Search Count: 2,771
All
Selected
 |
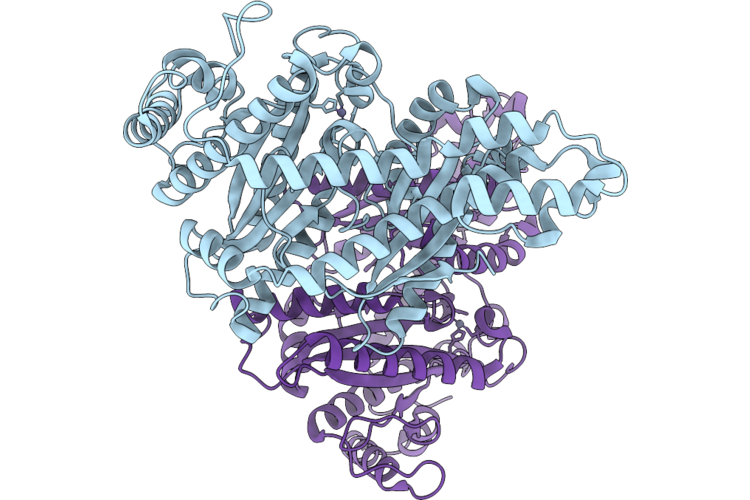
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Hexamer
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.13 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Dimer, Major State, Active Conformation
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.15 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Dimer, Minor State
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.45 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Reducing Conditions, Hexamer
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.06 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Reducing Conditions, Dimer, Major State, Inactive Conformation
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.12 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Reducing Conditions, Dimer, Minor State
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.27 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
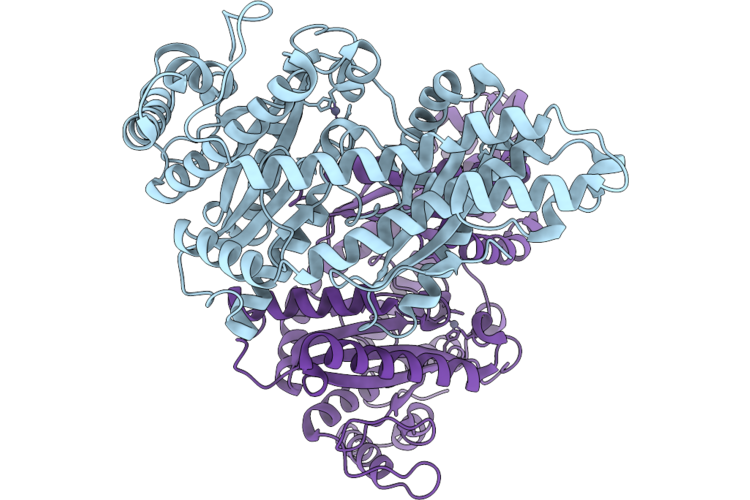
Cryo-Em Structure Of H. Neapolitanus Csosca C283A/C284A Inactive Mutant, Hexamer
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.08 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca C283A/C284A Inactive Mutant, Dimer, State 1
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.18 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca C283A/C284A Inactive Mutant, Dimer, State 2
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.22 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
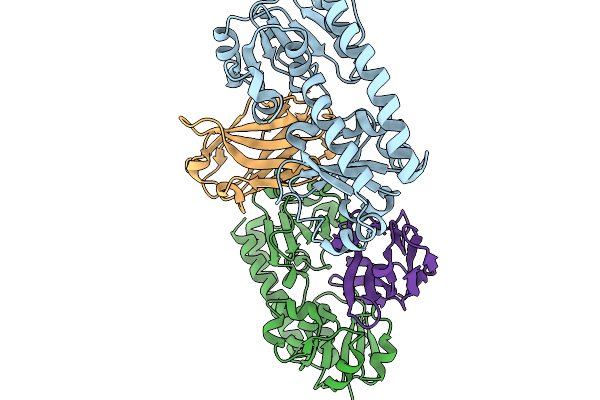
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: RIBOSOME Ligands: ZN |
 |
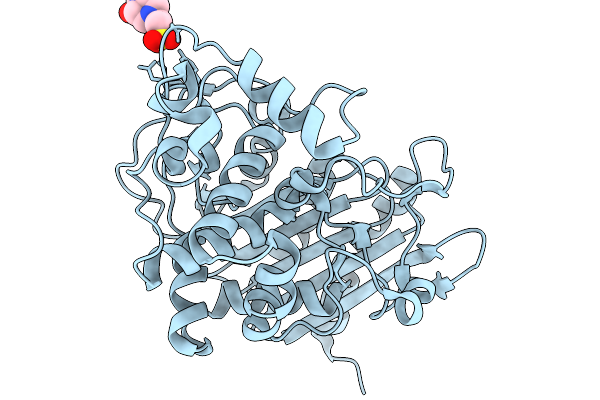
Crystal Structure Of Ftsb From Streptococcus Pyogenes In Complex With Nb1 Nanobody
