Deposition Date
2025-07-06
Release Date
2026-05-06
Last Version Date
2026-05-06
Entry Detail
PDB ID:
9PFT
Keywords:
Title:
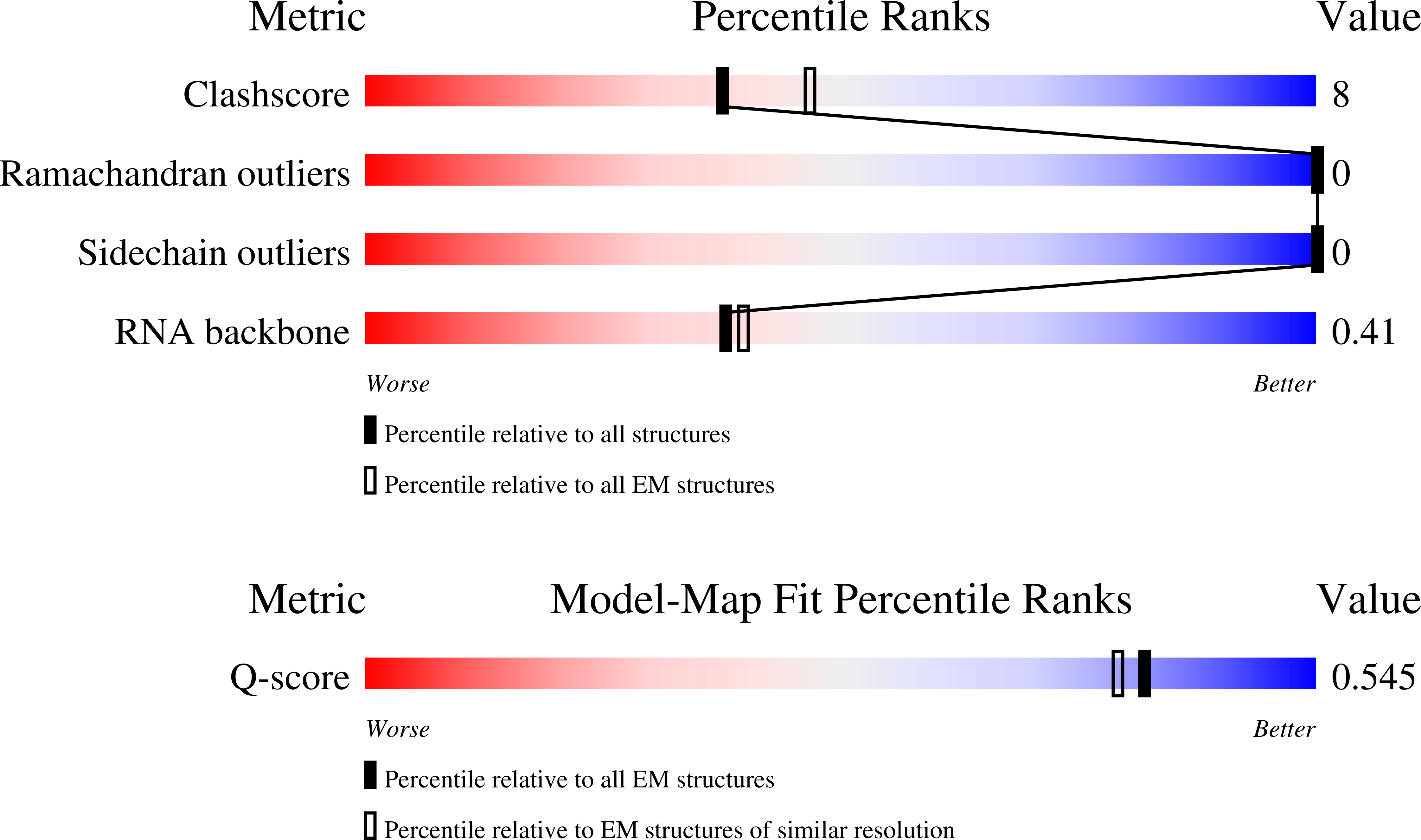
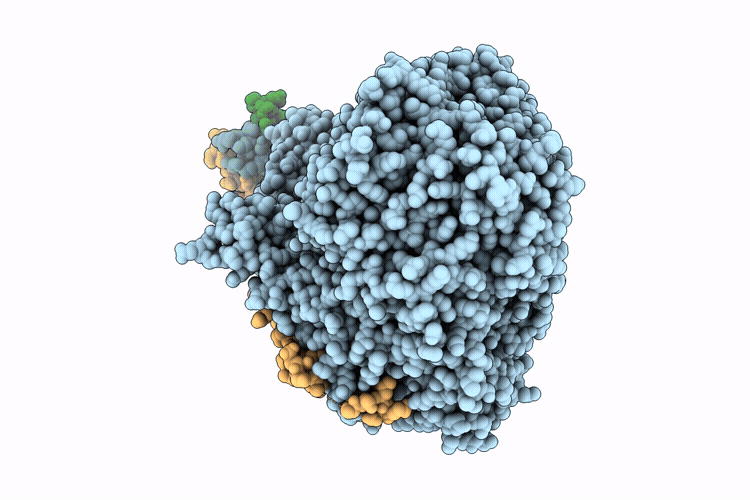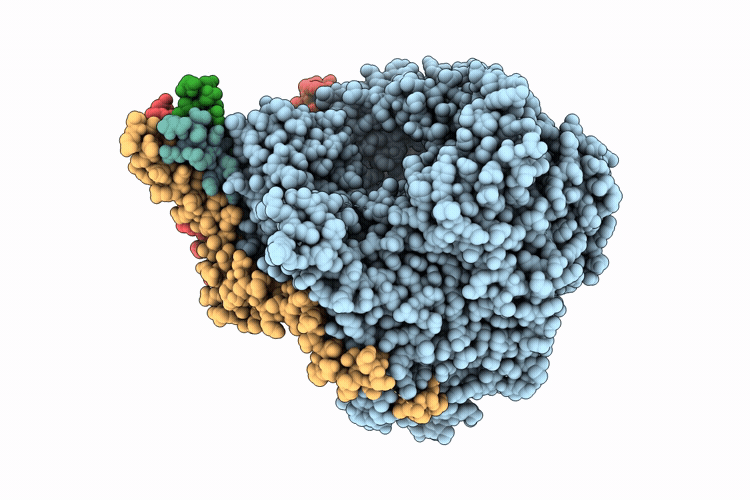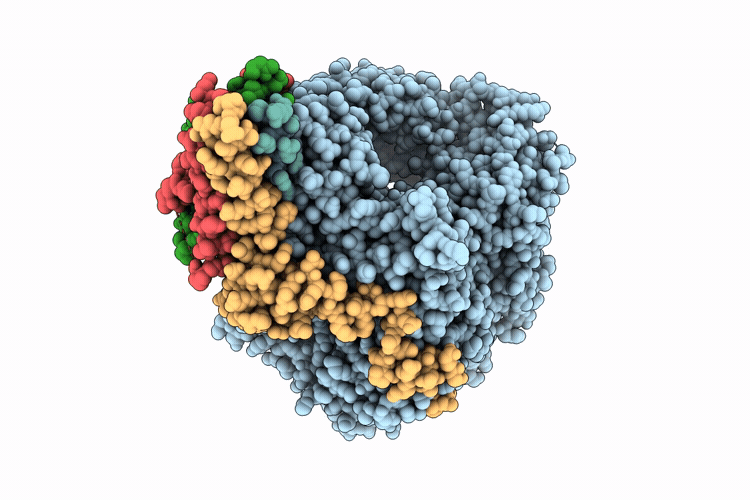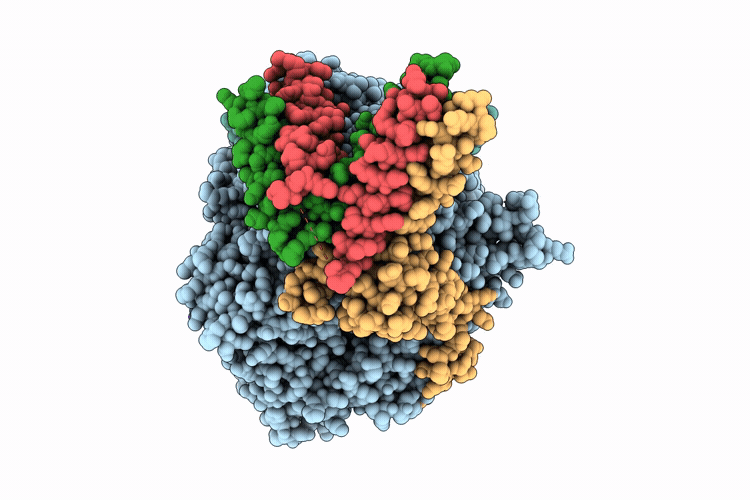
Cryo-EM structure of the respiratory syncytial virus polymerase (L:P) in pre-translocation elongation state
Biological Source:
Source Organism(s):
Respiratory syncytial virus (Taxon ID: 1972429)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
3.07 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE