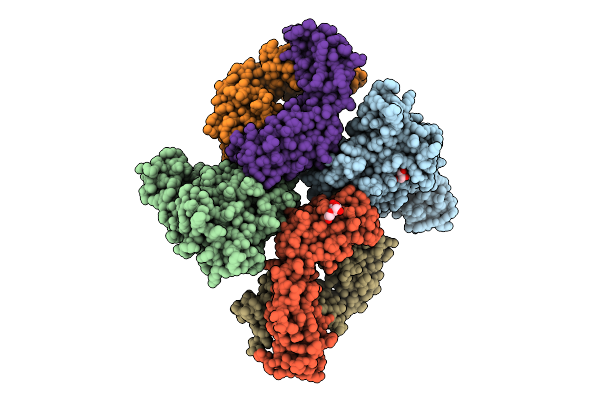 |
Organism: Rift valley fever virus, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2025-11-26 Classification: VIRAL PROTEIN Ligands: GOL, PEG |
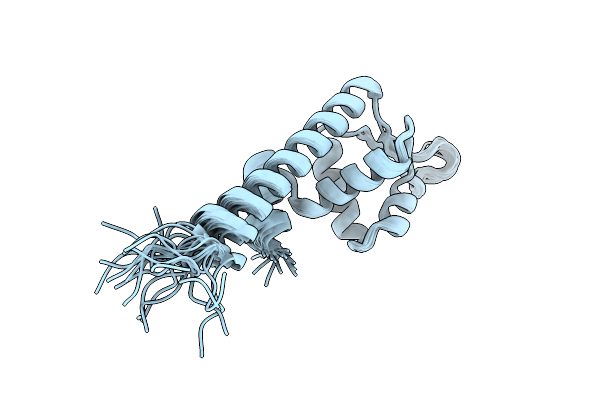 |
Organism: Staphylococcus aureus subsp. aureus n315
Method: SOLUTION NMR Release Date: 2025-10-15 Classification: DNA BINDING PROTEIN |
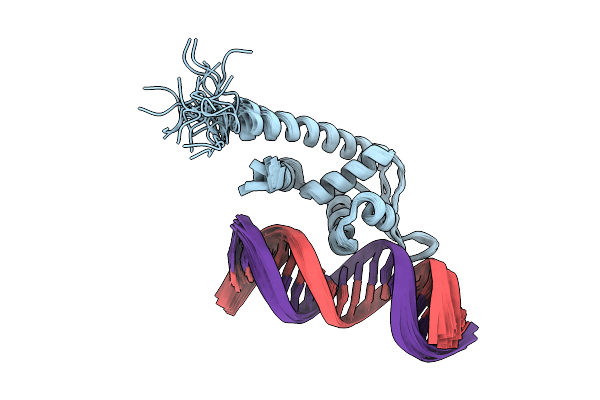 |
Organism: Staphylococcus aureus subsp. aureus n315, Synthetic construct
Method: SOLUTION NMR Release Date: 2025-10-15 Classification: DNA BINDING PROTEIN |
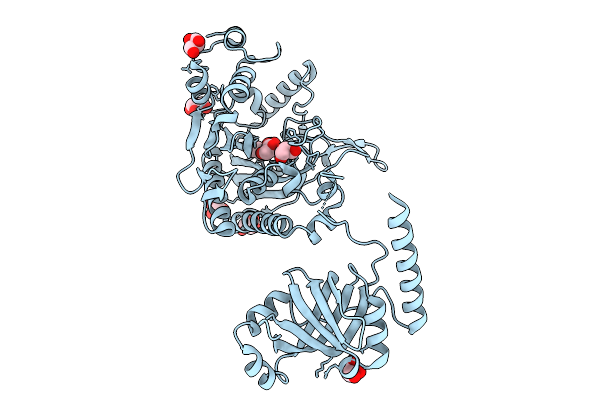 |
Organism: Staphylococcus aureus subsp. aureus mu50
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-09-10 Classification: LIGASE Ligands: LYS, GOL |
 |
Organism: Staphylococcus aureus subsp. aureus mu50
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2025-09-03 Classification: LYASE Ligands: SO4 |
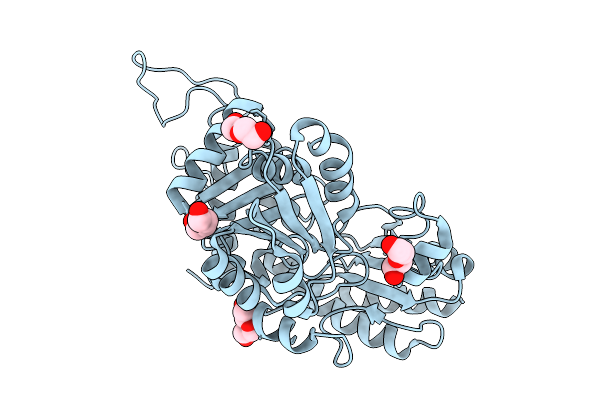 |
Organism: Staphylococcus aureus subsp. aureus mu50
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2025-09-03 Classification: LYASE Ligands: PEG |
 |
Organism: Staphylococcus aureus subsp. aureus mu50
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-09-03 Classification: LYASE Ligands: PGU |
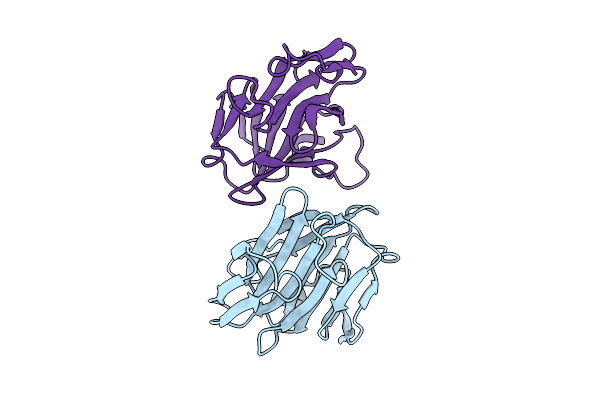 |
Vp8* Domain Of The Spike Protein Vp4 From Bovine P[3] Strain Of Rotavirus Species C