Organism: Streptococcus pyogenes, Vicugna pacos
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2026-03-25 Classification: IMMUNE SYSTEM |
 |
Organism: Acinetobacter baumannii
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: EPE |
 |
Organism: Acinetobacter baumannii
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: NXL |
 |
Organism: Acinetobacter baumannii
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: NXL |
 |
Organism: Acinetobacter baumannii
Method: X-RAY DIFFRACTION Resolution:2.11 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: NXL |
 |
Crystal Structure Of The Relseq N-Terminal Domain From Streptococcus Equisimilis In Complex With Pppgpp
Organism: Streptococcus dysgalactiae subsp. equisimilis
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2026-03-18 Classification: TRANSFERASE Ligands: GOL, MN, 0O2, CL |
 |
Crystal Structure Of The A/Puerto Rico/8/1934 (H1N1) Influenza Virus Hemagglutinin In Complex With Fusion Inhibitor Clips Peptide Cp121132 (Cp-S1)
Organism: Influenza a virus (a/puerto rico/8/1934(h1n1)), Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2026-03-18 Classification: VIRAL PROTEIN Ligands: NAG, PEG, FUC, SEZ |
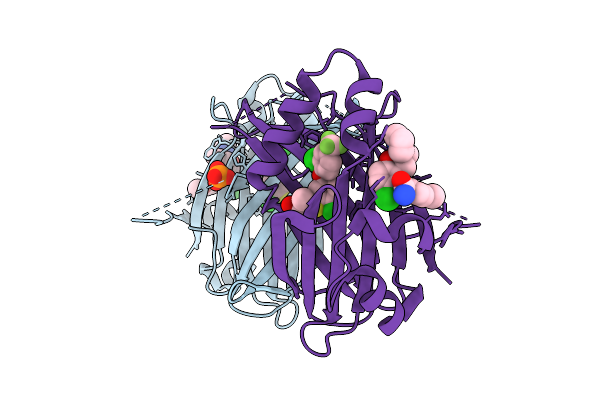 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-03-11 Classification: TRANSCRIPTION Ligands: U3O, A1ICN, PO4 |
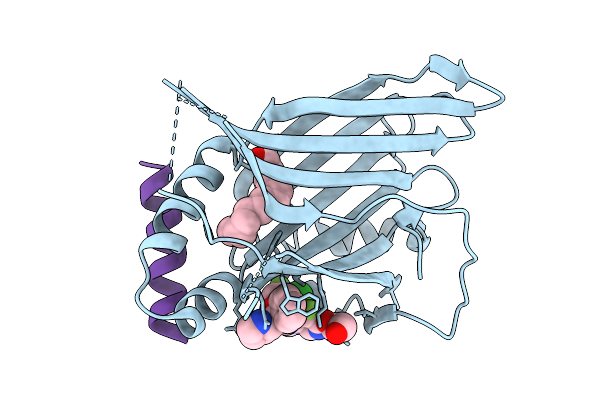 |
Organism: Homo sapiens, Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2026-03-11 Classification: TRANSCRIPTION Ligands: MYR, WCF |
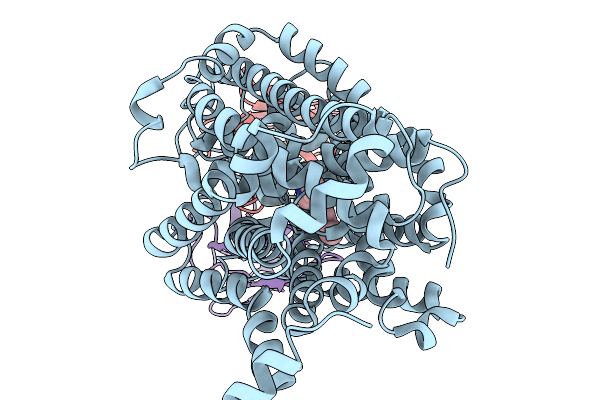 |
Structure Of Human Serotonin Transporter Bound To Small Molecule Zpzd In Lipid Nanodisc And Nacl
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: TRANSPORT PROTEIN/IMMUNE SYSTEM/INHIBITOR Ligands: CL, A1CIZ |

