Deposition Date
2025-09-05
Release Date
2025-11-12
Last Version Date
2025-12-17
Entry Detail
PDB ID:
9Y5Y
Keywords:
Title:
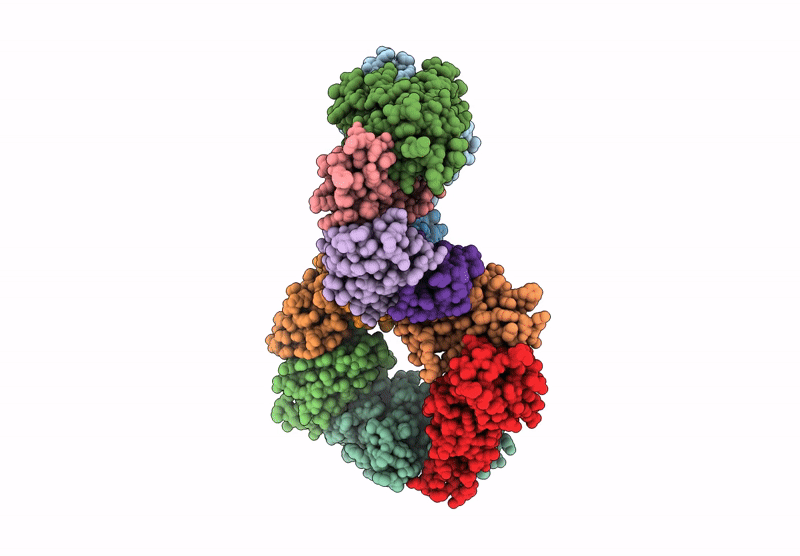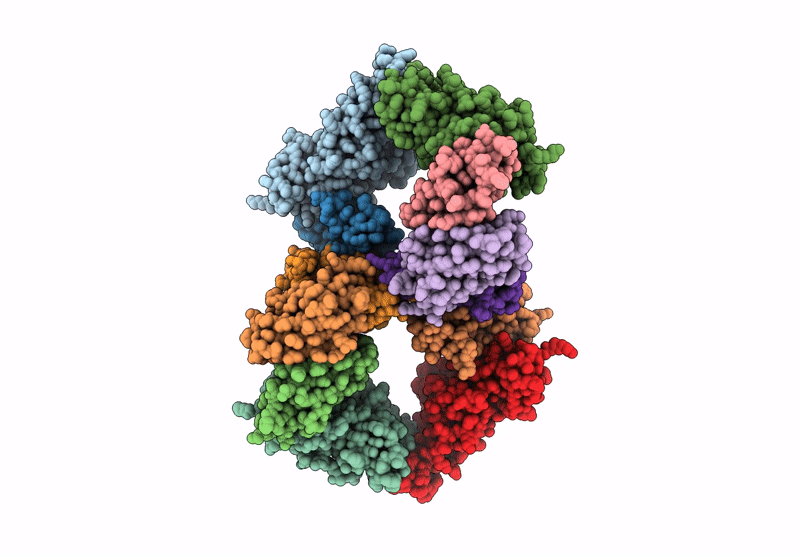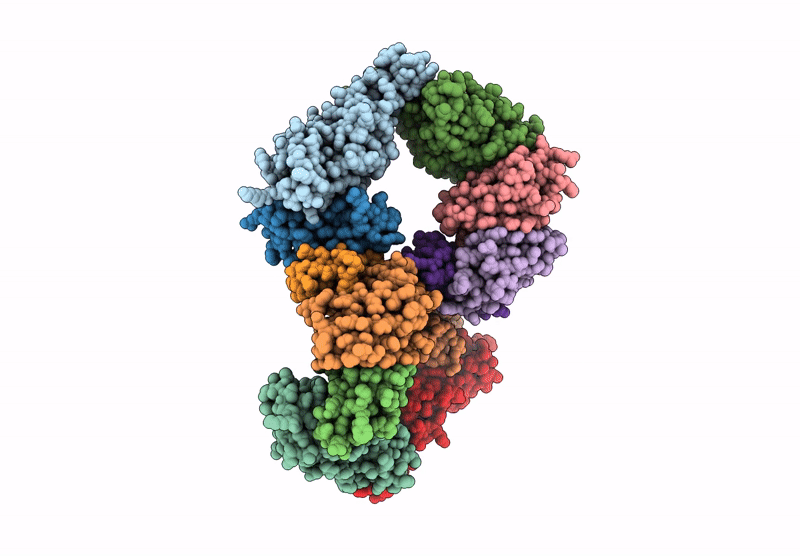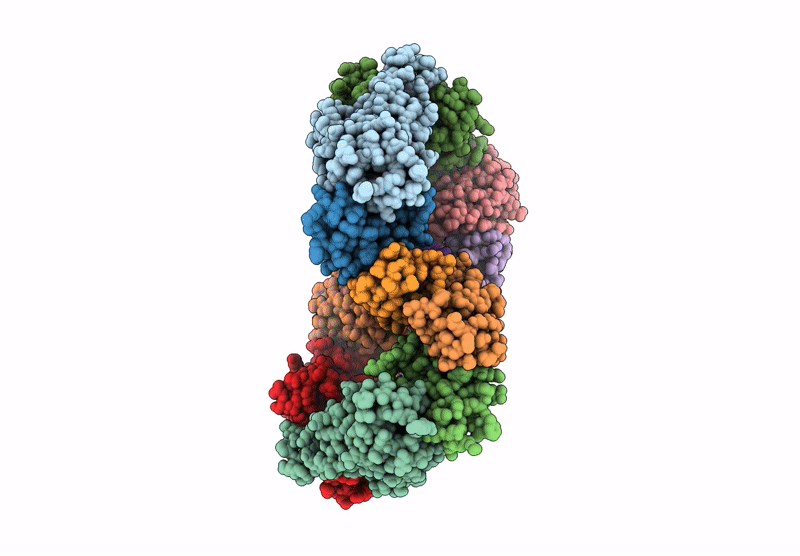
Structure of the Omicron Spike RBD bound by the monobody s19382 (local refinement from dimerized Spike protein ECDs)