Organism: Bovine rotavirus c
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2025-08-20 Classification: VIRAL PROTEIN |
 |
Cryo-Em Structure Of Outward State Anhydromuropeptide Permease (Ampg) Complex With Glcnac-1,6-Anhmurnac
Organism: Homo sapiens, Synthetic construct, Yokenella regensburgei, Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.97 Å Release Date: 2025-07-16 Classification: MEMBRANE PROTEIN Ligands: 2YP |
 |
Organism: Vicugna pacos, Mus musculus, Escherichia coli, Staphylococcus aureus subsp. aureus n315, Streptococcus sp. 'group g'
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2025-06-11 Classification: BIOSYNTHETIC PROTEIN |
 |
Organism: Yokenella regensburgei
Method: ELECTRON MICROSCOPY Release Date: 2025-05-21 Classification: TRANSPORT PROTEIN |
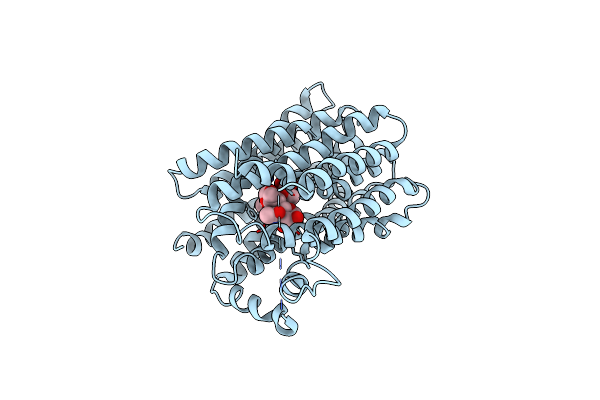 |
Cryo-Em Structure Of Inward-Facing Anhydromuropeptide Permease (Ampg) In Complex With Glcnac-1,6-Anhmurnac
Organism: Yokenella regensburgei
Method: ELECTRON MICROSCOPY Release Date: 2025-05-21 Classification: MEMBRANE PROTEIN Ligands: 2YP |
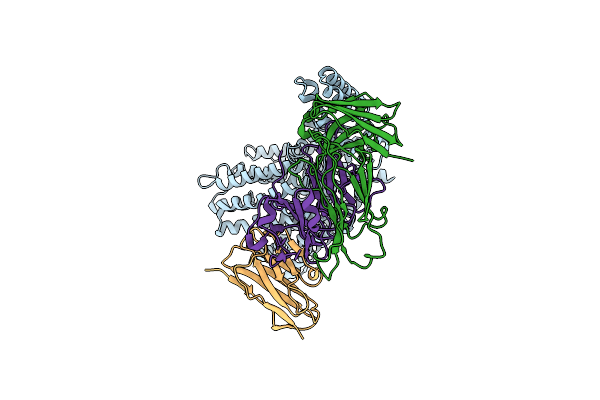 |
Cryo-Em Structure Of Outward State Anhydromuropeptide Permease (Ampg) G50W/L269W
Organism: Yokenella regensburgei, Escherichia coli, Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.11 Å Release Date: 2025-04-30 Classification: PEPTIDE BINDING PROTEIN |

