Search Count: 39,995
All
Selected
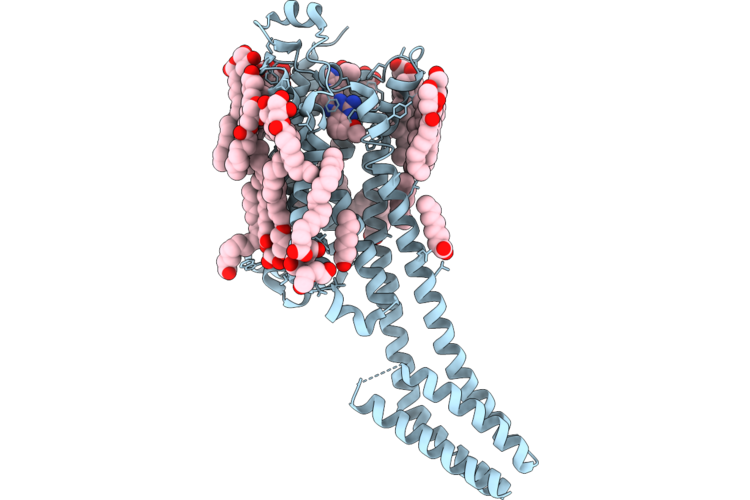 |
Crystal Structure Of A2A Adenosine Receptor A2Ar-Bril In Complex With Compound50
Organism: Homo sapiens, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.59 Å Release Date: 2026-05-20 Classification: MEMBRANE PROTEIN |
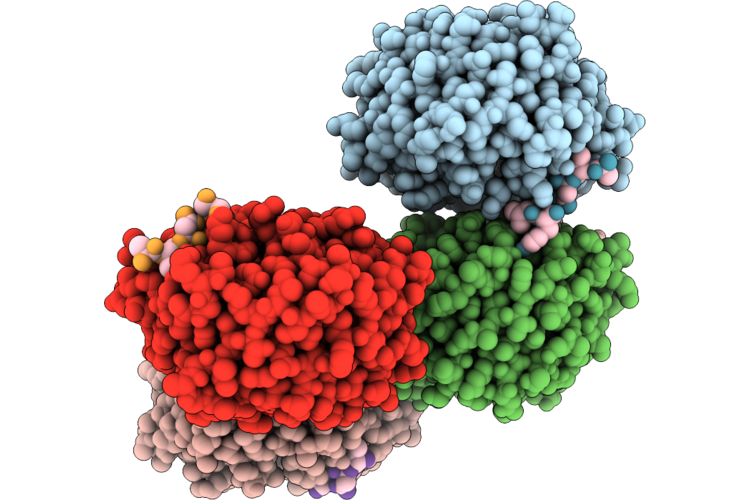 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-05-20 Classification: PEPTIDE BINDING PROTEIN |
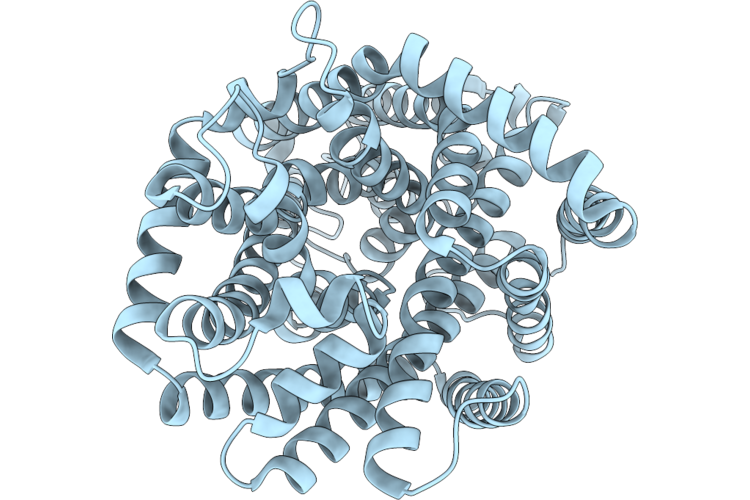 |
Organism: Psychrobacter sp. d2
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-05-20 Classification: TRANSPORT PROTEIN |
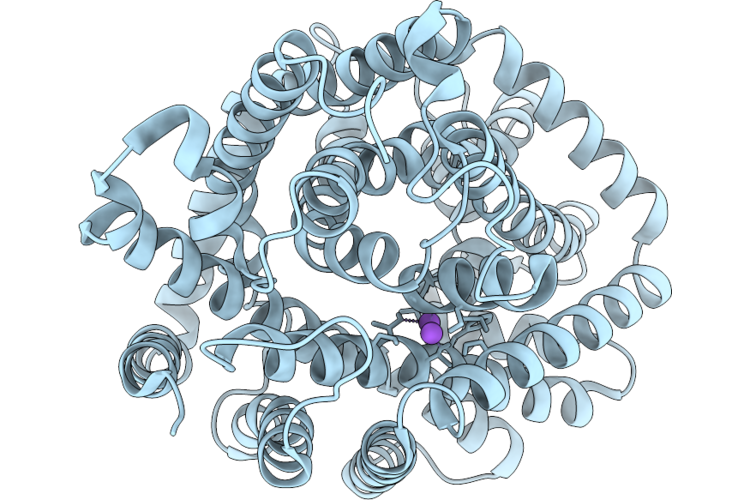 |
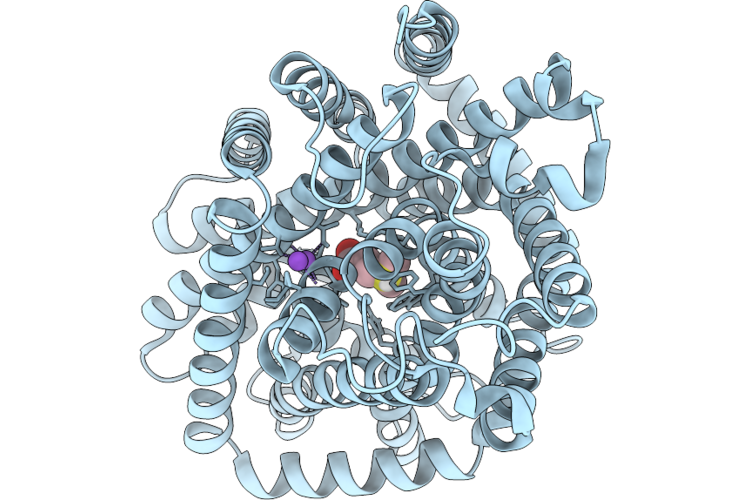
Cryo-Em Structure Of Dddt G101D In Substrate-Free Outward Open Conformation
Organism: Psychrobacter sp. d2
Method: ELECTRON MICROSCOPY Resolution:2.66 Å Release Date: 2026-05-20 Classification: TRANSPORT PROTEIN |