Biological Source:
Source Organism(s):
Severe acute respiratory syndrome coronavirus 2 (Taxon ID: 2697049)
synthetic construct (Taxon ID: 32630)
synthetic construct (Taxon ID: 32630)
Expression System(s):
Method Details:
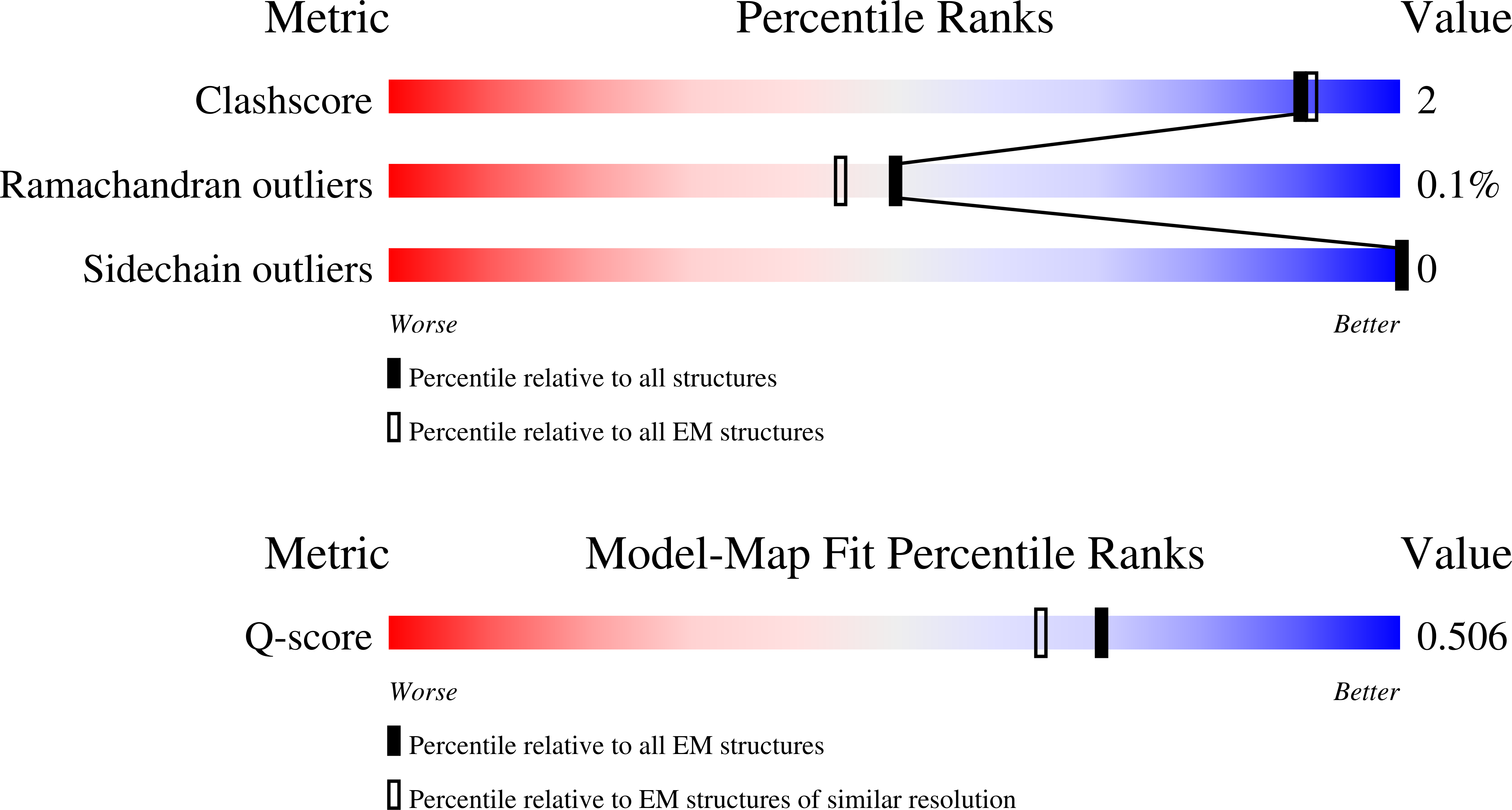
Experimental Method:
Resolution:
3.16 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE