Deposition Date
2025-09-26
Release Date
2026-02-18
Last Version Date
2026-04-15
Entry Detail
PDB ID:
9SSM
Keywords:
Title:
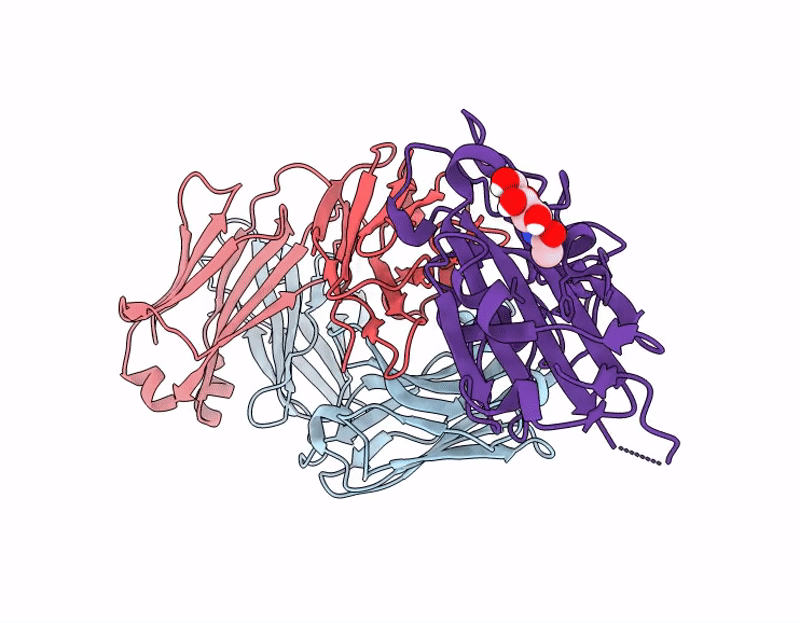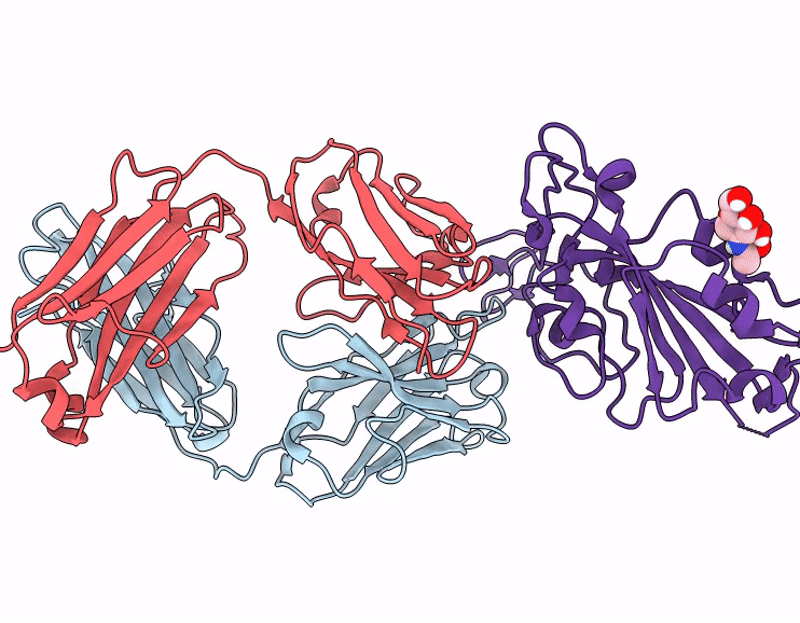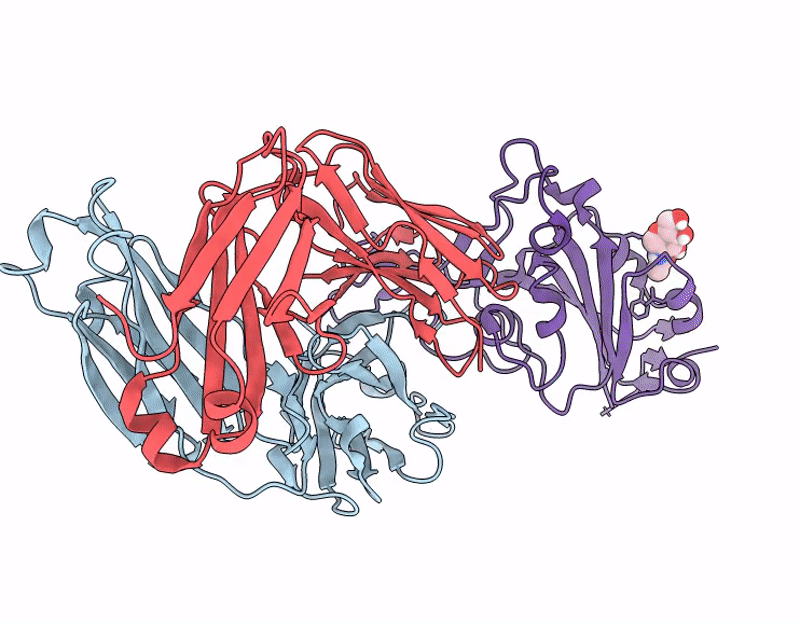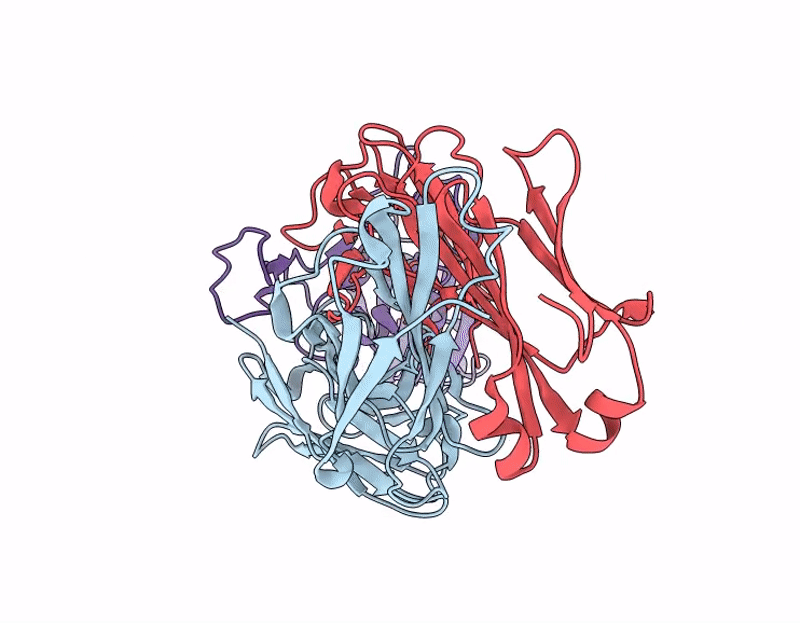
Crystal structure of 084-7D Fab bound to SARS-CoV-2 Beta RBD
Biological Source:
Source Organism(s):
Homo sapiens (Taxon ID: 9606)
Severe acute respiratory syndrome coronavirus 2 (Taxon ID: 2697049)
Severe acute respiratory syndrome coronavirus 2 (Taxon ID: 2697049)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.70 Å
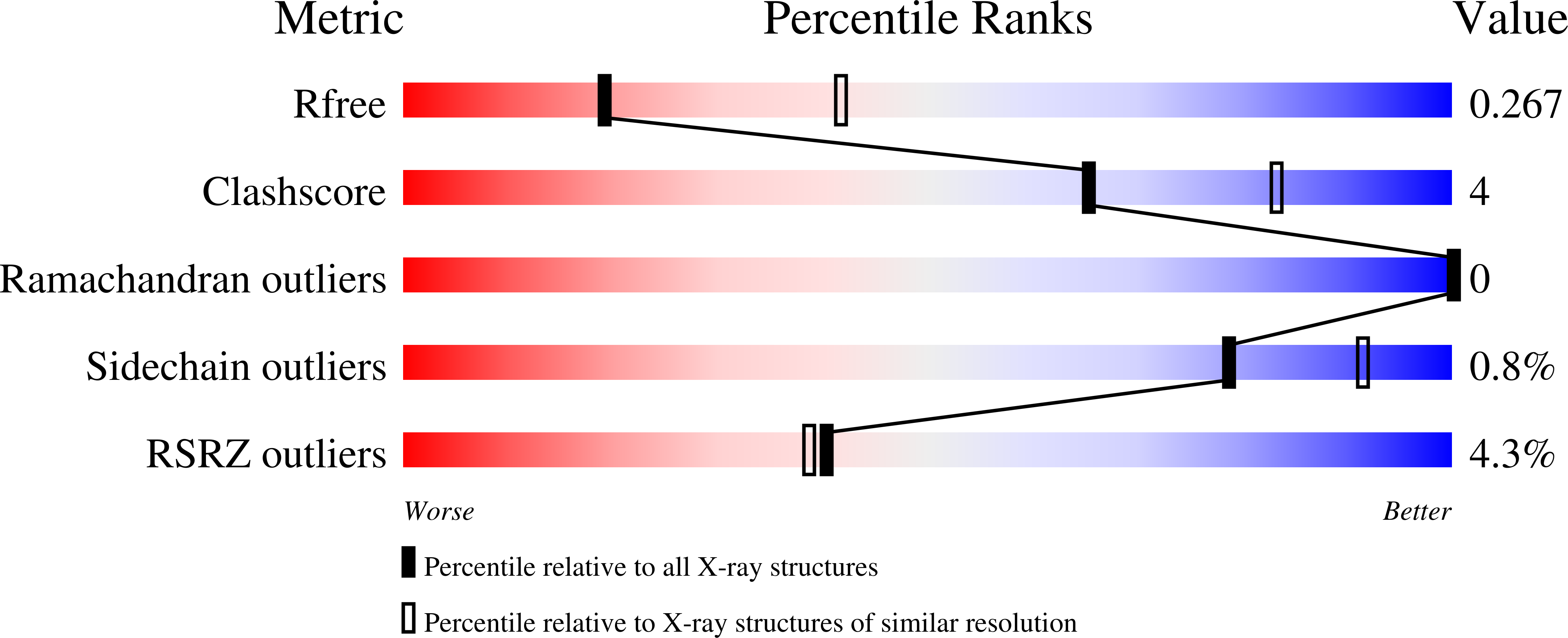
R-Value Free:
0.26
R-Value Work:
0.22
R-Value Observed:
0.22
Space Group:
P 21 21 21