 |
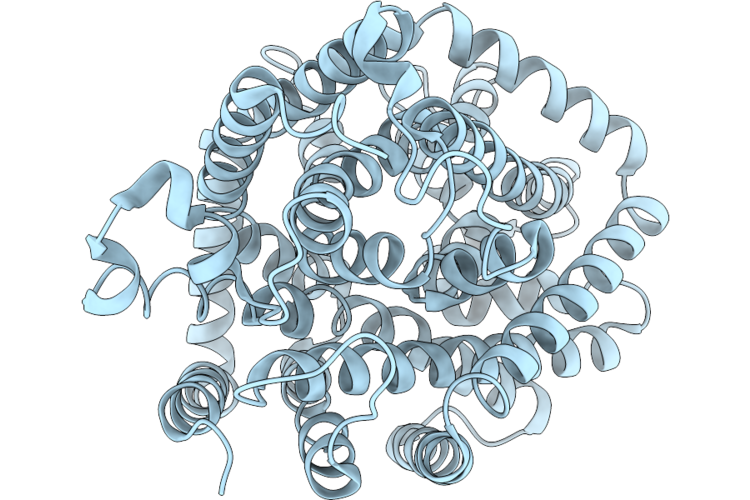
Organism: Psychrobacter sp. d2
Method: ELECTRON MICROSCOPY Resolution:2.52 Å Release Date: 2026-05-20 Classification: TRANSPORT PROTEIN Ligands: DQY, NA |
 |
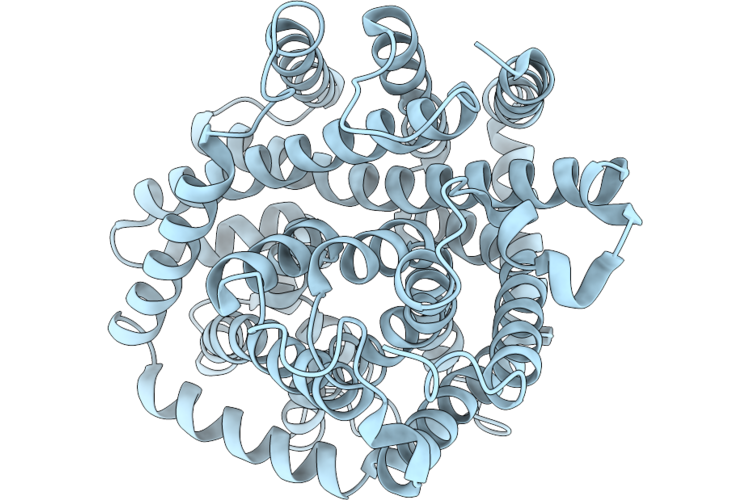
Cryo-Em Structure Of Dddt In Closed Substrate-Free Conformation In The Presence Of Potassium Ions And Dimethylsulfoniopropionate
Organism: Psychrobacter sp. d2
Method: ELECTRON MICROSCOPY Resolution:3.18 Å Release Date: 2026-05-20 Classification: TRANSPORT PROTEIN |
 |
Organism: Psychrobacter sp. d2
Method: ELECTRON MICROSCOPY Resolution:3.29 Å Release Date: 2026-05-20 Classification: TRANSPORT PROTEIN |
 |
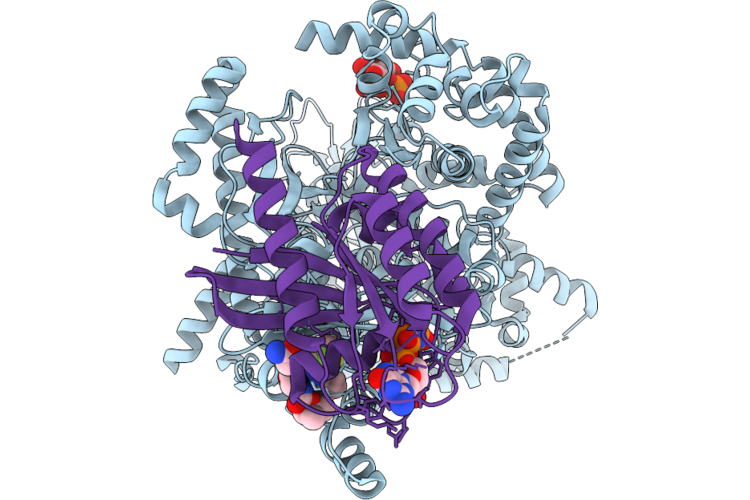
Cryo-Em Structure Of The Class 3 Pi3K Alpha/Kras Complex On Popc/Pops Nanodiscs
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: Transferase/Hydrolase Ligands: A1AZD, MG, GNP |
