Abstact
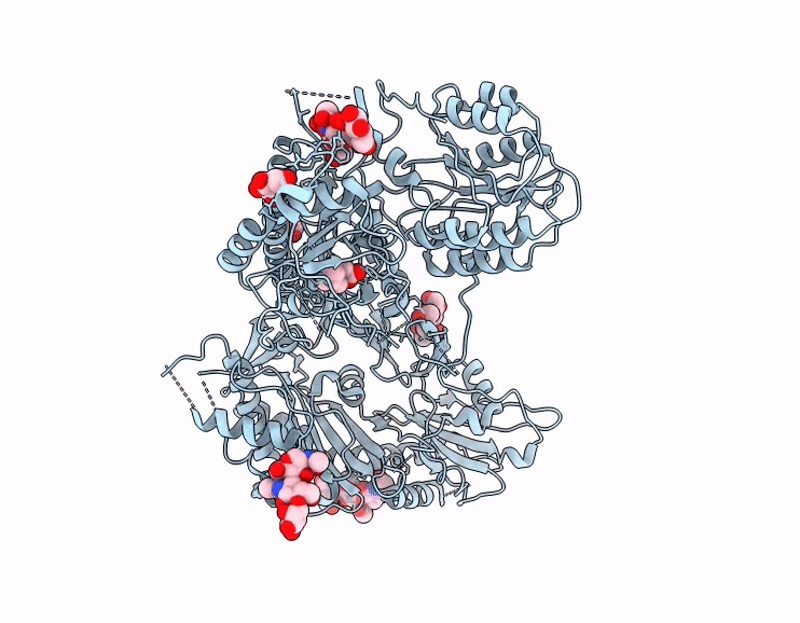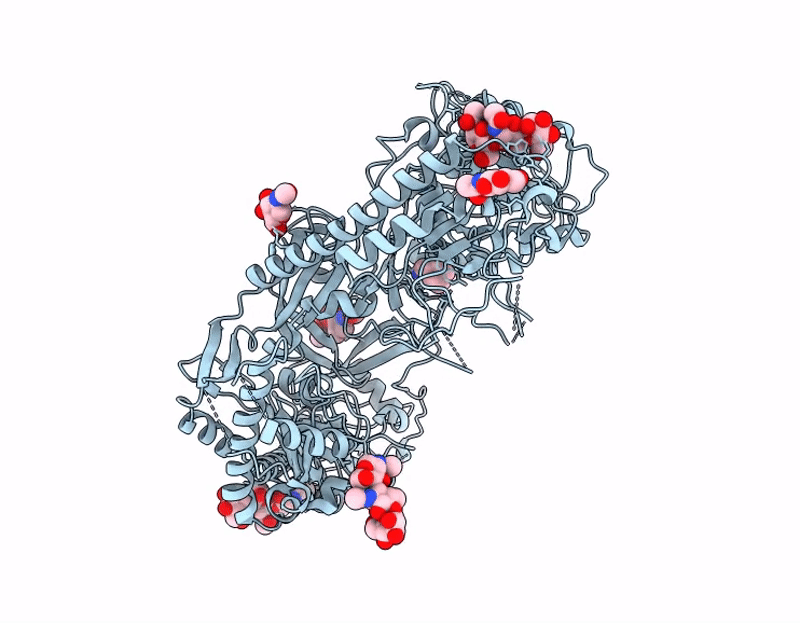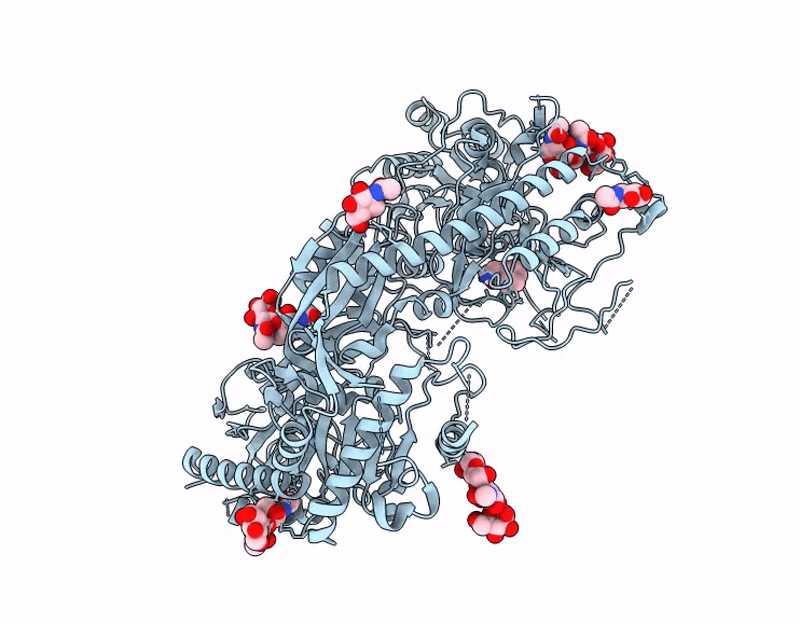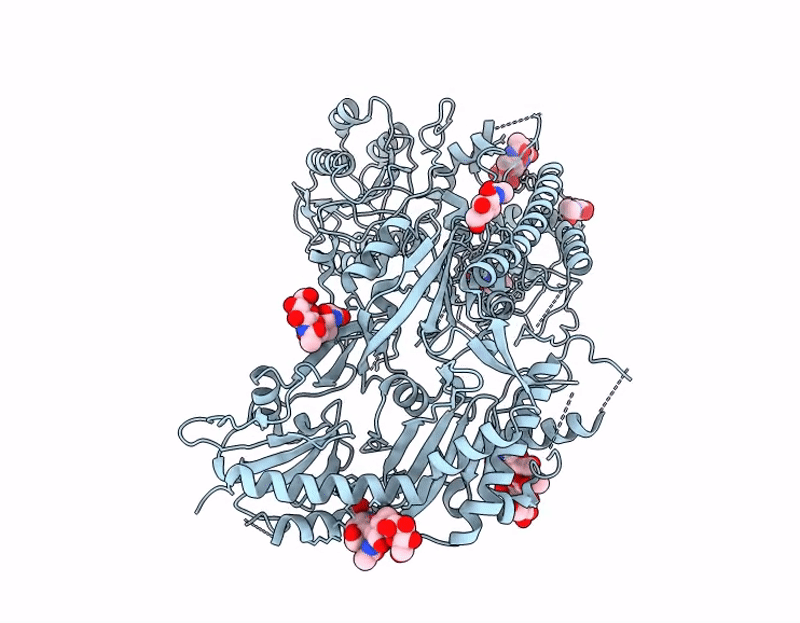
Crisugabalin, a recently approved third-generation GABA analogue with a unique cage-like tricyclic scaffold, shows superior efficacy and safety over pregabalin and mirogabalin for treating neuropathic pain. Through integrated biophysical, structural, and computational approaches, we elucidate the molecular basis of its enhanced pharmacological profile. Dissociation kinetic studies revealed that crisugabalin exhibited the slowest dissociation kinetics from the α2δ1 subunit (τ = 32.05, 80.00, 111.11 min for pregabalin, mirogabalin, and crisugabalin) but the fastest dissociation from the α2δ2 subunit (τ = 8.70, 16.39, 5.78 min for pregabalin, mirogabalin, and crisugabalin). Cryo-EM structures demonstrated crisugabalin's superior binding affinity for α2δ1 over gabapentin and l-leucine, driven by enhanced hydrogen bonding and hydrophobic contacts, alongside volumetric expansion of the l-leucine binding pocket. Molecular dynamics (MD) simulations identified significantly more persistent hydrogen bonding by crisugabalin (66.3% average occupancy) relative to pregabalin (28.3%). Random Acceleration Molecular Dynamics (RAMD) simulations revealed that ligand dissociation primarily proceeds via Pathway A (along the β2, β3, and β1 segments), and τRAMD calculations correctly ranked the ligand residence times, yielding values of 0.18 ns for pregabalin and 2.88 ns for crisugabalin. Furthermore, the binding free energies for pregabalin, mirogabalin, and crisugabalin were -21.64, -31.30, and -34.99 kcal/mol, calculated by MM/GBSA. The decomposition of the binding free energy components revealed that crisugabalin exhibits a dual-action mechanism characterized by enhanced hydrophobic interactions (-28.46 kcal/mol) and favorable entropic contributions (3.03 kcal/mol). This unique binding behavior stems from its cage-like tricyclic scaffold, an unprecedented substructure in drug molecules. These findings establish the cage-like tricyclic motif as a novel pharmacophore that simultaneously optimizes binding entropy and enthalpy, providing a blueprint for next-generation voltage-gated calcium channel modulators. MD, τRAMD, and MM-GBSA used in this study are powerful computational tools for rational drug design, particularly for optimizing compounds with prolonged target residence times.