 |
Cryo-Em Structure Of The Pi3K Alpha/Kras Complex On Popc/Pops/Pip2 Nanodiscs Low-Pass Filtered To 5 Angstroms
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: Transferase/Hydrolase Ligands: PBU, A1AZD, MG, GNP |
 |
Organism: Nocardia brasiliensis
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: TRANSFERASE Ligands: MG |
 |
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: CA |
 |
Structure Of Geobacillus Stearothermophilus Rnase P Ribozyme Sub-Conformation 1
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: CA |
 |
Structure Of Geobacillus Stearothermophilus Rnase P Ribozyme Sub-Conformation 2
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: CA |
 |
Structure Of Geobacillus Stearothermophilus Rnase P Ribozyme Sub-Conformation 3
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: CA |
 |
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: MG |
 |
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: MG |
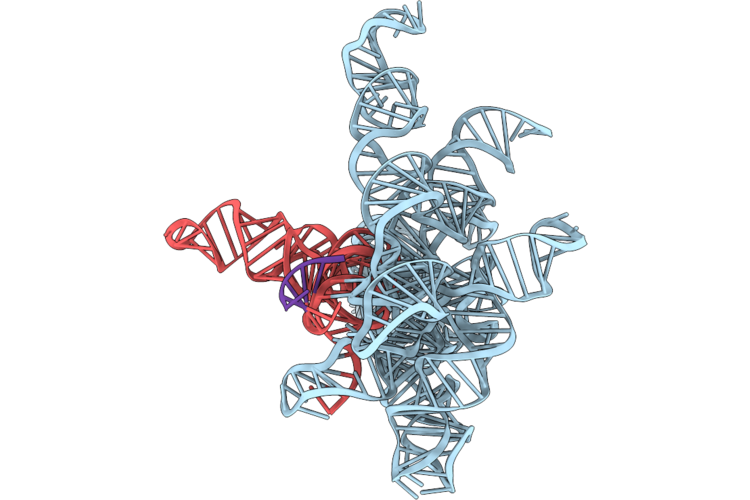 |
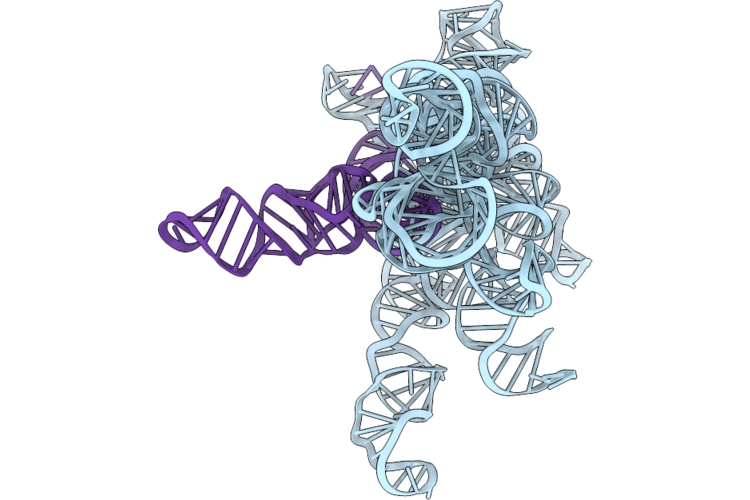
Structure Of Geobacillus Stearothermophilus Rnase P Ribozyme In Complex With Precursor Trna In 5 Mm Ca2+
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: CA |
 |
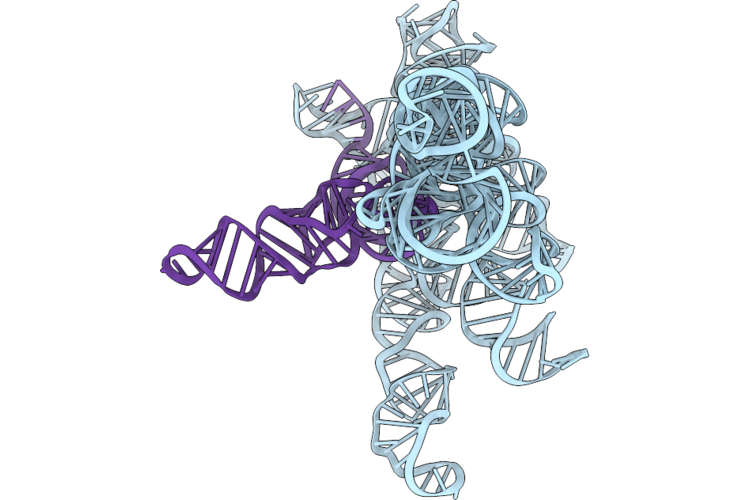
Structure Of Geobacillus Stearothermophilus Rnase P Ribozyme In Complex With Mature Trna In 5 Mm Ca2+
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: CA |
 |
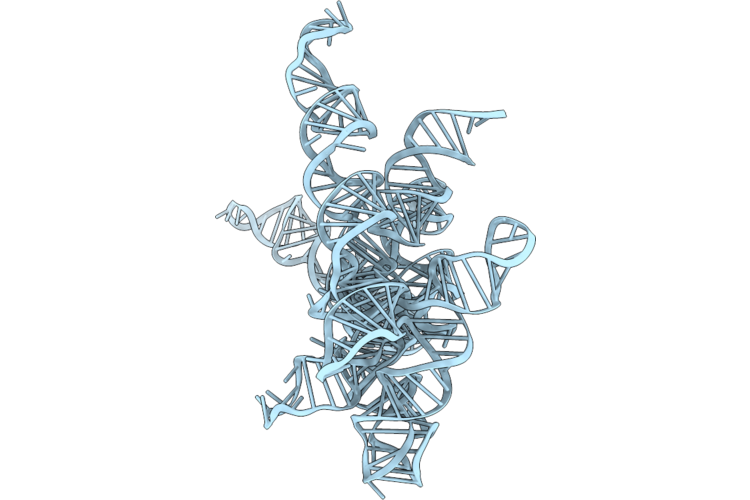
Structure Of Geobacillus Stearothermophilus Rnase P Ribozyme In Complex With Mature Trna In 10 Mm Ca2+
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: CA |
 |
Structure Of Geobacillus Stearothermophilus Rnase P Ribozyme Tetraloop Mutant (Sub-Conformation 1)
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: CA |

